Multimodal Data Parsing for Materials Informatics: Techniques, Applications, and Future Directions
This article provides a comprehensive overview of multimodal data parsing and its transformative impact on materials informatics.
Multimodal Data Parsing for Materials Informatics: Techniques, Applications, and Future Directions
Abstract
This article provides a comprehensive overview of multimodal data parsing and its transformative impact on materials informatics. It explores the foundational principles of processing heterogeneous data types—including spectral, microscopic, textual, and tabular information—to accelerate materials discovery and characterization. The content covers core methodologies like multimodal fusion and alignment, addresses practical challenges such as handling missing data and ensuring interoperability, and examines validation frameworks for assessing model performance. Designed for researchers, scientists, and drug development professionals, this review synthesizes current trends, highlights practical applications in biomedical research, and outlines a forward-looking perspective on integrating these techniques into intelligent, automated materials development pipelines.
Understanding Multimodal Data Parsing: Core Concepts and Material Science Imperatives
Defining Modality and Multimodal Parsing in Materials Science
Modern materials research generates complex, heterogeneous datasets that span multiple scales and data types, from atomic composition and processing parameters to macroscopic properties and performance characteristics. This inherent complexity necessitates a paradigm shift from single-modality analysis to multimodal learning, an approach that jointly analyzes these diverse data types—or modalities—to uncover deeper insights and overcome the limitations of data scarcity. In artificial intelligence (AI), a modality refers to a specific type or form of data representation and communication [1]. In the specific context of materials science, modalities encompass the diverse types of data generated throughout the material lifecycle, such as chemical composition, synthesis parameters, microstructural images from microscopy, spectral data, and mechanical property measurements [2] [3].
Multimodal parsing is the computational framework that enables the integration, alignment, and joint analysis of these disparate modalities. This process is crucial for modeling the complex, hierarchical relationships in material systems, often described by the processing-structure-properties-performance chain. By effectively parsing multimodal data, researchers can build more robust models that accelerate the discovery and design of novel materials, even when certain data types are incomplete—a common challenge in experimental materials science [2]. This technical guide explores the core concepts, methodologies, and applications of modality and multimodal parsing, providing a foundation for their implementation in advanced materials information research.
Core Concepts and Definitions
Modality in Context
In materials science, the concept of modality extends beyond simple data types to encompass the entire multi-scale characterization of a material. The following table categorizes common modalities encountered in materials research:
Table: Common Modalities in Materials Science Data
| Modality Category | Specific Examples | Typical Data Form |
|---|---|---|
| Composition & Processing | Chemical formula, synthesis parameters (e.g., temperature, flow rate) | Tabular data, numerical vectors [2] |
| Structure & Morphology | Crystal structure, micrographs (SEM, TEM), grain size distribution | Graph representations (for crystal structures), 2D images [2] [3] |
| Properties & Performance | Mechanical properties (e.g., yield strength, modulus), electronic properties, spectral data (XRD, FTIR) | Numerical vectors, line spectra (DOS), time-series data [2] [3] |
| Textual Descriptions | Scientific abstracts, machine-generated crystal descriptions (e.g., from Robocrystallographer) | Natural language text [3] |
The Principle of Multimodal Parsing
Multimodal parsing is the computational engine that transforms a collection of individual modalities into a unified, knowledge-rich representation. It involves several key processes:
- Feature Extraction: Converting raw, high-dimensional data from each modality into meaningful, lower-dimensional feature vectors using specialized encoders (e.g., Convolutional Neural Networks for images, Graph Neural Networks for crystal structures) [2] [3].
- Cross-Modal Alignment: Mapping these extracted features from different modalities into a shared latent space where semantically similar material representations are close to one another, regardless of their original form [2] [3].
- Joint Representation Learning: Fusing the aligned features to create a comprehensive representation that captures the complex, non-linear interactions between different aspects of the material, such as how processing parameters influence microstructure [2].
This integrated approach allows models to reason about materials in a holistic manner, leading to improved performance on predictive tasks and enabling novel discovery workflows.
A Framework for Implementation: MatMCL
The MatMCL (Multimodal Contrastive Learning for Materials) framework provides a concrete architecture for implementing multimodal parsing, specifically designed to handle the challenges of real-world materials data, such as missing modalities [2].
MatMCL employs a structure-guided pre-training (SGPT) strategy, which uses a contrastive learning objective to align representations from different modalities. The core architecture consists of the following components [2]:
- Unimodal Encoders: Specialized neural networks that convert raw data from each modality into a feature vector. For example, a table encoder (e.g., an FT-Transformer or MLP) processes numerical processing parameters, while a vision encoder (e.g., a Vision Transformer or CNN) processes microstructural images from SEM [2].
- Multimodal Encoder: A model (e.g., a transformer with cross-attention) that takes features from multiple modalities and learns a fused, joint representation of the material system [2].
- Projection Head: A shared neural network that maps all representations—unimodal and multimodal—into a joint latent space where contrastive learning is performed [2].
- Downstream Task Modules: Prediction, generation, or retrieval heads that utilize the pre-trained encoders for specific applications like property prediction or microstructure generation [2].
The Experimental Workflow
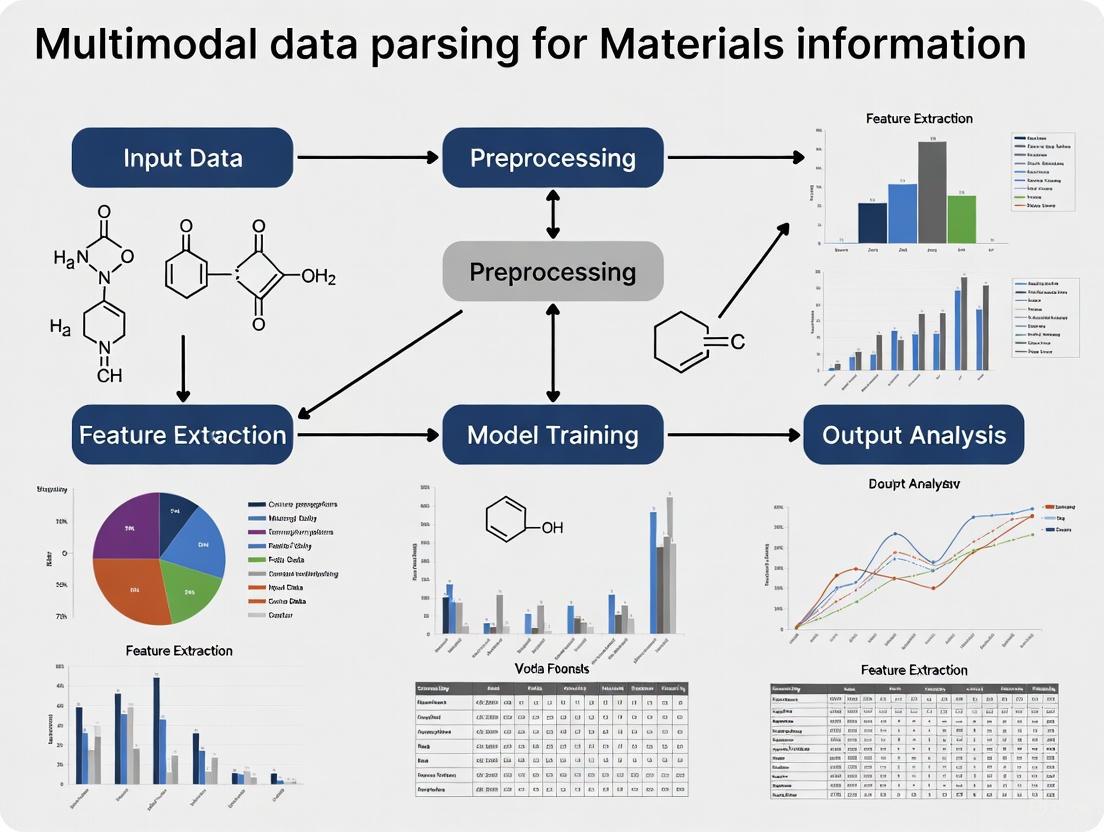
The following diagram illustrates the end-to-end experimental workflow of the MatMCL framework, from data preparation to downstream application:
Detailed Experimental Protocol: Structure-Guided Pre-Training
The SGPT phase is critical for teaching the model the fundamental relationships between modalities. The following protocol is adapted from the electrospun nanofiber case study [2]:
Objective: To learn a joint latent space where representations of the same material from different modalities (e.g., its processing parameters and its microstructure) are aligned closely together.
Input Data: A batch of ( N ) material samples, each with:
- Processing conditions ( { \mathbf{x}i^t }{i=1}^N ) (e.g., flow rate, voltage, concentration)
- Microstructural images ( { \mathbf{x}i^v }{i=1}^N ) (e.g., SEM micrographs)
- Fused input ( { (\mathbf{x}i^t, \mathbf{x}i^v) }_{i=1}^N ) for the multimodal anchor [2].
Procedure:
- Feature Encoding: For each sample ( i ) in the batch, process each modality through its respective encoder to obtain feature vectors:
- Table representation: ( \mathbf{h}i^t = ft(\mathbf{x}i^t) )
- Vision representation: ( \mathbf{h}i^v = fv(\mathbf{x}i^v) )
- Multimodal representation: ( \mathbf{h}i^m = fm(\mathbf{x}i^t, \mathbf{x}i^v) ) [2]
- Projection: Map all feature vectors into the joint latent space using a shared projection network ( g(\cdot) ):
- ( \mathbf{z}i^t = g(\mathbf{h}i^t) ), ( \mathbf{z}i^v = g(\mathbf{h}i^v) ), ( \mathbf{z}i^m = g(\mathbf{h}i^m) ) [2]
- Contrastive Learning: Use the fused representation ( \mathbf{z}_i^m ) as an anchor. Define positive and negative pairs:
- Positive pairs: ( (\mathbf{z}i^m, \mathbf{z}i^t) ) and ( (\mathbf{z}i^m, \mathbf{z}i^v) ) (different views of the same material).
- Negative pairs: ( (\mathbf{z}i^m, \mathbf{z}j^t) ) and ( (\mathbf{z}i^m, \mathbf{z}j^v) ) for ( i \neq j ) (representations from different materials) [2].
- Loss Calculation: Apply a contrastive loss function (e.g., NT-Xent) that minimizes the distance between positive pairs in the latent space while maximizing the distance between negative pairs. This jointly trains all encoders and the projector [2].
Output: Pre-trained and aligned encoders ( ft, fv, f_m ) that can generate meaningful representations even when some modalities are missing during downstream task execution.
The following diagram visualizes the flow of data and the contrastive learning process within the SGPT module:
Essential Research Reagents and Computational Tools
Implementing a multimodal parsing framework requires a suite of computational "reagents" and data sources. The following table details key components and their functions in the research workflow.
Table: Research Reagent Solutions for Multimodal Materials Informatics
| Category | Reagent / Tool | Function in the Workflow |
|---|---|---|
| Data Sources | Materials Project Database [3] | Provides curated, multi-property data on a vast number of crystalline structures, serving as a benchmark for training and validation. |
| Self-Constructed Datasets (e.g., Electrospun Nanofibers [2]) | Provides specialized, experimentally obtained multimodal data (processing, SEM, mechanical properties) for specific material classes. | |
| Encoders | Graph Neural Networks (e.g., PotNet [3]) | Acts as the crystal structure encoder, processing the graph representation of a crystal's atomic structure. |
| Vision Transformers (ViT) / Convolutional Neural Networks (CNN) [2] | Acts as the vision encoder, learning rich features directly from raw microstructural images (e.g., SEM, TEM). | |
| FT-Transformer / Multilayer Perceptron (MLP) [2] | Acts as the table encoder, modeling the non-linear effects of numerical processing parameters and compositions. | |
| Frameworks & Algorithms | Contrastive Learning (e.g., CLIP-inspired [2] [3]) | The core self-supervised algorithm for aligning different modalities in a shared latent space without explicit labels. |
| Multi-stage Learning (MSL) [2] | Extends the pre-trained framework to guide complex design tasks, such as composite material design. | |
| Validation & Analysis | Tensile Testing [2] | Provides ground-truth mechanical property data (e.g., fracture strength, elastic modulus) for model validation. |
| Cross-Modal Retrieval Module [2] | Enables quantitative testing of model understanding by retrieving relevant information across different modalities. |
Performance and Quantitative Validation
Rigorous validation is essential to demonstrate the superiority of multimodal parsing over traditional single-modality approaches. In the case study on electrospun nanofibers, the MatMCL framework was evaluated on several downstream tasks [2].
Table: Performance Metrics for Downstream Tasks in Multimodal Parsing
| Downstream Task | Key Metric | Performance / Outcome |
|---|---|---|
| Mechanical Property Prediction | Prediction Accuracy (with missing structural data) | MatMCL showed improved prediction of mechanical properties (e.g., fracture strength, elastic modulus) even when microstructural image data was unavailable during inference, highlighting its robustness to incomplete data [2]. |
| Conditional Structure Generation | Quality of Generated Microstructures | The framework successfully generated realistic microstructures from a given set of processing parameters, demonstrating its understanding of processing-structure relationships [2]. |
| Cross-Modal Retrieval | Retrieval Accuracy | MatMCL enabled accurate retrieval of relevant processing conditions when queried with a microstructure image, and vice-versa, proving effective knowledge extraction across modalities [2]. |
| Material Discovery | Identification of Stable, Novel Candidates | The MultiMat framework, a related approach, demonstrated novel material discovery by screening for stable materials with desired properties through latent space similarity searches [3]. |
The adoption of multimodal parsing represents a transformative advancement in computational materials science. By moving beyond single-modality analysis, frameworks like MatMCL and MultiMat provide a powerful methodology for modeling the complex, hierarchical relationships that define material behavior [2] [3]. The core strength of this approach lies in its ability to leverage the complementary nature of diverse data types, creating AI models that are not only more accurate but also more robust to the incomplete datasets typical of experimental research.
This paradigm enables novel scientific workflows, from predicting properties with missing data to generating new structures and discovering materials with targeted characteristics. As materials data continues to grow in volume and variety, the principles of defining modality and implementing effective multimodal parsing will become increasingly central to accelerating the design and discovery of next-generation materials.
The Critical Role of Parsing in Establishing Processing-Structure-Property Relationships
In materials science, the fundamental paradigm for designing new materials revolves around understanding the Processing-Structure-Property (PSP) relationships. Establishing these relationships requires integrating and interpreting diverse, complex data generated throughout the materials lifecycle. Parsing, the computational process of extracting structured information from raw, often heterogeneous data sources, serves as the critical foundation for this undertaking. Within the context of modern materials informatics, effective parsing enables the transformation of multimodal data into actionable knowledge, thereby accelerating the discovery and development of advanced materials [4] [5].
The challenge is particularly pronounced because materials data is inherently multimodal. It spans computational outputs from density functional theory (DFT) and molecular dynamics (MD) simulations [6], experimental characterizations from techniques like solid-state NMR [5], architectural drawings [7], and textual specifications. This article provides an in-depth technical examination of parsing methodologies that underpin the establishment of robust PSP relationships, framing them within a broader thesis on multimodal data parsing for materials information research.
Parsing Fundamentals for Multimodal Data
The core objective of parsing in materials science is to convert unstructured or semi-structured data into a structured, machine-readable format that can be integrated into predictive models. This process is the first and most critical step in the materials informatics pipeline, as the quality of the parsed data directly dictates the performance of downstream machine learning models [4] [8].
A sophisticated parsing workflow must handle several data modalities, each with its own unique structure and interpretation challenges. The following diagram illustrates a generalized parsing workflow for heterogeneous materials data, from raw input to structured knowledge.
Methodologies for Parsing Diverse Data Modalities
Parsing Computational Materials Data
Computational simulations like Density Functional Theory (DFT) and Molecular Dynamics (MD) generate complex, multi-step data. The FireWorks workflow software provides a structured approach to parsing this data by modeling computational workflows as Directed Acyclic Graphs (DAGs) [6]. Each computational job ("Firework") contains atomic tasks ("Firetasks") that execute sequentially, with dependencies explicitly defined. At the workflow's conclusion, an analysis FireTask parses all output files, extracts relevant properties, and generates a standardized JSON report or MongoDB document. This structured parsing transforms raw simulation outputs into a queryable database, directly linking computational processing conditions to predicted material structures and properties [6].
Parsing Spectral Data for Structure Analysis
Solid-state NMR (ssNMR) spectra provide critical information about domain structures in polymers but present parsing challenges due to broad, overlapping spectral peaks from domains with different molecular mobility. A developed parsing methodology uses Short-Time Fourier Transform (STFT) to decompose free-induction decay (FID) signals into time-frequency components [5]. The parsing workflow involves:
- Signal Transformation: Applying STFT to 1H-static ssNMR FIDs to obtain frequency, intensity, and T2 relaxation time.
- Component Separation: Fitting the time-frequency data to separate domain components based on distinct T2 relaxation times: Mobile (~0.96 ms), Intermediate (Mobile) (~0.55 ms), Intermediate (Rigid) (~0.32 ms), and Rigid (~0.11 ms).
- Ratio Calculation: Using 3D modeling and Bayesian optimization to calculate the volume ratios of each domain component, minimizing error between simulated and original data.
- Data Integration: The parsed domain ratios are integrated with meta-information (elements, functional groups, thermophysical properties) for subsequent analysis using self-organizing maps (SOM) and market basket analysis [5].
Parsing Textual and Geometric Data
Technical documents contain crucial processing information in textual and tabular formats. Parsing this data employs a hybrid approach:
- Textual Parameter Extraction: Combining regular expressions, domain-specific terminology dictionaries, and a BiLSTM-CRF deep learning model to achieve high-precision extraction of material parameters and properties from unstructured text [7]. This method reported a precision of 83.56% and recall of 86.91% for parameter extraction.
- Geometric Data Parsing: For DXF drawing files, parsing involves vector element analysis, layer semantic analysis, and spatial topological relationship reconstruction. This enables structured extraction of key component geometry, achieving F1 scores of 98.1% for wall line recognition and 92.2% for column recognition [7].
Table 1: Performance Metrics of Multimodal Data Parsing Methods
| Data Modality | Parsing Method | Key Performance Metric | Reported Value |
|---|---|---|---|
| Spectral Data (ssNMR) | STFT + Bayesian Optimization | T2 Relaxation Time Resolution | 4 distinct domains separated [5] |
| Textual Specifications | BiLSTM-CRF Model | Precision / Recall | 83.56% / 86.91% [7] |
| Architectural Drawings | Vector & Topology Analysis | F1 Score (Wall Lines) | 98.1% [7] |
| Architectural Drawings | Vector & Topology Analysis | F1 Score (Columns) | 92.2% [7] |
| Tabular Data | Multi-scale Sliding Window | Recall (Door/Window Params) | 95.0% [7] |
Establishing Processing-Structure-Property Relationships Through Parsed Data
Interpretable Deep Learning for Structure-Property Mapping
Once data is parsed into structured representations, it can be used to train models that map material structures to properties. The Self-Consistent Attention Neural Network (SCANN) architecture exemplifies this approach, using an attention mechanism to predict properties and interpret structure-property relationships [4]. SCANN operates by:
- Representing Local Structures: For each atom in a material structure, Voronoi tessellation identifies neighboring atoms. The geometrical influence of each neighbor is encoded as a vector based on Euclidean distance and Voronoi solid angle.
- Recursive Learning: A series of local attention layers recursively refine the representation of each atom's local environment by applying attention mechanisms to its neighbors' representations.
- Global Representation: A global attention layer combines these local representations into a complete material structure representation, quantitatively measuring the contribution (attention) of each local structure to the global property [4].
This architecture not only achieves prediction accuracy comparable to state-of-the-art models but also provides interpretability by identifying which local atomic environments most significantly influence specific properties like molecular orbital energies or formation energies.
Integrated Analysis for Polymer Design
In polymer science, parsing enables the direct linking of processing conditions to domain structures and final properties. The domain ratios parsed from ssNMR data serve as structural descriptors. For instance, analysis reveals that poly(ε-caprolactone) (PCL) contains a high proportion (37.7%) of Mobile domains, while poly(3-hydroxybutyrate-co-3-hydroxyhexanoate) (PHBH) is dominated by Rigid domains (50.5%) [5]. These parsed structural metrics are then integrated with processing parameters and performance data using methods like self-organizing maps (SOM) and market basket analysis to uncover complex, non-linear PSP relationships that guide the design of polymers with tailored properties.
Table 2: Experimental Protocol for Parsing-Based PSP Relationship Analysis
| Protocol Step | Technical Specification | Purpose/Function |
|---|---|---|
| Data Acquisition | 1H-static ssNMR, DFT/MD simulations, textual specifications | Generate raw, multimodal data on material processing and characterization [5] [6] |
| Data Preprocessing | Vector cleanup, layer standardization, signal filtering | Remove noise, correct misalignments, and standardize data formats [7] [5] |
| Modality-Specific Parsing | STFT, BiLSTM-CRF, FireWorks DAGs, geometric feature analysis | Extract structured descriptors (domain ratios, formation energies, geometric parameters) [5] [7] [6] |
| Data Integration | JSON/MongoDB documentation, feature vector concatenation | Create unified representation of processing, structure, and property parameters [6] [5] |
| Relationship Modeling | SCANN, SOM, Market Basket Analysis, XGBoost | Identify and model complex, non-linear PSP relationships [4] [5] [8] |
| Validation | First-principles calculations, train-test-split validation | Confirm predictive accuracy and physical meaningfulness of parsed descriptors and models [4] |
The Scientist's Toolkit: Essential Research Reagents & Solutions
Table 3: Key Research Reagent Solutions for Parsing-Driven Materials Research
| Reagent / Tool | Function / Application | Technical Notes |
|---|---|---|
| FireWorks Workflow Software | Parsing and managing DFT/MD computational workflows as Directed Acyclic Graphs (DAGs) | Enables execution tracking, data provenance, and standardized output parsing into JSON/MongoDB [6] |
| STFT + Bayesian Optimization Package | Deconvoluting overlapping ssNMR spectra to resolve domain-specific T2 relaxation times | Critical for parsing domain mobility distribution in polymers; requires optimization to minimize fitting error [5] |
| BiLSTM-CRF Model | Named Entity Recognition (NER) for extracting material parameters from unstructured text | Combines deep learning (BiLSTM) with rule-based constraints (CRF); domain dictionary enhances accuracy [7] |
| Vector & Topology Analysis Library | Parsing DXF files to extract component geometry and spatial relationships | Relies on layer semantic analysis and spatial topology reconstruction for automated 3D model generation [7] |
| SCANN Framework | Interpretable deep learning for structure-property prediction with attention mechanisms | Uses Voronoi tessellation for local structure definition; provides atomic-level insight into property determinants [4] |
| Self-Organizing Map (SOM) | Visualizing and clustering high-dimensional parsed material data | Reveals hidden patterns in the integrated space of processing parameters, structural descriptors, and properties [5] |
Parsing is the foundational enabler for establishing quantitative Processing-Structure-Property relationships in modern materials science. By transforming multimodal, heterogeneous data—from computational outputs and spectral signatures to textual descriptions and geometric layouts—into structured, interoperable descriptors, parsing bridges the gap between raw data and actionable knowledge. The methodologies and tools detailed in this technical guide, from interpretable deep learning architectures like SCANN to specialized parsing protocols for NMR and textual data, provide researchers with a reproducible framework to accelerate the design and discovery of next-generation materials. As materials data continues to grow in volume and complexity, the critical role of advanced parsing will only intensify, making it an indispensable component of the materials informatics paradigm.
In materials science and informatics, the systematic acquisition and analysis of data are fundamental to establishing the critical processing-structure-property-performance relationships that govern material behavior [9]. Modern research generates a plethora of data types, each capturing distinct aspects of material characteristics. This technical guide provides a comprehensive overview of four principal data modalities—spectral, microscopic, textual, and tabular—framed within the context of multimodal data parsing for accelerated materials discovery and development. The integration of these diverse data types through advanced artificial intelligence and machine learning methods is revolutionizing how researchers approach materials design, particularly in pharmaceutical development where material properties directly impact drug efficacy, stability, and delivery [10] [9].
Table 1: Core Material Data Modalities in Materials Informatics
| Data Modality | Primary Information Captured | Common Acquisition Techniques | Typical Data Structure |
|---|---|---|---|
| Spectral | Chemical composition, molecular structure, functional groups | UV-vis, NIR, IR, Raman, XRF, LIBS [11] | Hyperspectral cube (x, y, λ) [11] |
| Microscopic | Morphology, microstructure, spatial distribution | Optical microscopy, electron microscopy, scanning probe microscopy [12] | High-resolution 2D/3D image data [12] |
| Textual | Experimental observations, synthesis procedures, literature knowledge | Scientific publications, lab notebooks, technical manuals [13] | Unstructured or semi-structured text [14] |
| Tabular | Quantitative measurements, material properties, composition data | High-throughput experimentation, computational simulations [15] | Structured tables with rows and columns [15] |
Spectral Data
Fundamental Principles and Acquisition
Spectral data encompasses measurements of how materials interact with electromagnetic radiation across various wavelengths [16]. The foundational principle of spectroscopy involves studying light-matter interactions to obtain detailed information about reflectance, emission, or absorption properties [16]. Each material possesses a unique spectral signature—akin to a fingerprint—that enables identification based on chemical composition and physical characteristics [16]. Spectral sensors (spectrometers) capture and measure light reflected or emitted by objects in the form of reflectance spectra, which are typically presented as graphs of intensity versus wavelength [16].
Advanced spectroscopic imaging integrates spatial information with chemical or physical data, enabling comprehensive material characterization [11]. This is achieved through the creation of a hyperspectral data cube, where the X and Y axes represent spatial dimensions and the Z-axis represents spectral information across wavelengths [11]. The process involves systematically measuring spectra across sample surfaces, either through physical rastering, scanning optics with array detectors, or selective subsampling [11].
Experimental Methodology: Hyperspectral Data Cube Construction
Protocol Title: Construction of Hyperspectral Data Cubes for Material Characterization
Objective: To generate a three-dimensional hyperspectral data cube integrating spatial and spectral information for comprehensive material analysis.
Materials and Equipment:
- Illumination source covering relevant wavelength ranges
- Spectrometer with appropriate spectral range (UV-vis, NIR, IR, etc.)
- Positioning system for precise sample movement
- Data acquisition software
- Sample mounting apparatus
Procedure:
- Sample Preparation: Mount the target material securely to ensure stability during measurement.
- System Calibration: Perform wavelength and intensity calibration using standard reference materials.
- Data Acquisition:
- Illuminate the sample with a broadband light source
- Collect light interacting with the sample (reflected, emitted, or transmitted)
- Measure intensity (I) at each wavelength (λ) for an initial spatial point
- Systematically move the spectrometer across the sample surface using raster scanning
- Repeat spectral measurement for each spatial location on the XY-plane
- Data Cube Construction: Compile individual spectra into a stacked structure where:
- X and Y dimensions represent spatial coordinates
- Z dimension represents spectral wavelengths
- Each pixel contains complete spectral information for its spatial location
- Data Processing: Apply chemometrics and machine learning algorithms to extract chemical and physical parameters [11]
Technical Considerations: The specific wavelength range should be selected based on the application—UV (190-360 nm) for electronic transitions, visible (360-780 nm) for color analysis, NIR for overtone vibrations, and IR for molecular vibrations [11].
Research Reagent Solutions for Spectral Analysis
Table 2: Essential Resources for Spectral Data Acquisition and Analysis
| Resource Category | Specific Examples | Function/Purpose |
|---|---|---|
| Spectral Sensors | Point spectrometers, Imaging spectrometers (hyperspectral cameras) [16] | Capture and measure light spectra from materials |
| Wavelength Ranges | VNIR (400-1000 nm), NIR (1000-1700 nm), SWIR (1000-2500 nm), MWIR (3000-5000 nm), LWIR (8000-12000 nm) [16] | Target specific molecular vibrations and transitions |
| Accessories | Integrating spheres, cosine correctors, cuvettes, fiber optics [17] | Enable specific measurement geometries and sample types |
| Analysis Software | Chemometrics packages, machine learning algorithms [11] | Extract meaningful information from complex spectral data |
Microscopic Data
Imaging Modalities and Applications
Microscopic data provides structural information across multiple scales, from atomic arrangements to microstructural features [12]. Modern microscopy techniques generate high-resolution images that reveal critical insights into material morphology, phase distribution, grain boundaries, and defect structures [12]. The primary challenge in contemporary microscopy is the conversion of "big visual data" into interpretable information, as automated microscopy systems can acquire thousands of images within hours, far exceeding human analysis capacity [12].
Key microscopy modalities include:
- Optical Microscopy: For microstructural analysis at micrometer resolution
- Electron Microscopy: For nanoscale and atomic-level resolution
- Scanning Probe Microscopy: For surface topography and properties
- Fluorescence Microscopy: For specific component identification in biological materials
Experimental Methodology: Grain Segmentation in Polycrystalline Materials
Protocol Title: Deep Learning-Based Segmentation of Grain Structures in Microscopic Images
Objective: To accurately segment grain boundaries and instances in polycrystalline materials using a transfer learning approach with synthetic data augmentation.
Materials and Equipment:
- Polycrystalline material samples (e.g., iron)
- Optical microscope with digital imaging capabilities
- Sample preparation equipment (polishing, etching)
- Computing hardware with GPU acceleration
- Monte Carlo Potts simulation software
- Image style transfer model (GAN-based)
Procedure:
- Real Data Acquisition:
- Prepare material samples through polishing and etching
- Capture optical images of polycrystalline structure (e.g., 136 serial sections at 2800×1600 resolution)
- Manually annotate grain boundaries to create ground truth labels (2 semantic classes: grain and grain boundary)
- Split data into training (100 images) and test (36 images) sets
- Pre-process into 400×400 pixel patches for computational efficiency [18]
Simulated Data Generation:
- Establish 3D simulated model of polycrystalline materials using Monte Carlo Potts model
- Generate 2D images by slicing simulated 3D image in normal direction
- Extract boundary pixels of each grain to obtain simulated labels
- Ensure geometric and topological consistency with real data [18]
Synthetic Data Creation:
- Train image style transfer model (GAN) using real dataset
- Transform simulated label images into synthetic images incorporating real image features
- Validate that synthetic images maintain label information while adopting realistic appearance [18]
Model Training and Validation:
- Train segmentation models (e.g., U-Net architecture) using combinations of real, simulated, and synthetic data
- Evaluate performance on held-out test set using accuracy metrics
- Demonstrate competitive performance with model trained on synthetic data plus 35% real data versus 100% real data [18]
Technical Considerations: This approach addresses the data scarcity problem in material microscopy by leveraging physical simulations and transfer learning, significantly reducing experimental burden while maintaining segmentation accuracy [18].
Research Reagent Solutions for Microscopic Analysis
Table 3: Essential Resources for Microscopic Data Acquisition and Analysis
| Resource Category | Specific Examples | Function/Purpose |
|---|---|---|
| Microscopy Platforms | Optical microscopes, Electron microscopes (SEM, TEM), Scanning probe microscopes [12] | High-resolution imaging at appropriate scales |
| Sample Preparation | Polishing equipment, Etching solutions, Coating systems | Prepare samples for optimal imaging quality |
| Analysis Software | ImageJ, Cell tracking algorithms, Deep learning frameworks (PyTorch, TensorFlow) [12] | Automated analysis of complex microscopic data |
| Simulation Tools | Monte Carlo Potts model, Phase-field simulations [18] | Generate synthetic data for training models |
Textual Data
Characteristics and Applications in Materials Science
Textual data in materials science encompasses a wide range of unstructured and semi-structured information, including scientific publications, experimental protocols, laboratory notebooks, technical manuals, and patent documents [13]. This modality captures critical contextual knowledge about synthesis procedures, experimental observations, material processing conditions, and research outcomes that may not be fully represented in structured data formats.
The primary challenge with textual materials lies in designing them for optimal usability by specific target audiences [13]. Effective text design must consider audience characteristics, purpose, and context of use—what works for expert researchers may not be suitable for cross-cultural applications or those with different linguistic, educational, or intellectual backgrounds [13].
Text Design and Management Methodologies
Protocol Title: Structured Approach to Technical Textual Material Design and Management
Objective: To create usable and effective textual materials tailored to specific audience needs and research purposes within materials science contexts.
Materials and Equipment:
- Content management systems
- Text analysis software
- Digital publishing platforms
- Cross-cultural review resources
Procedure:
- Audience Analysis:
- Identify primary and secondary user groups
- Assess linguistic capabilities, educational backgrounds, and technical expertise
- Determine cultural considerations for international audiences
Purpose Definition:
- Clarify primary use cases (learning, reference, procedure execution)
- Avoid purpose confusion that leads to usability compromises
- Establish clear success metrics for textual effectiveness
Content Structuring:
- Select appropriate format based on purpose:
- Integrated text-graphics for procedures
- Vertical/horizontal division for balanced content
- Variable illustration sizing for complex concepts
- Sparse graphics with heavy text for expert users
- Implement consistent organizational frameworks
- Select appropriate format based on purpose:
Usability Enhancement:
- Incorporate learning principles: advanced organizers, concrete examples, self-check questions
- Utilize formatting options: headings, bold/italic emphasis, quotes, URLs
- Support multiple text value types: String (short text), Text (multi-line), Text with layout, Text with tags and layout [14]
Validation and Iteration:
- Conduct usability testing with target audience representatives
- Gather feedback on clarity, effectiveness, and accessibility
- Revise materials based on empirical observations
Technical Considerations: The four common formats for technical procedural information include: (1) completely integrated text and graphics, (2) vertically or horizontally divided frames, (3) variable-sized illustrations supporting text, and (4) sparse graphics with extensive text for familiar content [13].
Tabular Data
Structure and Standards in Materials Informatics
Tabular data represents structured information organized in rows and columns, forming the backbone of quantitative materials informatics [15]. This modality efficiently captures measured properties, compositional information, processing parameters, and performance characteristics in a format amenable to statistical analysis and machine learning applications.
The preferred format for tabular data in materials informatics is comma-separated values (CSV), which offers cross-platform compatibility and programmatic accessibility compared to proprietary spreadsheet formats [15]. The pandas library in Python has emerged as the standard tool for manipulating tabular data, providing powerful structures like DataFrames (for 2D heterogeneous data) and Series (for 1D homogeneous data) [15].
Methodologies for Tabular Data Management
Protocol Title: Best Practices for Tabular Data Structure and Analysis in Materials Research
Objective: To create, manage, and analyze tabular materials data following FAIR (Findable, Accessible, Interoperable, Reusable) principles for maximum research impact.
Materials and Equipment:
- Python programming environment with pandas library
- Data visualization tools (Matplotlib, Plotly, Seaborn)
- Version control system (Git)
- Data repositories for sharing and preservation
Procedure:
- Data Structure Design:
- Define clear column names representing specific variables
- Ensure consistent data types within columns
- Include metadata describing units, measurement conditions, and uncertainties
- Establish primary keys for data record uniqueness
Data Creation and Import:
- Use pd.DataFrame() constructor with dictionary of columnname:valuelist pairs
- Alternatively, create from 2D arrays with explicit column naming
- For existing data, utilize pd.read_csv() with parameters:
- sep=',' for comma separation (or other delimiters)
- skiprows to exclude header comments
- names for column renaming
- nrows for large file handling
Data Validation:
- Verify data types and ranges for each column
- Check for missing values and inconsistencies
- Confirm unit consistency across related datasets
- Validate against known physical constraints and relationships
Exploratory Data Analysis:
- Compute descriptive statistics (mean, median, standard deviation)
- Generate visualizations (scatter plots, histograms, correlation matrices)
- Identify outliers and anomalous patterns
- Explore relationships between processing parameters and properties
Data Export and Sharing:
- Export to CSV for broad accessibility
- Include comprehensive metadata documentation
- Apply appropriate data licenses and citation information
- Deposit in community-recognized repositories
Technical Considerations: Tabular data provides the foundation for establishing processing-structure-property-performance relationships in materials science [15]. Properly structured tables enable efficient implementation of machine learning algorithms for materials prediction and optimization [9].
Multimodal Data Integration Framework
The true power of materials informatics emerges from the integration of multiple data modalities into a unified analytical framework. Deep learning methods have demonstrated remarkable capabilities in processing and correlating heterogeneous data types, including atomistic, image-based, spectral, and textual information [9]. This multimodal approach enables the establishment of comprehensive structure-property relationships that would be difficult to discern from individual data sources alone.
Integrated Experimental Methodology
Protocol Title: Multimodal Data Parsing for Materials Property Prediction
Objective: To integrate spectral, microscopic, textual, and tabular data modalities for accelerated materials discovery and property prediction.
Materials and Equipment:
- Cross-modal data integration platform
- Deep learning framework (PyTorch, TensorFlow)
- High-performance computing resources
- Multimodal materials dataset
Procedure:
- Data Collection:
- Acquire spectral data capturing chemical composition
- Collect microscopic images revealing microstructure
- Extract textual information from synthesis protocols and literature
- Compile tabular data containing measured properties
Data Preprocessing:
- Normalize each modality to common scale and representation
- Extract features using modality-specific encoders:
- Convolutional Neural Networks (CNNs) for images
- Recurrent Neural Networks (RNNs) or transformers for text
- Spectral convolution networks for spectroscopic data
- Create aligned multimodal representations
Model Architecture Design:
- Implement cross-modal attention mechanisms
- Design fusion layers for integrated representation learning
- Incorporate uncertainty quantification methods
- Enable interpretability for scientific insight generation
Training and Validation:
- Utilize transfer learning to address data scarcity
- Apply multi-task learning for related property prediction
- Implement cross-validation strategies
- Benchmark against physics-based models and experimental results
Knowledge Extraction:
- Identify dominant features influencing target properties
- Discover new material design rules
- Generate hypotheses for experimental validation
- Update models with new data in continuous learning framework
Technical Considerations: Multimodal data integration faces challenges including data heterogeneity, varying scales, and different levels of uncertainty [9]. Deep learning approaches can help bridge these gaps through representation learning and cross-modal alignment, potentially uncovering previously unrecognized relationships in materials behavior [10] [9].
The systematic characterization and integration of spectral, microscopic, textual, and tabular data modalities represents a transformative approach to materials research and development. Each modality offers unique insights—spectral data reveals chemical composition, microscopic data captures structural features, textual data provides contextual knowledge, and tabular data enables quantitative analysis. The emerging paradigm of multimodal data parsing, powered by advanced deep learning methods, is accelerating materials discovery by establishing comprehensive processing-structure-property relationships across diverse data types. For researchers in pharmaceutical development and materials science, mastering these data modalities and their integration is becoming increasingly essential for addressing complex challenges in drug formulation, delivery system design, and material performance optimization. As materials informatics continues to evolve, the development of standardized protocols, shared data resources, and interoperable analysis frameworks will further enhance our ability to extract meaningful insights from multimodal materials data.
The convergence of artificial intelligence (AI) and materials science has given rise to the field of materials informatics, which promises to significantly accelerate the discovery and design of novel materials [19]. However, the real-world application of AI in materials science faces three fundamental challenges: the scarcity of high-quality experimental data, the inherent heterogeneity of multimodal data, and the multiscale complexity of material systems [2]. These challenges are particularly acute when seeking to establish processing-structure-property-performance relationships, a core objective of materials research. This whitepaper details these challenges and presents advanced computational frameworks, including multimodal learning and transfer learning, which are designed to overcome these obstacles within the context of multimodal data parsing for materials informatics.
Data Scarcity in Materials Science
Data scarcity is a pervasive issue that substantially limits the predictive reliability of AI models in materials science. This scarcity primarily stems from the high cost and complexity of material synthesis and characterization, which naturally limits the volume of available data [2].
Strategies to Overcome Data Scarcity
Transfer Learning is a powerful technique for bridging sparse datasets. It involves using information from one dataset to inform a model on another, which preserves contextual differences in underlying measurements. The table below summarizes three key transfer learning architectures and their effectiveness in different materials science contexts [20].
Table 1: Transfer Learning Architectures for Overcoming Data Scarcity
| Architecture Type | Description | Best-Suited Application |
|---|---|---|
| Multi-task | Simultaneously learns multiple related tasks, sharing representations between them. | Most improves classification performance (e.g., of color with band gaps). |
| Difference | Models the difference between data sources or fidelity levels. | Most accurate for multi-fidelity data (e.g., mixed DFT and experimental band gaps). |
| Explicit Latent Variable | Learns an explicit latent variable representing hidden contextual factors. | Most accurate for complex relationships; enables cancellation of errors in functions depending on multiple tasks (e.g., activation energies in NO reduction). |
Multimodal Learning (MML) also serves as a potent remedy for data scarcity. By integrating multiple types of data (modalities), such as processing parameters and microstructural images, MML enhances the model's understanding of complex material systems and mitigates the limitations of small datasets [2].
Experimental Protocol: Leveraging Transfer Learning
- Data Preparation: Assemble the primary dataset (
Dataset A) for the target property, which is small, and a larger, related secondary dataset (Dataset B). - Model Pre-training: Train a base model on
Dataset B. This process allows the model to learn general, transferable features and patterns. - Model Fine-tuning: Use the weights from the pre-trained model to initialize a new model for the target task. Fine-tune this model on
Dataset A. This allows the model to adapt its previously learned knowledge to the specific, data-scarce problem [20].
Data Heterogeneity
Data heterogeneity in materials informatics refers to the challenges of managing and processing data that varies dramatically in format, size, content, and structure, often distributed across multiple institutions [21]. This includes the model heterogeneity and data heterogeneity encountered when training complex AI models.
Manifestations of Data Heterogeneity
- Multimodal, Multi-institutional Data: Combinatorial materials science produces large, complex datasets from synthesis, processing, and characterization (e.g., X-ray diffraction). These datasets are often distributed and vary substantially in format and content, creating a significant data management and integration challenge [21].
- Model Heterogeneity in AI Training: In Multimodal Large Language Models (LLMs), different modules (e.g., modality encoders, LLM backbones, modality generators) vary dramatically in size and operator complexity. This heterogeneity introduces severe pipeline bubbles during training, leading to poor GPU utilization [22].
- Data Heterogeneity in AI Training: The intricate and unstructured nature of multimodal input data leads to training "stragglers," where some data batches take longer to process than others. This prolongs training duration and exacerbates pipeline bubbles [22].
Solutions for Data Heterogeneity
Technical Framework: The MMScale framework is designed to address heterogeneity in multimodal LLM training. Its core techniques are [22]:
- Adaptive Resource Allocation (for Model Heterogeneity): This technique tailors resource allocation and parallelism strategies separately for each model module (encoder, backbone, generator) to minimize pipeline bubbles.
- Data-Aware Reordering (for Data Heterogeneity): This involves strategically reordering training data at two levels—inter-microbatch and intra-microbatch—to evenly distribute computational load and minimize training delays.
Data Management Infrastructure: Developing standardized dashboards and data infrastructures is crucial for organizing, analyzing, and visualizing large "data lakes" composed of heterogeneous combinatorial datasets [21]. The guiding principle is to prioritize standardisation and the creation of FAIR (Findable, Accessible, Interoperable, and Reusable) data repositories [19].
Multiscale Complexity
Real-world material systems exhibit a hierarchical nature, characterized by multiple scales of information—from atomic composition and microstructure to macroscopic properties [2]. This multiscale complexity poses a significant challenge for AI models to accurately represent and integrate these correlated features.
A Framework for Multiscale, Multimodal Learning
The MatMCL framework is a versatile multimodal learning approach specifically designed to tackle multiscale complexity [2].
Table 2: Core Modules of the MatMCL Framework
| Module Name | Function | Key Benefit |
|---|---|---|
| Structure-Guided Pre-training (SGPT) | Aligns processing and structural modalities via a fused material representation using contrastive learning. | Enables robust property prediction even when structural information is missing. |
| Property Prediction | Predicts material properties from the aligned multimodal representations. | Improves prediction accuracy without requiring complete structural data. |
| Cross-Modal Retrieval | Allows for querying and extracting knowledge across different modalities. | Uncovers processing-structure-property relationships. |
| Conditional Structure Generation | Generates microstructures from given processing parameters. | Facilitates the inverse design of materials. |
Experimental Protocol: Structure-Guided Pre-training for Multiscale Learning
- Data Input: For a batch of N samples, input the processing conditions ( {\mathbf{x}{i}^{t}}{i=1}^{N} ) and microstructure images ( {\mathbf{x}{i}^{v}}{i=1}^{N} ).
- Modality Encoding:
- Process the tabular processing parameters with a table encoder (e.g., an FT-Transformer or MLP) to get representations ( {\mathbf{h}{i}^{t}}{i=1}^{N} ).
- Process the microstructure images with a vision encoder (e.g., a Vision Transformer or CNN) to get representations ( {\mathbf{h}{i}^{v}}{i=1}^{N} ).
- Multimodal Fusion: Fuse the processing and structure representations ( {\mathbf{x}{i}^{t}, \mathbf{x}{i}^{v}}{i=1}^{N} ) using a multimodal encoder (e.g., a Transformer with cross-attention) to obtain fused embeddings ( {\mathbf{h}{i}^{m}}_{i=1}^{N} ).
- Contrastive Alignment: Project all representations into a joint latent space using a shared projector. Use the fused representation ( \mathbf{z}{i}^{m} ) as an anchor and align it with its corresponding unimodal representations ( \mathbf{z}{i}^{t} ) and ( \mathbf{z}_{i}^{v} ) (positive pairs), while pushing away representations from other samples (negative pairs). This self-supervised step forces the model to learn the underlying correlations between processing conditions and microstructure [2].
The Scientist's Toolkit: Essential Research Reagents & Solutions
The following table details key computational tools and resources essential for implementing the advanced frameworks discussed in this whitepaper.
Table 3: Key Research Reagents & Computational Solutions
| Tool/Resource Name | Type | Function/Purpose |
|---|---|---|
| MMScale [22] | Training Framework | An efficient and adaptive framework for training Multimodal LLMs that addresses model and data heterogeneity to achieve high efficiency on large-scale GPU clusters. |
| MatMCL [2] | Multimodal Learning Framework | A versatile MML framework that handles missing modalities and facilitates cross-modal interaction and transformation for multiscale material systems. |
| Multi-task/Difference/Explicit Latent Variable Architectures [20] | Transfer Learning Model | Specific neural network architectures designed to effectively perform transfer learning, overcoming data scarcity by leveraging information from related datasets. |
| FT-Transformer & Vision Transformer (ViT) [2] | Neural Network Architecture | Advanced encoder architectures used within MatMCL for modeling tabular data and image data, respectively, to capture complex, non-linear relationships. |
| StepCCL [22] | Computational Library | A custom collective communication library designed to hide communication overhead within computation, optimizing large-scale distributed training. |
| FAIR Data Repositories [19] | Data Infrastructure | Standardized data repositories adhering to the Findable, Accessible, Interoperable, and Reusable (FAIR) principles, which are critical for addressing data heterogeneity. |
The intertwined challenges of data scarcity, heterogeneity, and multiscale complexity represent significant but surmountable barriers in materials informatics. As detailed in this whitepaper, the strategic application of transfer learning, robust data management infrastructures, and advanced multimodal learning frameworks like MatMCL and MMScale provides a clear pathway forward. Progress will depend on the continued development of modular AI systems, the widespread adoption of standardized FAIR data practices, and sustained cross-disciplinary collaboration. By addressing these core challenges, the materials science community can unlock transformative advances in the discovery and design of novel functional materials.
The convergence of high-throughput experimentation (HTE) and the FAIR (Findable, Accessible, Interoperable, and Reusable) data principles represents a fundamental shift in materials science and pharmaceutical research. This transformation is driven by the need to accelerate discovery while ensuring the growing volumes of complex, multimodal data remain valuable for future reuse. HTE enables the rapid exploration of vast chemical and materials spaces through automated, parallelized experimentation, dramatically increasing the pace of data generation. However, this acceleration creates a critical bottleneck in data management, where the value of experimental outputs depends entirely on how well they can be integrated, analyzed, and reused. The FAIR principles provide a framework to overcome this bottleneck by ensuring data are machine-actionable and semantically rich, enabling both human understanding and computational analysis. Within the context of multimodal data parsing for materials informatics, this integration is essential for building comprehensive processing-structure-property-performance relationships that drive innovation in functional materials and drug development.
High-Throughput Experimentation: Methodologies and Quantitative Impact
High-Throughput Experimentation employs automated, parallelized platforms to efficiently explore experimental parameter spaces, drastically reducing the time and resources required for discovery and optimization. In pharmaceutical development, HTE has become indispensable for candidate screening and reaction optimization. A 20-year implementation journey at AstraZeneca demonstrates the profound impact of systematized HTE, where automation of powder dosing using systems like CHRONECT XPR enabled the screening of diverse solids—including transition metal complexes, organic starting materials, and inorganic additives—with high precision: deviations of <10% at sub-milligram masses and <1% at masses >50 mg [23]. This technological advancement reduced manual weighing time from 5-10 minutes per vial to under 30 minutes for an entire experiment, while simultaneously eliminating significant human errors associated with manual handling at small scales [23].
The experimental workflow for pharmaceutical HTE typically involves several standardized protocols. Library Validation Experiments (LVEs) screen building block chemical space against variables like catalyst type and solvent choice in 96-well array manifolds at milligram scales [23]. Reaction screening employs inert atmosphere gloveboxes and robotic liquid handling systems with resealable gaskets to prevent solvent evaporation [23]. In oncology discovery, implementation of integrated HTE workflows at AstraZeneca facilities increased average quarterly screen sizes from ~20-30 to ~50-85, while the number of conditions evaluated grew from <500 to approximately 2000 per quarter over a seven-quarter period [23].
In materials chemistry, flow chemistry has emerged as a powerful HTE tool that addresses limitations of traditional batch-based screening. Flow HTE enables investigation of continuous variables (temperature, pressure, reaction time) dynamically throughout experiments, providing wider process windows and improved safety profiles for hazardous chemistry [24]. The methodology is particularly valuable in photochemical reaction optimization, where flow reactors enable efficient photochemical processes by minimizing light path length and precisely controlling irradiation time, overcoming challenges of poor light penetration and non-uniform irradiation in batch systems [24]. Automated platforms integrate flow chemistry with inline/real-time process analytical technologies (PAT), creating efficient HTE workflows requiring less material and human intervention [24].
Table 1: Quantitative Impact of HTE Implementation in Pharmaceutical Research
| Metric | Pre-Automation Performance | Post-Automation Performance | Implementation Context |
|---|---|---|---|
| Weighing Time | 5-10 minutes per vial | <30 minutes for complete experiment | Solid dosing with CHRONECT XPR [23] |
| Weighing Accuracy | Significant human error at small scales | <10% deviation (sub-mg to low mg)<1% deviation (>50 mg) | Wide range of solid materials [23] |
| Quarterly Screen Size | ~20-30 screens per quarter | ~50-85 screens per quarter | AstraZeneca Oncology Discovery [23] |
| Conditions Evaluated | <500 per quarter | ~2000 per quarter | AstraZeneca Oncology Discovery (7 quarters) [23] |
| Photochemical Scale-up | Laboratory scale (grams) | 6.56 kg per day throughput | Photoredox fluorodecarboxylation [24] |
Diagram 1: High-Throughput Experimentation Workflow
FAIR Data Principles: Implementation Frameworks for Materials Science
The FAIR principles provide a structured framework for scientific data management, emphasizing machine-actionability to handle the increasing volume, complexity, and creation speed of research data [25]. FAIR represents four foundational pillars: Findability (easy location of data and metadata through persistent identifiers and rich descriptions), Accessibility (retrieval using standardized protocols with authentication where necessary), Interoperability (integration with other data through common languages and vocabularies), and Reusability (comprehensive description with clear provenance and licensing) [25]. These principles address critical challenges in modern materials research, where multimodal datasets from combinatorial materials science are often too large and complex for human reasoning alone, distributed across institutions, and variable in format, size, and content [21].
Implementation of FAIR principles occurs through structured "FAIRification" processes. The NOMAD Laboratory platform exemplifies this approach through its schema-based Metainfo system, which preserves structural semantics of standardized data formats like NeXus and makes them interoperable across materials science domains [26]. Recent extensions to NeXus standards developed through FAIRmat collaboration include NXapm for atom probe microscopy, NXem for electron microscopy, and application definitions for various spectroscopy techniques, creating cross-domain standards for experimental materials data [26]. These standardization efforts follow transparent, community-driven processes where definitions are openly discussed and refined on GitHub before official adoption [26].
Best practices for FAIR implementation include incorporating FAIR considerations at project inception, utilizing domain-specific metadata standards, applying clear usage licenses, and engaging data stewards with specialized knowledge in data governance and lifecycle management [27]. The Datatractor framework addresses tool discoverability and inconsistent usage instructions that hinder FAIR implementation by providing a curated registry of data extraction tools with standardized, lightweight schema descriptions, enabling machine-actionable installation and use [28]. This approach addresses inefficiencies of tool reimplementation while offering both public-facing data extraction services and integration capabilities for research data management systems [28].
Table 2: FAIR Data Implementation Frameworks and Standards
| Framework/Standard | Primary Domain | Key Features | Implementation Examples |
|---|---|---|---|
| NOMAD Metainfo | Materials Science | Schema-based system for metadataPreserves structural semanticsEnables cross-platform interoperability | Oasis for community data sharingNeXus format integration [26] |
| NeXus Standard Extensions | Experimental Materials Science | NXapm (atom probe microscopy)NXem (electron microscopy)Optical spectroscopy definitions | Community-driven development via GitHubFull NOMAD platform integration [26] |
| Datatractor | Chemical & Materials Sciences | Curated registry of extraction toolsStandardized schema descriptionsMachine-actionable installation | Public data extraction servicesRDM system integration [28] |
| Perovskite JSON Schema | Hybrid Perovskite Materials | Standardized composition reportingIUPAC-compliant descriptionsMachine-readable representations | Hybrid Perovskite Ions DatabaseNOMAD API accessibility [26] |
Diagram 2: FAIR Data Principles Implementation Ecosystem
Multimodal Data Parsing: Bridging HTE and FAIR Data
Multimodal data parsing provides the critical technical bridge connecting high-throughput experimentation with FAIR data ecosystems, transforming heterogeneous experimental outputs into structured, interoperable information. In materials science, this involves integrating diverse data modalities including synthesis conditions, characterization results (e.g., XRD patterns), property measurements, and computational descriptors [21]. The parsing challenge is particularly acute for legacy building documentation in construction materials, where differences in design standards and drafting conventions across historical periods create diverse and complex representations that resist automated processing [7]. Similar challenges exist in materials informatics, where heterogeneous data formats and incomplete parameter information hinder the development of comprehensive materials databases.
Advanced parsing methodologies employ hybrid approaches combining rule-based and machine-learning techniques. For architectural and materials data, vector element parsing with layer semantic analysis enables structured extraction of key component geometry, while spatial topological relationship analysis improves modeling accuracy [7]. In text parsing, combining regular expressions, domain-specific terminology dictionaries, and BiLSTM-CRF deep learning models significantly improves extraction accuracy of unstructured parameters from scientific literature and experimental documentation [7]. For complex nested tables common in materials characterization data, multi-scale sliding windows with geometric feature analysis enable automatic detection and parameter extraction [7].
Experimental results demonstrate the effectiveness of these approaches. In architectural data parsing, F1 scores for wall line, wall, and column recognition reach 98.1%, 84.9%, and 92.2% respectively, while door and window recognition achieves 74.3% and 76.2% F1 scores [7]. For text parameter extraction, the PENet model achieves precision of 83.56% and recall of 86.91%, and table parameter extraction recalls for doors/windows and structure reach 95.0% and 96.7% respectively [7]. These parsing capabilities enable what is described as "Beyond 3D" multi-dimensional BIM integration, where drawings provide geometry and topology, text contributes materials and performance data, and tables supply identifiers and specifications [7].
Integrated Workflow: From Experimentation to Data-Driven Discovery
The complete integration of HTE, multimodal parsing, and FAIR data management creates a powerful ecosystem for accelerated discovery. This workflow begins with automated experimental execution, where platforms like the CHRONECT XPR system handle powder dosing of diverse solid materials with minimal deviation from target masses [23]. In parallel, liquid handling systems prepare reagent solutions in multi-well plates within inert atmosphere gloveboxes. The experimental phase employs either batch-based approaches in 96- or 384-well plates or continuous flow systems that enable investigation of continuous variables like temperature, pressure, and reaction time [24].
Following experimental execution, multimodal data parsing extracts and standardizes parameters from heterogeneous sources. For photochemical reactions, this includes parsing reaction conditions, light intensity parameters, conversion metrics, and spectral data [24]. The parsed data then undergoes FAIRification through platforms like NOMAD, where it is enriched with standardized metadata using domain-specific schemas, assigned persistent identifiers, and registered in searchable resources [26]. This process employs application definitions like NXoptical_spectroscopy for optical spectroscopy data or NXmpes for photoemission spectroscopy, ensuring semantic interoperability across experimental techniques [26].
The resulting FAIR data ecosystem enables advanced data mining and machine learning applications. For perovskite materials research, a standardized JSON schema following IUPAC recommendations enables both human- and machine-readable descriptions of over 300 identified perovskite ions, capturing descriptors including composition, molecular formula, SMILES representation, IUPAC name, and CAS number [26]. Similar approaches for metal-organic frameworks (MOFs), electrospun PVDF piezoelectrics, and 3D printed mechanical metamaterials facilitate the mapping of complex structure-property-processing relationships [19]. The curated Hybrid Perovskite Ions Database, accessible via the NOMAD API, demonstrates how standardized data enables researchers worldwide to upload, share, and reuse consistent materials data in line with FAIR principles [26].
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Essential Research Reagents and Platforms for HTE and FAIR Data Workflows
| Tool/Platform | Function | Application Context |
|---|---|---|
| CHRONECT XPR Workstation | Automated powder dispensing (1 mg to grams)Handles free-flowing to electrostatic powdersCompact footprint for glovebox integration | High-throughput screening of solid catalysts, reactants, and additives in pharmaceutical and materials synthesis [23] |
| NOMAD Laboratory Platform | FAIR data management, storage, and sharingSchema-based Metainfo systemNeXus standard integration | Materials science data repository enabling cross-platform interoperability and data reuse across computational and experimental domains [26] |
| Flow Photochemical Reactors | Enables photochemical HTE with controlled irradiationMinimized light path lengthPrecise residence time control | Photoredox reaction screening and optimization, including flavin-catalyzed fluorodecarboxylation and cross-electrophile coupling [24] |
| Datatractor Framework | Curated registry of data extraction toolsStandardized schema descriptionsMachine-actionable installation | Metadata extraction from scientific literature and experimental documentation for chemical and materials sciences [28] |
| BiLSTM-CRF Models | Named Entity Recognition (NER) for textual dataDomain-specific terminology integrationUnstructured parameter extraction | Parsing architectural texts, material specifications, and experimental protocols for multimodal data integration [7] |
| Multi-well Plate Reactors | Parallel reaction screening (96/384-well)Miniaturized reaction volumes (~300 μL)Integrated mixing and cooling | Initial reaction condition screening, catalyst evaluation, and solvent optimization in pharmaceutical and materials chemistry [24] |
The integration of high-throughput experimentation with FAIR data principles through advanced multimodal parsing represents a paradigm shift in materials and pharmaceutical research. This synergy addresses both the acceleration of discovery and the long-term value preservation of experimental data, creating a foundation for sustainable, data-driven scientific progress. The transformation is evidenced by quantitative improvements in pharmaceutical screening throughput, precision gains in experimental execution, and robust frameworks for data interoperability across research communities.
Future developments will focus on enhancing semantic interoperability, advancing autonomous experimentation systems, and refining hybrid parsing models that combine rule-based and machine-learning approaches. Initiatives like FAIR 2.0 aim to extend the FAIR guiding principles to address semantic interoperability challenges more comprehensively, ensuring data and metadata are not only accessible but also meaningful across different systems and contexts [27]. Similarly, the development of FAIR Digital Objects (FDOs) seeks to standardize data representation, facilitating seamless data exchange and reuse globally [27]. In computational materials science, hybrid models combining the strengths of traditional neural network potentials with foundation model concepts show promise for improving predictive accuracy and computational efficiency [28]. As these technologies mature, the research community moves closer to fully autonomous discovery systems where HTE, multimodal parsing, and FAIR data management create a continuous cycle of hypothesis generation, experimental validation, and knowledge extraction.
Multimodal Parsing Techniques and Real-World Material Applications
In the field of materials information research, the integration of heterogeneous data—from atomic-scale microscopy and spectral analysis to macroscopic mechanical properties and scientific literature—presents a significant computational challenge. Effectively parsing this multimodal data is crucial for accelerating the discovery and development of new materials and pharmaceuticals. Two competing artificial intelligence (AI) paradigms have emerged to address this complexity: the well-established modular pipeline and the increasingly prominent end-to-end vision-language model.
This whitepaper provides an in-depth technical comparison of these two approaches, framing them within the specific context of multimodal data parsing for materials science and drug development. It is structured to equip researchers and scientists with the knowledge to select and implement the optimal strategy for their specific research challenges, supported by quantitative data, detailed experimental protocols, and practical toolkits.
Defining the Approaches
Modular Pipelines
A modular pipeline decomposes a complex task, such as analyzing a material's structure-property relationship, into a sequence of discrete, specialized components or sub-tasks. Each component is designed, optimized, and validated independently, with the output of one module serving as the input for the next [29]. In a typical materials science workflow, this might involve a series of steps such as data ingestion, preprocessing, feature extraction, and predictive modeling.
End-to-End Models
An end-to-end model seeks to directly map raw, multimodal inputs (e.g., a scanning electron microscopy image and a textual description of processing parameters) to a desired output (e.g., a prediction of tensile strength) using a single, unified model, most often a deep neural network [29]. This approach minimizes human intervention in intermediate stages, relying on the model's internal architecture to learn optimal representations and sub-tasks from the data.
Comparative Analysis: Performance and Practical Considerations
The choice between modular and end-to-end approaches involves trade-offs across multiple dimensions, including performance, resource requirements, and operational flexibility. The following table synthesizes a comparative analysis based on recent implementations and research.
Table 1: Comparative analysis of modular pipeline and end-to-end approaches
| Aspect | Modular Pipelines | End-to-End Models |
|---|---|---|
| Performance Metrics | High reliability in controlled tasks (e.g., 92.3% success rate for template-based DAG generation) [30]. Excels in precision-focused tasks like variant calling in genomics [31]. | Often leads in overall accuracy on complex tasks (e.g., state-of-the-art precision/recall) [29]. Superior on integrated reasoning benchmarks like MatVQA [32]. |
| Data Requirements | Can be effective with smaller, well-defined datasets for individual components. | Data-intensive; requires large amounts of high-quality, multimodal data for training [29]. |
| Computational Cost | Inference can be efficient; total cost depends on pipeline complexity. | Training is computationally expensive and often intractable; inference can also be costly [29]. |
| Explainability & Debugging | High; failures are easily diagnosable to specific components, facilitating correction [29]. | "Black box" nature makes it difficult to locate the source of errors or understand decisions [29]. |
| Flexibility & Updating | Updating a single component is straightforward; however, output/input format changes can require downstream revisions [29]. | Highly flexible; can be retrained for new tasks with new data, often without architectural changes [29]. |
| Development Effort | High; requires significant design choices and expertise to define components and interactions [29]. | Lower initial effort; avoids thorny component design problems but requires deep learning expertise [29]. |
| Optimization | Suboptimal; components are optimized independently, errors accumulate, and downstream info cannot inform upstream components [29]. | Optimal for the global task; the entire model is jointly optimized, allowing all parts to co-adapt [29]. |
| Risk Mitigation | Easier to validate and control individual components, reducing risks of biased or incorrect output [29]. | Higher risk of biased, incorrect, or offensive output derived directly from training data [29]. |
Experimental Protocols for Materials Research
To ground this comparison in practical research, below are detailed methodologies for implementing each approach in a scenario involving the prediction of a material's properties from its processing conditions and microstructure.
Protocol 1: Modular Pipeline for Structure-Property Prediction
This protocol is inspired by established data management and bioinformatics principles [33] [31].
Objective: To predict the mechanical properties of an electrospun nanofiber material based on processing parameters and SEM microstructural images, using a modular, reproducible pipeline.
Workflow Overview: The following diagram illustrates the sequence of discrete, containerized modules in this pipeline.
Methodology Details:
Data Ingestion & Curation:
- Inputs: A dataset containing processing parameters (e.g., flow rate, voltage, concentration) and corresponding SEM images [2].
- Tools: Data is managed following FAIR principles (Findable, Accessible, Interoperable, Reusable) using platforms like Pymicro, which provides a high-level interface for building complex, multimodal datasets [33].
- Output: A standardized and version-controlled dataset.
Data Preprocessing:
- Tabular Data: Normalize and scale numerical processing parameters. Encode categorical variables.
- Image Data: Apply image normalization, resizing, and data augmentation techniques to SEM images to improve model robustness.
Feature Extraction:
- From Processing Parameters: Use a Multilayer Perceptron (MLP) or a FT-Transformer to convert tabular data into a feature vector [2].
- From SEM Images: Use a Convolutional Neural Network (CNN) or a Vision Transformer (ViT) pretrained on a relevant task to extract a feature vector representing microstructural characteristics [2].
Multimodal Fusion:
- Process: Concatenate or use cross-attention mechanisms to combine the feature vectors from the processing parameters and SEM images into a unified representation [2].
- Output: A single, fused embedding that captures the joint information from both modalities.
Property Prediction:
- Model: A final predictor module, such as a fully connected neural network, takes the fused embedding as input and outputs predictions for target properties (e.g., yield strength, elastic modulus) [2].
Validation:
- Method: Perform rigorous cross-validation. Benchmark the pipeline's performance using established metrics (e.g., Mean Absolute Error, R² score) on a held-out test set.
Protocol 2: End-to-End Multimodal Learning with MatMCL
This protocol is based on the MatMCL framework, designed to handle multimodal data even when some modalities are missing [2].
Objective: To train a single, unified model that can directly ingest processing parameters and SEM images to predict mechanical properties, and to demonstrate its capability for cross-modal tasks like generating structures from processing conditions.
Workflow Overview: The end-to-end process involves pre-training a model to understand the relationships between modalities before fine-tuning it for specific downstream tasks.
Methodology Details:
Model Architecture:
Structure-Guided Pre-training (SGPT):
- Objective: Align the representations of different modalities in a shared latent space without using property labels. This is a self-supervised step.
- Process: For a batch of data, the fused representation (from both processing and structure) is used as an anchor. The corresponding unimodal representations are positive samples, and all others are negatives. A contrastive loss (e.g., InfoNCE) is used to maximize agreement between positive pairs and minimize it for negatives [2].
- Outcome: The model learns a robust, shared embedding space where semantically similar materials are close, even if they have different missing modalities.
Downstream Task Fine-tuning:
- Property Prediction with Missing Data: The pre-trained encoders are frozen. A small predictor network is trained on top of the fused or unimodal embeddings. Crucially, due to SGPT, the model can make accurate predictions even when the SEM image modality is missing, by using only the processing parameters [2].
- Conditional Structure Generation: The model can be extended with a generative component (e.g., a decoder) to generate synthetic SEM microstructures based on a given set of processing parameters [2].
- Cross-Modal Retrieval: The aligned embedding space allows for querying with one modality (e.g., a SEM image) to retrieve materials with similar processing parameters, or vice-versa [2].
Validation:
- Method: Evaluate on benchmark datasets like MatVQA, which requires fine-grained visual and textual reasoning [32]. For property prediction, standard regression metrics apply. For generation, use Fréchet Inception Distance (FID) to assess image quality.
The following table details key software and data resources essential for implementing the aforementioned experimental protocols.
Table 2: Key resources for multimodal materials informatics
| Category | Tool / Resource | Function | Relevant Context |
|---|---|---|---|
| Workflow Orchestration | Snakemake [31] | A workflow management system to create reproducible and scalable data analyses. | Used in modular bioinformatics pipelines for WES and RNA-Seq analysis [31]. |
| Apache Airflow DAGs [30] | Programmatically author, schedule, and monitor workflows as Directed Acyclic Graphs. | Generated automatically from natural language via the Prompt2DAG methodology [30]. | |
| Containerization | Docker [31] | Containers package software and its dependencies into a standardized unit, ensuring consistency across environments. | Critical for deploying modular pipelines in cloud environments like the Google Cloud Platform [31]. |
| Multimodal Learning Frameworks | MatMCL [2] | A versatile multimodal learning framework for materials science that handles missing modalities and enables cross-modal tasks. | Core framework for the end-to-end protocol described in this paper [2]. |
| Benchmarks & Data | MatVQA [32] | A benchmark for evaluating research-level multimodal reasoning on materials science imagery and text. | Used to rigorously test the capabilities of end-to-end MLLMs on structure-property-performance reasoning [32]. |
| Data Management | Pymicro [33] | An open-source Python package offering a high-level interface to build complex multimodal datasets, complying with FAIR principles. | Used for managing 4D multimodal mechanics data for material microstructures [33]. |
| Core ML Libraries | PyTorch / TensorFlow | Foundational open-source libraries for building and training deep learning models, including transformers and CNNs. | Essential for implementing both modular components and end-to-end models. |
The comparison between modular pipelines and end-to-end models reveals a clear, context-dependent trade-off. Modular pipelines offer superior control, explainability, and reliability for well-defined, sequential tasks with high-stakes outcomes, such as clinical biomarker analysis [31]. Their structured nature makes them ideal for environments where reproducibility and regulatory compliance are paramount.
Conversely, end-to-end models excel in tackling complex, integrated reasoning tasks where optimal performance is the primary goal and the "black-box" nature is an acceptable trade-off [29]. Their ability to learn directly from raw, multimodal data and to generalize across tasks makes them exceptionally powerful for discovery-driven research, such as uncovering novel processing-structure-property relationships [2] [32].
For the materials and drug development researcher, the optimal path forward may not be a binary choice but a strategic hybrid. One could leverage modular pipelines for robust, standardized data preprocessing and validation, while integrating end-to-end models for specific, high-complexity prediction and generation tasks. As frameworks like MatMCL continue to evolve, they will further blur the lines between these paradigms, offering more flexible and powerful tools to drive the next generation of scientific breakthroughs.
The acceleration of materials discovery and development increasingly hinges on the ability to extract and utilize structured knowledge from a vast, heterogeneous corpus of scientific literature. This literature, often stored in legacy formats like PDFs, contains critical experimental data, synthesis protocols, and characterization results locked within complex layouts, tables, and figures. Document parsing has emerged as an essential technological solution, serving as the foundational step in converting unstructured and semi-structured documents into structured, machine-readable data suitable for computational analysis and AI-driven discovery platforms [34]. In the specific context of materials informatics, the paradigm of data-driven research is fundamentally constrained by the inaccessibility of historical knowledge; effective parsing directly addresses this bottleneck by transforming published findings into a computable format [35].
The core challenge in materials science literature is its multimodal nature. A typical research article combines dense textual descriptions with complex tables of properties, spectral data, microstructural images, and graphical representations of chemical structures. A effective parsing system must therefore not only recognize individual elements but also understand their contextual relationships—for instance, linking a micrograph to its corresponding caption and the discussion of its properties in the text. This process, generally categorized into layout analysis, content extraction, and relation integration, forms the essential pipeline for unlocking this knowledge [34]. Subsequent sections of this whitepaper will provide a detailed technical examination of each component, present experimental protocols and benchmarks relevant to materials science, and illustrate integrated systems that are currently advancing materials research.
Core Technical Components
A comprehensive document parsing system can be architected through two primary methodologies: the traditional modular pipeline and the emerging end-to-end approach leveraging large vision-language models. The following diagram illustrates the high-level workflow of the modular pipeline system, which remains a dominant and highly interpretable paradigm.
Layout Analysis
Layout Analysis serves as the critical first step in the document parsing pipeline, responsible for identifying and locating the structural elements of a document. Its primary function is to segment a document image into semantically distinct regions—such as text blocks, paragraphs, headings, images, tables, and mathematical expressions—and to determine their spatial coordinates and logical reading order [34]. The accuracy of this stage is paramount, as errors in layout understanding propagate through subsequent extraction and integration steps, compromising the entire parsing outcome.
The technological evolution of layout analysis mirrors advances in computer vision. Early systems relied on rule-based methods and statistical techniques to analyze simple document structures [34]. The field was transformed by the adoption of deep learning, particularly Convolutional Neural Networks (CNNs). Models adapted from object detection, such as Mask R-CNN, proved highly effective for detecting page objects like text blocks and tables [34]. More recently, Transformer-based methods have demonstrated superior capability in capturing global document context. Architectures like the Document Image Transformer (DiT) process document images as sequences of patches, enabling a more nuanced understanding of complex layouts that are commonplace in scientific publications [34]. For materials science documents, which often feature multi-column layouts with intricate arrangements of figures, tables, and equations, these advanced models are essential for robust performance.
Content Extraction
Following layout analysis, the Content Extraction phase processes the identified regions to convert their visual information into machine-readable digital content. This is a multimodal task that requires specialized techniques for different types of content, each presenting unique challenges, especially in the technical domain of materials science.
Text Extraction: This process primarily leverages Optical Character Recognition (OCR) technology to convert images of text into encoded characters. While a mature technology, OCR in materials science must accurately handle extensive domain-specific terminology, symbols, and units (e.g., "MPa", "kV", "at.%") [34]. Modern OCR systems are integrated with language models to improve accuracy for specialized vocabularies.
Table Data and Structure Extraction: Tables in materials science papers are repositories of critical numerical data, such as alloy compositions, mechanical properties, and processing parameters. Table recognition involves two sub-tasks: table structure recognition, which identifies the layout of cells and the relationships between rows and columns, and table content recognition, which extracts the textual and numerical data from each cell [34]. The output is typically structured into formats like HTML, LaTeX, or JSON to preserve the relational nature of the data.
Mathematical Expression Extraction: The quantification of material behavior is expressed through mathematical equations. Extracting these requires detecting mathematical symbols and understanding their two-dimensional spatial arrangements (e.g., superscripts, subscripts, fractions). The goal is to convert these detected regions into standardized formats like LaTeX or MathML, which can be interpreted by computational engines [34].
Chart and Figure Recognition: Materials science is a highly visual field, relying on micrographs, spectra, and phase diagrams. Chart recognition goes beyond simple image extraction; it aims to interpret the chart type (e.g., SEM, XRD, stress-strain curve) and extract the underlying data points and labels, converting visual information back into a structured data table or JSON format [34].
Relation Integration
The final component, Relation Integration, is the synthesizing step that reassembles the individually extracted content elements into a coherent, unified structure that faithfully represents the original document. This process ensures that the spatial and semantic relationships between elements are preserved [34]. Without effective relation integration, the output is merely a collection of disjointed data points, lacking the context necessary for knowledge extraction.
This stage relies on the spatial coordinates generated during layout analysis to establish the physical layout of the document. Rule-based systems or specialized reading order models are then applied to infer the logical flow of content, determining, for example, that a figure caption belongs to the image above it, or that a paragraph of text references a specific table [34]. In the context of materials science, this is crucial for linking a discussion of "the microstructure shown in Figure 3a" to the actual image and its corresponding data, or for associating a specific dataset in a table with the graph that visualizes it. The final output of this integrated pipeline is a structured document in a format like JSON, XML, or Markdown, which can be seamlessly ingested by databases, knowledge graphs, or AI models for further analysis [34] [36].
Experimental Protocols and Benchmarking
Validating the performance of document parsing components requires rigorous evaluation on standardized datasets and benchmarks. The following table summarizes key quantitative results from recent evaluations in the field, highlighting the performance of different approaches on specific tasks.
Table 1: Performance Benchmarks for Document Parsing Components
| System / Model | Domain / Dataset | Task | Key Metric & Result | Reference |
|---|---|---|---|---|
| MERMaid Pipeline | Chemical Reaction PDFs (3 domains) | End-to-end reaction data extraction | 87% End-to-End Accuracy | [37] |
| Docling Parser | Financial Reports (Tesla Q3) | Table Structure & Content Extraction | High-Fidelity Table Reconstruction (Qualitative) | [36] |
| MatQnA Benchmark | Materials Characterization (10 techniques) | Multi-modal MLLM Evaluation | ~90% Accuracy (Top MLLMs on Objective Questions) | [38] |
| CRESt System | Materials Science (Fuel Cell Catalysts) | Automated Experimentation & Analysis | 9.3x improvement in power density per dollar; 3,500 tests conducted | [39] |
Protocol: End-to-End Knowledge Graph Generation (MERMaid)
The MERMaid pipeline provides a exemplary protocol for extracting structured knowledge from scientific PDFs, with a focus on chemical reactions, a task directly analogous to materials synthesis information extraction [37].
- Image Segmentation: The PDF is first processed to identify and segment all graphical elements, which are the primary carriers of complex scientific information.
- VLM-Powered Parsing: Each segmented graphic is processed by a Vision-Language Model (VLM) to parse the visual content. The VLM is tasked with identifying relevant entities and relationships, such as reactants, products, and catalysts in a reaction scheme.
- Context Completion and Coreference Resolution: The system performs self-directed context completion by cross-referencing parsed data with the document's text to resolve abbreviations and link references (e.g., "see Figure 1").
- Graph Generation: The extracted and resolved entities and relations are assembled into a coherent knowledge graph, representing the final structured output.
This protocol's effectiveness is demonstrated by its 87% end-to-end accuracy across 100 articles from three distinct chemical domains, proving its robustness to layout and stylistic variability [37].
Protocol: Automated Materials Experimentation (CRESt)
The CRESt platform demonstrates a advanced application where document parsing integrates with robotic experimentation in a closed-loop system for materials discovery [39].
- Multimodal Knowledge Ingestion: The system ingests and processes diverse information sources, including scientific literature, personal experience, and input from colleagues, using multimodal models to create a knowledge base.
- Hypothesis and Experiment Design: Using active learning, specifically Bayesian optimization, the system designs new material recipes and experiments. This process is augmented by the ingested knowledge, which helps define a more efficient search space.
- Robotic Synthesis and Testing: A liquid-handling robot and a carbothermal shock system synthesize the proposed materials. An automated electrochemical workstation then tests their performance.
- Characterization and Feedback: The synthesized materials are characterized using automated electron microscopy. The results are fed back into the active learning model to refine the next round of experiments.
This protocol enabled the exploration of over 900 chemistries and the conduction of 3,500 tests, leading to the discovery of a multi-element fuel cell catalyst with a record power density [39].
Emerging Architectures: The Role of Large Vision-Language Models
While modular pipelines offer precision and interpretability, a significant shift is underway toward end-to-end approaches based on Large Vision-Language Models. These models, such as GPT-4V, Claude, and specialized variants like Nougat, can simultaneously process visual and textual data, potentially simplifying the complex multi-stage pipeline into a single, unified model [34] [40]. The following diagram contrasts the traditional modular approach with this emerging VLM-based paradigm.
VLMs possess emergent capabilities in visual reasoning and contextual understanding, allowing them, in principle, to perform layout analysis, content extraction, and relation integration in a single, integrated step. This is particularly promising for handling documents with novel or highly complex layouts where predefined rules might fail. Their ability to follow natural language instructions also makes them highly flexible for extracting different types of information without retraining the core model. However, challenges remain, including high computational costs, potential "hallucinations" where the model generates incorrect content, and difficulties in handling high-density text and complex tables with perfect accuracy [34]. The optimal architecture for many enterprise-grade applications, including in materials science, may therefore be a hybrid approach, leveraging the precision of specialized modular components for well-defined tasks like OCR, while utilizing VLMs for higher-level reasoning and integration.
The Scientist's Toolkit: Research Reagents for Digital Extraction
The following table details key software and data "reagents" essential for constructing a modern document parsing pipeline for materials science research.
Table 2: Essential Research Reagents for Document Parsing Pipelines
| Reagent / Tool | Type | Primary Function | Application in Materials Research |
|---|---|---|---|
| Docling | Open-Source Parser | Converts PDFs/DOCX into structured JSON/Markdown; layout-aware. | Extracting text, tables, and figures from technical datasheets and historical papers for data curation. [36] |
| axe-core | Accessibility Engine | Checks color contrast and other UI rules programmatically. | Ensuring parsed visualizations and web-based data dashboards are accessible to all researchers. [41] |
| MatQnA Dataset | Benchmark Dataset | Multi-modal benchmark for evaluating LLMs on materials characterization. | Testing and validating the performance of AI models on domain-specific interpretation tasks. [38] |
| Pathway | Streaming Framework | Enables real-time processing of live data streams. | Building a continuously updating knowledge base from streaming scientific publications or lab instrument data. [36] |
| Vision-Language Model (VLM) | AI Model (e.g., GPT-4V) | End-to-end document understanding and question-answering. | Rapidly querying a corpus of parsed documents to find synthesis methods or property data. [40] [37] |
The core technical components of document parsing—layout analysis, content extraction, and relation integration—collectively form a critical technological foundation for the future of data-driven materials science. As evidenced by systems like CRESt and MERMaid, the ability to automatically convert unstructured scientific literature into structured, actionable knowledge is no longer a theoretical concept but a practical tool that is already accelerating discovery and innovation [39] [37]. The ongoing integration of more powerful and sophisticated Large Vision-Language Models promises to further enhance the robustness and scope of these systems, enabling them to tackle an even wider array of document types and complex scientific reasoning tasks. For researchers and professionals in materials science and drug development, mastering and contributing to these technologies is paramount to unlocking the full potential of the vast digital knowledge resources at their disposal.
In the domain of materials information research, the characterization of complex material systems relies on heterogeneous data streams from multiple analytical techniques. Spectroscopic, chromatographic, imaging, and sensor modalities each provide partial views of material properties and behaviors. Multimodal machine learning addresses the fundamental challenge of integrating these disparate data sources to form a unified representation that captures complementary, redundant, and cooperative information [42]. The fusion of such multimodal data is particularly crucial in pharmaceutical development, where predicting material properties, stability, and bioavailability requires synthesizing information across structural, compositional, and behavioral measurements.
The core thesis of this technical guide posits that effective data fusion strategies must be strategically selected based on data characteristics, computational constraints, and research objectives specific to materials science applications. Within multimodal machine learning, three principal paradigms have emerged: early fusion, late fusion, and coordinated representation learning, each with distinct mechanistic properties and application domains [43] [44] [45]. This guide provides an in-depth technical examination of these fusion strategies, their experimental implementations, and their relevance to materials informatics.
Theoretical Foundations of Multimodal Fusion
Core Concepts and Taxonomy
Multimodal learning fundamentally addresses the representation learning challenge of processing and relating information from different signal types or modalities [45]. In materials research, modalities may include spectral data (IR, NMR, Raman), structural information (XRD, microscopy), compositional analysis (MS, chromatography), and physical property measurements. The heterogeneity of these data sources presents significant challenges for integration.
The togetherness of multimodal signals during processing defines the core taxonomy of fusion approaches [45]. In joint representation learning, multimodal inputs are combined and projected into a unified representation space, allowing the model to learn cross-modal relationships directly [46]. In coordinated representation learning, separate models process each modality, with their representations coordinated through constraint-based learning to enable cross-modal reasoning without shared representation space [45].
Mathematical Formulations
From a mathematical perspective, fusion strategies can be formalized within the generalized linear model framework [47]. For early fusion with K modalities, the combined feature set X = (x₁,...,xₘ) where ⋃ᵢ₌₁ᵏ Xᵢ = X, the model satisfies:
g_E(μ) = η_E = Σᵢ₌₁ᵐ wᵢxᵢ
where gE(·) is the link function in early fusion, ηE is the output, wᵢ is the weight coefficient (wᵢ ≠ 0), and the final prediction is gE⁻¹(ηE) [47].
For late fusion, separate models are trained for each modality:
g_Lₖ(μ) = η_Lₖ = Σⱼ₌₁ᵐₖ wⱼₖxⱼₖ, k=1,2,...,K, xⱼₖ ∈ X
with the final decision being:
output_L = f(g_L₁⁻¹(η_L₁), g_L₂⁻¹(η_L₂), ..., g_Lₖ⁻¹(η_Lₖ))
where f(·) is the fusion function that aggregates decisions from each modality [47].
Fusion Architecture Paradigms
Early Fusion (Feature-Level Fusion)
Early fusion, also known as feature-level fusion, integrates raw data or feature representations from multiple modalities at the input stage before model training [43] [44]. This approach involves concatenating or otherwise combining feature vectors from different modalities into a unified representation that serves as input to a single machine learning model.
The experimental workflow for early fusion typically involves:
- Feature extraction from each modality using modality-specific encoders
- Feature concatenation into a single composite vector
- Joint model training on the combined feature set
In materials research, early fusion might combine spectral features, morphological descriptors, and compositional data into a unified feature vector for predicting material properties [48]. The convolutional LSTM architecture described in [49] demonstrates this approach, where audio and visual inputs are fused in the initial network layer, resulting in improved robustness to noise in both modalities.
Late Fusion (Decision-Level Fusion)
Late fusion, or decision-level fusion, employs separate models for each modality and combines their predictions at the decision stage [43] [44]. Each modality is processed independently, with fusion occurring only after individual models have generated their outputs.
The experimental protocol for late fusion involves:
- Individual model training on separate modalities
- Prediction generation from each modality-specific model
- Decision aggregation using voting, averaging, or weighted summation
In pharmaceutical applications, late fusion might combine predictions from separate models trained on chemical structure data, bioavailability measurements, and stability test results [44]. This approach allows domain experts to develop optimized models for each data type while leveraging complementary information at the decision level.
Intermediate Fusion and Hybrid Approaches
Intermediate fusion represents a hybrid approach where modalities are integrated after some processing but before final decision-making [42]. This strategy balances the benefits of early interaction between modalities with the flexibility of late fusion. The recently proposed gradual fusion method processes modalities in a stepwise manner based on their interrelationships, fusing highly correlated modalities first [47].
In complex materials characterization, intermediate fusion might initially combine structural and compositional data, then integrate spectroscopic information, and finally incorporate temporal stability measurements in a hierarchical manner.
Comparative Analysis of Fusion Strategies
Performance Characteristics and Trade-offs
Table 1: Comparative Analysis of Fusion Strategies for Materials Data
| Feature | Early Fusion | Late Fusion | Coordinated Representation |
|---|---|---|---|
| Fusion Level | Feature level [43] | Decision level [43] | Representation level [45] |
| Inter-modality Interaction | Direct interaction during feature extraction [43] | Limited interaction; models work separately [43] | Representations are aligned through constraints [45] |
| Data Handling | Integrates raw data/features at input level [43] | Integrates predictions from independent models [43] | Learns separate representations coordinated through loss functions [46] |
| Robustness to Missing Modalities | Low [44] | High [44] | Moderate to high [42] |
| Computational Efficiency | Single training process [43] | Multiple models trained independently [43] | Multiple encoders with coordination mechanism [46] |
| Dimensionality Challenges | High-dimensional feature space [43] | Avoids high-dimensional issues [43] | Moderate dimensionality [42] |
| Materials Science Application Examples | Combining spectral and structural features for crystal phase identification [48] | Ensemble of property prediction models [44] | Cross-modal retrieval of materials with similar properties [45] |
Selection Criteria for Fusion Strategies
The optimal fusion strategy depends on multiple factors specific to the materials research context:
Modality relationships: Early fusion excels when modalities are closely related and contain complementary information at the feature level [43] [46]. Late fusion is preferable when modalities are distinct or heterogeneous [43].
Data availability and quality: Late fusion demonstrates greater robustness to missing modalities, a common challenge in experimental materials science [44]. Early fusion requires complete multimodal datasets for training.
Computational constraints: Early fusion employs a single model, reducing training complexity, while late fusion enables parallel development of modality-specific models [43].
Task requirements: For tasks requiring rich cross-modal interactions (e.g., relating spectral signatures to mechanical properties), early or intermediate fusion is preferable. For problems where modalities provide independent evidence (e.g., ensemble property prediction), late fusion is more effective [44].
Recent theoretical work has established that the performance dominance between early and late fusion can reverse at a critical sample size threshold, with early fusion generally performing better with large datasets and late fusion showing advantages with limited data [47].
Implementation Methodologies
Experimental Protocols for Fusion Strategies
Early Fusion Implementation
The protocol for implementing early fusion in materials characterization involves:
- Feature extraction: Use modality-specific encoders to extract relevant features (e.g., CNN for microstructural images, autoencoders for spectral data).
- Feature normalization: Apply appropriate scaling to ensure compatibility across modalities.
- Feature concatenation: Combine normalized feature vectors into a unified representation.
- Joint model training: Train a prediction model on the concatenated features using a single loss function.
The study by Barnum et al. [49] demonstrates this approach with convolutional LSTM networks that immediately fuse audio and visual inputs, resulting in enhanced noise robustness.
Late Fusion Implementation
The experimental protocol for late fusion includes:
- Individual model training: Develop optimized models for each modality using domain-specific architectures.
- Prediction generation: Obtain outputs from each modality-specific model.
- Decision aggregation: Combine predictions using methods such as:
- Weighted averaging based on model confidence
- Meta-learners that optimize combination weights
- Voting schemes for classification tasks
In biomedical data fusion, late fusion often outperforms early approaches when modalities have different statistical properties or sampling rates [42].
Coordinated Representation Learning
Implementation of coordinated representations involves:
- Separate encoder training: Train modality-specific encoders to generate embeddings.
- Coordination objective: Apply coordination constraints (e.g., similarity measures, canonical correlation analysis) to align representation spaces.
- Joint optimization: Balance modality-specific losses with coordination objectives.
This approach is particularly valuable for cross-modal retrieval tasks in materials databases, where users might search for compounds with similar properties using different query modalities [45].
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Tools for Multimodal Fusion in Materials Research
| Tool/Category | Function | Example Implementations |
|---|---|---|
| Modality-Specific Encoders | Extract meaningful features from raw data | CNN for images (ResNet, VGGNet) [48], Transformers for sequences (BERT) [48], Graph NNs for molecular structures [42] |
| Fusion Architectures | Combine information from multiple modalities | Early fusion (concatenation), Late fusion (voting, averaging), Attention mechanisms [50] |
| Alignment Techniques | Coordinate representation spaces across modalities | Contrastive loss [51], Canonical Correlation Analysis [48], Similarity constraints [45] |
| Multimodal Benchmarks | Evaluate fusion performance | Amazon Reviews [44], MIntRec [50], Materials Project datasets |
| Implementation Frameworks | Develop and test fusion models | PyTorch, TensorFlow, Hugging Face Transformers [49] |
Applications in Materials Information Research
Pharmaceutical Development Applications
In drug development, multimodal fusion strategies enable more accurate prediction of compound properties and behaviors:
Early fusion applications include integrating structural descriptor data with physicochemical properties to predict bioavailability, where direct feature-level interactions enhance model accuracy [42].
Late fusion approaches combine predictions from separate models trained on chemical structure, in vitro assay results, and pharmacokinetic data to estimate drug efficacy and toxicity [44].
Coordinated representation learning enables cross-modal retrieval between chemical structures and biological activity profiles, facilitating drug repurposing and similarity search in large compound databases [45].
Materials Characterization and Discovery
Multimodal fusion accelerates materials discovery and characterization through:
Property prediction from multimodal characterization data, where early fusion of spectral and structural features improves prediction accuracy for mechanical and thermal properties [48].
Quality control systems that employ late fusion to combine results from multiple inspection techniques (e.g., spectroscopic, imaging, thermal analysis) for comprehensive material assessment [44].
Accelerated materials screening using coordinated representations that enable efficient similarity search across compositional, structural, and property spaces [45].
The field of multimodal fusion continues to evolve with several promising research directions specifically relevant to materials informatics:
Knowledge-guided fusion incorporates domain expertise to inform fusion architecture design, such as prioritizing certain modality interactions based on known material relationships [50]. Cross-modal generation techniques can synthesize plausible material structures from property specifications or predict spectra from structural data [51]. Resource-efficient fusion addresses computational challenges associated with large-scale multimodal materials data through techniques like modular networks and transfer learning [42].
As materials research increasingly relies on multimodal characterization techniques, the strategic selection and implementation of data fusion strategies becomes crucial for extracting maximum insight from complex, heterogeneous datasets. The optimal fusion approach depends critically on the specific research objectives, data characteristics, and computational resources available, requiring researchers to carefully consider the trade-offs outlined in this technical guide.
Cross-modal alignment is a foundational technique in multimodal machine learning, designed to integrate and harmonize data from diverse sources such as images, text, and genomic sequences. The core challenge lies in the heterogeneous nature of these data modalities, which often reside in disparate feature spaces, making direct comparison and fusion ineffective [52] [53]. By mapping these different modalities into a shared latent space, cross-modal alignment enables machines to understand and leverage the complementary information each modality provides. Within materials science and biomedical research, this approach is revolutionizing the analysis of complex, multimodal datasets, from integrating microstructural images with simulation data for new material discovery to combining histopathological images with genomic profiles for enhanced cancer survival prediction [54] [53] [3].
This technical guide provides an in-depth examination of contrastive learning and optimization algorithms for cross-modal alignment. It is framed within the context of multimodal data parsing for materials information research, offering a detailed exploration of core methodologies, their applications, and practical experimental protocols.
Core Concepts and Technical Foundations
Contrastive Learning for Cross-Modal Alignment
Contrastive learning has emerged as a powerful self-supervised framework for aligning multimodal data. Its fundamental objective is to learn an embedding space where similar data points (positive pairs) are pulled closer together, while dissimilar ones (negative pairs) are pushed apart [53].
In a typical cross-modal setup, a positive pair consists of data points from different modalities that are semantically related, such as a whole-slide pathology image and its corresponding genomic profile from the same patient [53]. Negative pairs are formed by associating a data point from one modality with non-matching data points from the other modality. The model, typically comprising encoder networks for each modality, is trained to maximize the similarity for positive pairs and minimize it for negative pairs. A common loss function used for this purpose is the Normalized Temperature-scaled Cross Entropy (NT-Xent) loss [3].
The success of this approach hinges on the careful construction of positive and negative pairs and the choice of a distance metric, often cosine similarity, which measures the alignment in the shared latent space.
Optimization Algorithms and Advanced Fusion Mechanisms
While contrastive learning establishes the initial alignment, advanced optimization and fusion mechanisms are required to model the complex, fine-grained interactions between modalities.
- Cross-Modal Attention Mechanisms: These modules allow features from one modality to dynamically inform and refine the features of another. For instance, in the CPathomic framework for cancer survival prediction, a cross-attention module is used to enhance the intermodal sharing of information, thoroughly exploring the correlations between histopathological and genomic data [53]. This enables the model to focus on the most relevant aspects of one modality when processing the other.
- Gated Attention Fusion: This mechanism regulates the expressiveness of each modality's representation before integration. By learning a set of gates that control the flow of information from each modality, the model can dynamically decide how much to rely on each data source, leading to a more robust and balanced fused representation [53].
These algorithms work in concert with contrastive learning to not only align the modalities but also to enable a rich, interactive fusion that captures the complex, non-linear relationships inherent in multimodal scientific data.
Applications in Materials and Biomedical Research
The outlined methodologies have demonstrated significant impact in several scientific domains, particularly in materials science and biomedicine. The table below summarizes key applications and their outcomes.
Table 1: Key Applications of Cross-Modal Alignment in Scientific Research
| Application Domain | Integrated Modalities | Core Methodology | Reported Outcome |
|---|---|---|---|
| Materials Science [3] | Crystal structure, Density of States (DOS), Charge density, Textual descriptions | MultiMat framework (CLIP-inspired multimodal contrastive learning) | State-of-the-art property prediction; Novel material discovery via latent space similarity. |
| Cancer Survival Prediction [52] [53] | Histopathological images (WSIs), Genomic data | CPathomic (Cross-modal contrastive learning & gated attention) | Consistently outperformed alternative multimodal survival prediction methods on TCGA datasets. |
| Text-to-Image Person Re-Identification [55] | Person images, Text descriptions | Interactive Cross-modal Learning (ICL) with MLLMs | Remarkable performance improvement on benchmarks like CUHK-PEDES and RSTPReid. |
| Vision-Language Tracking [56] | Visual sequences, Textual instructions | Text Heatmap Mapping (THM) for spatial alignment | Improved robustness to semantic ambiguity and multi-instance interference on OTB99 and LaSOT. |
These applications underscore the versatility of cross-modal alignment. In materials science, the MultiMat framework enables the construction of a foundation model that can seamlessly relate a material's structure to its various properties, accelerating the discovery of materials with desired functionalities [3]. In biomedicine, the integration of pathology and genomics through models like CPathomic provides a more holistic view of a patient's disease, leading to more accurate prognostic assessments [53].
Experimental Protocols and Methodologies
Workflow for a Cross-Modal Alignment Experiment
The following diagram illustrates a generalized experimental workflow for implementing and evaluating a cross-modal alignment system, synthesizing common elements from the cited research.
Detailed Methodological Breakdown
Data Preparation and Feature Encoding
The initial phase involves curating and processing multimodal datasets. For materials science, this could involve obtaining crystal structures, density of states, and automated textual descriptions from databases like the Materials Project [3]. In cancer research, this entails collecting paired Whole Slide Images (WSIs) and genomic data from sources like The Cancer Genome Atlas (TCGA) [53].
- Image Encoding: For high-resolution WSIs, a common approach is to use a pre-trained CNN (e.g., ResNet) to extract features from smaller image patches, which are then aggregated using a Multiple Instance Learning (MIL) encoder to form a global image representation [53]. For materials, a state-of-the-art Graph Neural Network (GNN) like PotNet can be used to encode the crystal structure [3].
- Sequence/Vector Encoding: Genomic data, often represented as feature vectors, can be processed with fully connected neural networks. Textual descriptions are typically encoded using transformer-based models like BERT or a pre-trained language encoder from a model like CLIP [55] [3].
Alignment and Fusion Implementation
This is the core technical phase where modalities are aligned and integrated.
- Contrastive Learning Setup: The encoded features from each modality are projected into a shared latent space using a projection head. The model is then trained with a contrastive loss function. For example, the CPathomic model uses a cross-modal representational contrastive learning module to minimize intermodal differences [53]. The MultiMat framework employs a CLIP-inspired loss to align the latent spaces of its four material modalities [3].
- Attention-Based Fusion: Following alignment, a cross-modal attention module is often deployed. This allows features from one modality (e.g., genomics) to attend to and refine features from another (e.g., pathology), capturing intricate intermodal relationships [53]. A gated attention mechanism can subsequently be used to weight and combine these refined features into a unified representation for the final prediction task.
Evaluation Metrics and Validation
Rigorous evaluation is critical. The standard methodology involves benchmarking the proposed model against established baselines on relevant downstream tasks.
Table 2: Quantitative Results of CPathomic on TCGA Datasets
| Cancer Type (TCGA Dataset) | Concordance Index (C-Index) | Comparison to Baselines |
|---|---|---|
| Bladder Urothelial Carcinoma (BLCA) | Reported | Consistently outperformed existing multimodal survival prediction methods [53]. |
| Breast Invasive Carcinoma (BRCA) | Reported | Effectively bridged modality gaps, leading to more accurate predictions [53]. |
| Uterine Corpus Endometrial Carcinoma (UCEC) | Reported | Surpassed existing methodologies using pathology, genomics, or both [53]. |
For materials property prediction, common metrics include Mean Absolute Error (MAE) and accuracy. The predictive performance of the MultiMat framework was shown to achieve state-of-the-art results on these tasks [3]. Furthermore, the quality of the learned latent space can be validated by performing material discovery screenings, searching for stable materials with desired properties based on latent space similarity [3].
The Scientist's Toolkit: Research Reagent Solutions
The following table details key computational "reagents" and resources essential for building cross-modal alignment systems in a research context.
Table 3: Essential Research Reagents for Cross-Modal Alignment Experiments
| Item / Resource | Function / Description | Exemplar Use Case |
|---|---|---|
| Pre-trained Encoder Models (e.g., ResNet, BERT, GNNs) | Provides a strong foundation for feature extraction from raw data (images, text, graphs), reducing data needs and training time. | Encoding histopathology image patches [53] or textual descriptions of materials [3]. |
| Multimodal Large Language Models (MLLMs) | Serves as a source of external knowledge for data augmentation, query refinement, and interactive learning. | Generating fine-grained textual descriptions for person re-identification [55] or enriching material data [54]. |
| Contrastive Learning Framework (e.g., CLIP) | The algorithmic blueprint for aligning multiple modalities in a shared latent space without dense supervision. | Aligning crystal structure with density of states in the MultiMat framework [3]. |
| Cross-Modal & Gated Attention Modules | Neural network components that enable fine-grained, dynamic interaction and fusion between aligned modalities. | Facilitating interactive learning between pathology and genomic data in CPathomic [53]. |
| Public Datasets (e.g., TCGA, Materials Project) | Large-scale, curated sources of multimodal scientific data for training and benchmarking models. | Training and evaluating cancer survival models [53] and materials foundation models [3]. |
Architectural Diagram: The MultiMat Framework
The MultiMat framework provides an excellent example of a modern, scalable architecture for cross-modal alignment in science. Its design, which can handle an arbitrary number of modalities, is illustrated below.
The accelerated discovery and development of advanced materials are increasingly reliant on the intelligent integration of experimental data. This whitepaper presents a detailed technical examination of three pivotal areas in materials science—electrospun nanofibers, composite materials, and thermoelectric discovery—framed within the context of multimodal data parsing for materials informatics research. The convergence of high-throughput experimentation, advanced manufacturing techniques, and data-driven methodologies is fundamentally reshaping the materials design timeline [57]. However, transformative advances require addressing significant deficiencies in materials informatics, particularly the lack of standardized experimental data management for complex, multi-institutional datasets [57] [58].
This document provides researchers, scientists, and drug development professionals with both theoretical frameworks and practical experimental protocols. It emphasizes the critical importance of establishing processing-structure-property-performance (PSPP) relationships through comprehensive data integration across the entire materials lifecycle [57]. The case studies presented herein demonstrate how multimodal data management infrastructures can bridge the gap between traditional materials development and next-generation, data-accelerated discovery.
Electrospun Nanofibers: Fundamentals and Fabrication Control
Electrospinning is an electrohydrodynamic process that utilizes high-voltage electrostatic force to stretch a polymer solution into nanofibers with diameters typically ranging from several nanometers to micrometers [59]. This technology has gained significant traction due to its simple, inexpensive setup and ability to produce nanofibers with high specific surface area, high porosity, and excellent processability [59]. The origins of electrospinning date back to 1745 with electrostatic atomization principles, with significant refinement occurring after the 1990s through the work of Reneker's group at the University of Akron [59].
Electrospun polymer nanofibers (EPNFs) have become particularly valuable for biomedical applications including tissue engineering, drug delivery, wound dressings, and various sensor types [59] [60]. Their ability to mimic the extracellular matrix (ECM) and provide a cell-friendly environment makes them ideal for creating scaffolds in regenerative medicine [59]. Additionally, their high surface area-to-volume ratio enables efficient loading and delivery of therapeutic agents in drug delivery systems [60].
Critical Processing Parameters and Control Strategies
The formation and characteristics of electrospun nanofibers are influenced by a complex interplay of parameters that must be carefully controlled to achieve desired fiber morphology and properties. These parameters can be categorized into solution properties, process conditions, and environmental factors [59] [60].
Table 1: Key Parameters Influcing Electrospun Nanofiber Quality
| Parameter Category | Specific Factor | Impact on Fiber Morphology | Optimal Control Strategy |
|---|---|---|---|
| Solution Properties | Polymer Molecular Weight | Affects chain entanglement and solution viscosity; inappropriate MW causes defects or irregular diameters [59] | Use polymers with appropriate relative molecular weight to balance chain interactions and solution fluidity [59] |
| Solution Viscosity | Determines fiber diameter and continuity; too high prevents extrusion, too low causes droplet formation [59] | Maintain viscosity within polymer-specific optimal range (e.g., 1-20 wt% for various polymers) [59] | |
| Electrical Conductivity | Influences jet stability and fiber stretching; higher conductivity produces thinner fibers [61] | Add ionic salts or use conductive polymers to modulate conductivity [61] | |
| Process Conditions | Applied Voltage | Controls electrostatic force for jet initiation; affects fiber diameter and stability [59] | Optimize voltage to maintain stable Taylor cone without instabilities (typically 10-30 kV) [59] |
| Flow Rate | Determines solution supply rate; affects fiber diameter and potential bead formation [60] | Lower flow rates generally produce finer fibers; balance with voltage [60] | |
| Collector Distance | Influences solvent evaporation and fiber stretching; insufficient distance causes wet fibers [61] | Adjust distance (typically 10-20 cm) based on solvent volatility and electric field strength [61] | |
| Environmental Factors | Temperature | Affects solvent evaporation rate and solution viscosity [59] | Maintain constant temperature appropriate for polymer-solvent system [59] |
| Humidity | Influences solvent evaporation and fiber morphology; high humidity can cause porous structures [61] | Control humidity based on desired fiber morphology (typically 30-50%) [61] |
The relationship between solution viscosity and successful electrospinning has been systematically studied for various biomedical polymers, with optimal concentration ranges identified for different material systems [59]:
Table 2: Viscosity and Concentration Requirements for Common Electrospinning Polymers
| Polymer | Optimal Concentration Range | Viscosity Relationship | Typical Application |
|---|---|---|---|
| PLGA | Varies by molecular weight | Higher viscosity increases fiber diameter | Tissue engineering scaffolds [59] |
| PVA | 1-20 wt% | Lower viscosity produces finer fibers | Drug delivery, wound dressings [59] |
| PEO | Dependent on MW | Must exceed critical entanglement concentration | Template for composite fibers [59] |
| PLLA | Specific to solvent system | Optimal range prevents defects | Biomedical implants [59] |
Experimental Protocol: Standardized Electrospinning Procedure
Materials and Equipment:
- High-voltage power supply (5-30 kV capability)
- Syringe pump with precise flow control (0.1-5.0 mL/h)
- Stainless steel blunt-ended needles (18-22 gauge)
- Polymer solutions prepared with appropriate solvents
- Grounded collector (static or rotating)
- Environmental chamber (for humidity and temperature control)
Step-by-Step Methodology:
Polymer Solution Preparation:
- Dissolve polymer in appropriate solvent at room temperature with continuous stirring for 12-24 hours until complete dissolution.
- Filter solution through 0.45 μm filter to remove particulate matter.
- Measure viscosity, conductivity, and surface tension prior to electrospinning.
Electrospinning Setup Configuration:
- Fill syringe with polymer solution and secure in syringe pump.
- Connect needle to high-voltage power supply.
- Set collector distance based on solvent volatility (typically 10-20 cm).
- Adjust environmental controls to maintain constant temperature (20-25°C) and humidity (30-50%).
Parameter Optimization:
- Initiate with moderate voltage (12-15 kV) and flow rate (1.0 mL/h).
- Observe Taylor cone formation; adjust voltage to maintain stable cone without dripping.
- Collect preliminary fibers and analyze morphology by SEM.
- Systematically vary parameters using design of experiments (DOE) approach to optimize fiber diameter and morphology.
Fiber Collection and Characterization:
- Collect fibers on appropriate substrate (aluminum foil, glass slides, or specialized collectors).
- Analyze fiber morphology by scanning electron microscopy (SEM).
- Measure average fiber diameter and distribution from multiple SEM images (n>100 measurements).
- Assess porosity and pore size distribution using mercury porosimetry or image analysis.
Electrospinning Experimental Workflow
Composite Materials in Thermoelectric Applications
Thermoelectric Principles and Performance Metrics
Thermoelectric generators (TEGs) represent a solid-state energy conversion technology that transforms heat directly into electricity through the Seebeck effect, operating in a noiseless, environmentally friendly manner with minimal maintenance requirements [62]. The efficiency of thermoelectric materials is governed by the dimensionless figure of merit (ZT), defined as ZT = (S²σT)/κ, where S is the Seebeck coefficient, σ is electrical conductivity, T is absolute temperature, and κ is thermal conductivity [62].
Recent breakthroughs in thermoelectric materials have focused on developing compositionally complex systems that enable independent control of electronic and thermal transport properties. A notable advancement comes from researchers at Queensland University of Technology (QUT) and Fudan University, who developed a novel multinary alloy of silver, copper, tellurium, selenium, and sulfur, designated AgCu(Te,Se,S) [63]. This composite material achieves a ZT of approximately 0.83 at 343 K while maintaining exceptional mechanical flexibility—withstanding up to 10% strain—making it ideal for wearable applications [63].
Vacancy Engineering and Microstructural Control
The exceptional performance of advanced thermoelectric composites stems from strategic vacancy engineering, which involves deliberate manipulation of atomic vacancies in the crystal structure to fine-tune electrical transport properties and thermal conductivities [63]. In the AgCu(Te,Se,S) system, the incorporation of selenium and sulfur increased charge carrier concentration, reduced lattice thermal conductivity, and facilitated the formation of flexible Ag-S bonds [63].
This vacancy engineering approach enables the decoupling of typically interdependent electronic and thermal transport properties, allowing researchers to optimize electrical conductivity while minimizing thermal conductivity through enhanced phonon scattering at engineered interfaces and defects. The resulting materials demonstrate both high thermoelectric performance and mechanical durability necessary for practical applications in flexible electronics.
Device Integration and Performance Validation
The transition from material development to functional devices has been successfully demonstrated through the fabrication of thin-film thermoelectric generators incorporating the novel p-type AgCu(Te,Se,S) alloy with established n-type Bi₂Te₃ [63]. When mounted on a human arm, these devices generated approximately 126 µW/cm² under a temperature difference of 25 K, maintaining stable voltage output during bending and movement [63].
This successful integration validates the practical potential of these composite materials for wearable applications, including self-powered fitness trackers, health monitors, and other on-skin electronics that can harvest body heat without relying on conventional batteries. The simple and scalable synthesis of these composite films further enhances their suitability for both laboratory research and commercial development [63].
Multimodal Data Parsing for Materials Informatics
The Data Management Challenge in Combinatorial Materials Science
The convergence of high-performance computing, automation, and machine learning has significantly accelerated materials discovery, but transformative advances require addressing critical deficiencies in materials informatics, particularly the lack of standardized experimental data management [57]. This challenge is especially pronounced in combinatorial materials science, where automated experimental workflows generate datasets that are too large and complex for human reasoning [57] [58].
The multimodal and multi-institutional nature of modern materials research further compounds these challenges, with datasets distributed across multiple institutions in varying formats, sizes, and content types [57]. Establishing meaningful processing-structure-property-performance (PSPP) relationships requires comprehensive data integration across the entire materials lifecycle, from synthesis conditions through characterization results to property measurements [57].
Case Study: Multi-Institutional Data Management Dashboard
A representative case study in multimodal data management comes from the ThermoElectric Compositionally Complex Alloys (TECCA) project, a 4.5-year multi-institutional effort focused on thermoelectric materials discovery [57]. This project developed a specialized data dashboard to address the limitations of existing data management tools for collaborative and persistent data analysis and visualization [57].
The dashboard architecture features:
- A graphical user interface (GUI) built using the Svelte JavaScript framework
- A backend server utilizing Flask Python micro web framework
- REST API for programmatic data access
- Integration with Globus cloud file storage for secure, scalable data management
- Custom ingestion scripts to standardize diverse data formats and naming conventions
This infrastructure enables researchers to organize, search, filter, and visualize multimodal datasets—including synthesis parameters, X-ray diffraction patterns, and property measurements—without requiring local data downloads [57]. The implementation has successfully facilitated collaboration across institutional boundaries while maintaining data security for pre-publication research [57].
Multimodal Data Parsing Infrastructure
Bimodal Learning for Enhanced Property Prediction
Beyond data management, multimodal machine learning approaches show significant promise for enhancing materials property predictions. The Composition-Structure Bimodal Network (COSNet) represents a novel approach that leverages both composition and structural information to predict experimentally measured materials properties, even with incomplete structural data [64].
This bimodal learning framework has demonstrated significant error reduction across diverse materials properties including Li conductivity in solid electrolytes, band gap, refractive index, dielectric constant, energy, and magnetic moment, consistently outperforming composition-only learning methods [64]. The success of this approach hinges on strategic data augmentation based on modal availability, highlighting the importance of comprehensive data collection strategies in materials informatics.
Integrated Experimental Protocols
Thermoelectric Composite Synthesis and Testing Protocol
Research Reagent Solutions and Essential Materials:
Table 3: Key Reagents for Thermoelectric Material Development
| Material/Reagent | Specifications | Function in Research |
|---|---|---|
| High-Purity Elements | Ag, Cu, Te, Se, S (99.99+% purity) | Base constituents for multinary thermoelectric alloys [63] |
| Lab Crucibles | High-purity alumina or graphite | Containment during high-temperature synthesis [63] |
| Laboratory Furnaces | Programmable with controlled atmosphere | Melting and annealing during material preparation [63] |
| Bismuth Telluride (Bi₂Te₃) | n-type thermoelectric material | Counterpart for p-type alloys in device fabrication [63] |
| Characterization Tools | XRD, SEM, EDS capabilities | Structural, compositional, and morphological analysis [63] |
Step-by-Step Synthesis Methodology:
Precursor Preparation:
- Weigh high-purity elemental constituents (Ag, Cu, Te, Se, S) according to target stoichiometry in an inert atmosphere glovebox.
- For AgCu(Te,Se,S) system, typical ratios correspond to (AgCu)₀.₉₉₈Te₀.₈Se₀.₁S₀.₁ [63].
Alloy Synthesis:
- Load mixed precursors into high-purity crucibles and seal under inert gas or vacuum.
- Heat using programmable furnace with specific thermal profile:
- Ramp to 600°C at 5°C/min
- Hold at 600°C for 12 hours for homogenization
- Slow cooling to room temperature at 1-2°C/min
- For flexible composites, additional annealing at 300-400°C may be required to optimize microstructure.
Material Processing:
- Grind synthesized ingots to fine powders using mortar and pestle or mechanical milling.
- For thin-film devices, use hot pressing or spark plasma sintering to consolidate powders.
- Optimize processing conditions to achieve target density >95% of theoretical.
Characterization and Testing:
- Perform XRD analysis to confirm crystal structure and phase purity.
- Measure electrical conductivity and Seebeck coefficient using commercial systems (e.g., Ulvac ZEM-3).
- Determine thermal conductivity via laser flash analysis or comparative method.
- Calculate ZT values across temperature range (300-500K).
Multimodal Data Collection and Management Protocol
Essential Infrastructure Components:
- Web-based dashboard platform (Svelte/Flask stack)
- Globus cloud storage endpoint
- Standardized data templates for different measurement types
- Automated data ingestion pipelines
Standardized Data Collection Workflow:
Experimental Metadata Recording:
- Document synthesis conditions: precursors, temperatures, durations, atmospheres.
- Record processing parameters: milling times, pressing conditions, sintering profiles.
- Note environmental conditions: temperature, humidity, measurement dates.
Structural Characterization Data:
- Collect XRD patterns with standardized parameters (scan range, step size).
- Perform SEM imaging at multiple magnifications with consistent operating conditions.
- Conduct EDS analysis for compositional verification.
Property Measurement Data:
- Measure thermoelectric properties with complete calibration information.
- Perform mechanical testing with detailed protocol documentation.
- Conduct stability assessments under operational conditions.
Data Integration and Analysis:
- Upload raw data to designated Globus endpoint with standardized naming.
- Execute automated ingestion scripts to process and index new data.
- Validate data quality through dashboard visualization tools.
- Export standardized datasets for machine learning applications.
The case studies presented in this whitepaper demonstrate the powerful synergy between advanced materials systems and data-driven research methodologies. Electrospun nanofibers continue to offer exceptional versatility for biomedical applications, while novel composite materials like the AgCu(Te,Se,S) system are expanding the possibilities for flexible thermoelectric devices. However, maximizing the potential of these advanced materials requires robust infrastructures for multimodal data management that can accommodate the complexity and volume of modern combinatorial materials science.
The successful implementation of specialized data dashboards, as demonstrated in the TECCA project, provides a template for future materials informatics initiatives. By integrating comprehensive data management with bimodal machine learning approaches, researchers can accelerate the establishment of meaningful processing-structure-property-performance relationships across diverse materials systems. This integrated approach represents the future of materials discovery—one where sophisticated experimental techniques are enhanced by equally sophisticated data management and analysis capabilities to dramatically reduce development timelines and unlock new materials functionalities.
Overcoming Practical Challenges in Multimodal Material Data Integration
Addressing Missing Modalities and Incomplete Material Characterization Data
In materials science and drug development, research increasingly relies on multimodal data to characterize complex material systems. However, practical experimental constraints often result in missing or incomplete data modalities, creating significant analytical challenges. The ability to robustly handle these imperfections is crucial for advancing materials informatics and accelerating discovery pipelines. This technical guide examines the core challenges of missing modalities within multimodal data parsing frameworks and provides actionable methodologies for researchers to overcome these limitations while maintaining data integrity and analytical rigor.
The Problem of Missing Data in Materials Research
Fundamental Challenges
Incomplete data occurs routinely in materials characterization due to sensor limitations, sample preparation variability, equipment failures, or privacy constraints [65] [66]. In molecular epidemiology studies, for instance, a review found inconsistent disclosure of missing data and limited use of statistical methods specifically designed for incomplete data [66]. The consequences propagate through analysis, potentially biasing model predictions, reducing statistical power, and limiting generalizability of findings.
Taxonomy of Missing Data Mechanisms
Understanding why data is missing is essential for selecting appropriate handling strategies. Three primary classifications exist:
- Missing Completely at Random (MCAR): The probability of missingness is independent of both observed and unobserved data [67]. For example, a broken sensor randomly failing to record measurements.
- Missing at Random (MAR): Missingness depends on observed variables but not unobserved values [66] [67]. For instance, certain material properties are missing because researchers didn't measure them for specific material classes that are documented in other modalities.
- Missing Not at Random (MNAR): Missingness depends on the unobserved values themselves [67]. This occurs when, for example, difficult-to-synthesize materials lack certain characterization data precisely because of their challenging synthesis conditions.
Table 1: Classification of Missing Data Mechanisms
| Mechanism | Definition | Example in Materials Science | Ignorability |
|---|---|---|---|
| MCAR | Missingness independent of any data | Sensor failure during random intermittent periods | Ignorable |
| MAR | Missingness depends only on observed data | Certain tests not performed based on documented material class | Ignorable with appropriate methods |
| MNAR | Missingness depends on unobserved values | Difficult measurements skipped for problematic samples | Non-ignorable |
Methodological Frameworks for Handling Missing Modalities
Data Processing Approaches
Modality Imputation
Modality imputation operates at the raw data level, filling missing information by compositing or generating absent modalities from available ones [65]. The fundamental premise is that accurately imputed data enables downstream analysis as if complete modalities were available.
- Modality Composition: Techniques include using available modalities to mathematically construct reasonable approximations of missing ones, such as inferring structural characteristics from compositional data [65].
- Modality Generation: Advanced methods leverage generative models to create plausible missing modality data, often using generative adversarial networks (GANs) or variational autoencoders (VAEs) trained on complete multimodal datasets [65].
Representation-Focused Models
These approaches address missingness at the feature representation level rather than raw data:
- Coordinated Representation Methods: Apply specific constraints to align representations of different modalities in semantic space, enabling effective training even with missing modalities [65].
- Representation Generation: Generate missing modality representations directly from available data rather than imputing raw data [65].
- Representation Fusion: Combine representations from existing modalities to fill informational gaps [65].
Strategy Design Approaches
Architecture-Focused Models
These methods design flexible model architectures that dynamically adapt to available modalities during training and inference [65]. This includes modular neural networks that can process variable input combinations and still produce consistent output representations.
Model Combinations
Ensemble approaches strategically combine multiple specialized models, each handling different modality availability patterns [65]. These external model combinations provide robustness through diversity of architectural assumptions.
Advanced Algorithmic Approaches
Multiple Imputation Methods
Multiple imputation addresses uncertainty in missing values by creating multiple plausible datasets, analyzing them separately, then combining results [66] [67]. Techniques include:
- Multiple Imputation by Chained Equations (MICE): Iteratively imputes missing values using conditional distributions [67].
- Fully Conditional Specification (FCS): Imputes missing values one variable at a time, conditional on observed data [67].
Domain-Informed Approaches
Incorporating domain knowledge significantly improves missing data handling. In healthcare, for example, missing tests might indicate the test was medically unnecessary rather than truly absent [67]. Similarly, in materials science, understanding synthesis constraints can inform why certain characterizations are missing.
Experimental Protocols and Validation Frameworks
Data Extraction with Conversational LLMs
The ChatExtract methodology demonstrates a sophisticated approach to handling incomplete information in scientific literature [68]. This protocol enables accurate data extraction from research papers despite variability in reporting formats:
Workflow Stages:
- Initial Classification: A relevancy prompt identifies sentences containing target data, filtering out irrelevant text [68].
- Text Expansion: Creates a passage containing the paper title, preceding sentence, and target sentence to capture material context [68].
- Single vs. Multiple Value Processing: Implements separate extraction strategies based on data complexity [68].
- Uncertainty-Inducing Redundant Prompts: Follow-up questions encourage negative responses when appropriate, reducing hallucination [68].
- Structured Response Enforcement: Yes/No answer formats improve automated processing reliability [68].
Multimodal Table Understanding
For materials science specifically, tables contain approximately 85% of composition-property relationships [69]. Experimental protocols for handling missing tabular data include:
Input Modality Comparisons:
- GPT-4-Vision on Table Images: Direct image processing preserves visual layout cues but may miss textual nuances [69].
- GPT-4 on OCR Extraction: Converts table images to text but loses structural information [69].
- GPT-4 on Structured Table Data: Processes CSV formats maintaining table structure but dependent on extraction accuracy [69].
Table 2: Performance of Different Input Modalities for Table Extraction
| Input Modality | Accuracy Composition Extraction | F₁ Score Property Extraction | Key Advantages | Key Limitations |
|---|---|---|---|---|
| GPT-4-Vision (Image) | 0.910 | 0.863 (flexible) 0.419 (exact) | Preserves visual layout and spatial relationships | Dependent on image quality and resolution |
| GPT-4 + OCR (Text) | Not reported | Not reported | Extracts raw text content | Loses table structure and relationships |
| GPT-4 + Structured (CSV) | Not reported | Not reported | Maintains tabular structure | Dependent on accurate table parsing |
Implementation and Tooling
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools for Handling Missing Materials Data
| Tool/Category | Primary Function | Application Context | Key Features |
|---|---|---|---|
| Multiple Imputation by Chained Equations (MICE) | Statistical imputation | Handling missing values in multivariate data | Creates multiple plausible datasets, accounts for uncertainty |
| Conversational LLMs (GPT-4, ChatGPT) | Data extraction from literature | Processing incomplete or variably reported research findings | Zero-shot learning, contextual understanding, conversational refinement |
| Scikit-learn Imputation Modules | Machine learning preprocessing | Preparing incomplete datasets for modeling | SimpleImputer, KNN imputation, integration with ML pipelines |
| Scientific Data Visualization (CDD Vault) | Data exploration and visualization | Identifying patterns in incomplete materials data | Interactive graphing, filtering, side-by-side visualization [70] |
| ColorBrewer & Scientific Color Maps | Accessible visualization | Communicating results from incomplete data analysis | Color-blind friendly palettes, perceptual uniformity [71] |
Evaluation and Sensitivity Analysis
Robust evaluation is essential when working with missing data. Recommended practices include:
- Sensitivity Analysis: Testing how results vary under different imputation methods or missingness assumptions [67].
- Performance Metrics: Assessing model performance, bias, and variation introduced by missing data handling techniques [67].
- Domain Validation: Corroborating findings with domain expertise to identify implausible patterns arising from data imperfections [67].
Addressing missing modalities in material characterization requires a multifaceted approach combining statistical rigor, domain knowledge, and advanced computational techniques. As materials research increasingly relies on heterogeneous multimodal data, robust methodologies for handling incompleteness will become ever more critical. The frameworks presented here provide researchers with principled approaches to maintain analytical integrity while leveraging partially available information. Future directions include developing more sophisticated cross-modal generative models, creating standardized benchmarks for evaluating missing data handling techniques specific to materials science, and establishing reporting standards for documenting data incompleteness in materials research publications.
The paradigm of scientific discovery is increasingly driven by data-intensive research, particularly in fields like materials science and drug development. A significant challenge in this landscape is managing data heterogeneity from multi-institutional sources and formats. Modern research often requires integrating diverse data modalities—including structured numerical data, semi-structured operational logs, unstructured textual documentation, spectral data, and microscopic images—from collaborating institutions that utilize different instrumentation, protocols, and metadata standards [72]. This heterogeneity creates critical bottlenecks in knowledge extraction, data reproducibility, and AI model development. Effectively addressing these challenges requires sophisticated frameworks for data fusion, standardization, and interpretation that can transform fragmented data into coherent, machine-actionable knowledge. The emergence of multimodal artificial intelligence and advanced data mining techniques now offers promising pathways to overcome these historical barriers, enabling researchers to unlock the full potential of distributed scientific data.
Understanding Data Heterogeneity in Scientific Research
Data heterogeneity in multi-institutional research manifests across several interconnected dimensions, each presenting distinct challenges for integration and analysis. The primary dimensions of heterogeneity include:
Format Variability: Scientific data exists in diverse formats ranging from structured numerical measurements and semi-structured operational logs to unstructured textual documentation, images, and spectral data [72]. This variability is compounded by the prevalence of legacy formats like PDFs, which lack semantic structure despite being a primary medium for disseminating scientific findings [37].
Modality Differences: Research data encompasses multiple modalities including textual descriptions, molecular structures, spectral signatures, microscopic images, and experimental parameters. Each modality requires specialized processing approaches while maintaining contextual relationships between them [38].
Protocol Disparities: Different institutions employ varying experimental protocols, instrumentation, acquisition parameters, and sampling rates, leading to fundamental incompatibilities in data structure and quality [73]. This includes differences in calibration standards, measurement precision, and environmental conditions.
Metadata Inconsistencies: The absence of standardized metadata schemas across institutions results in incompatible annotation practices, terminology variations, and incomplete contextual information, hindering effective data curation and discovery [73].
Impact on Research and Development
The failure to adequately address data heterogeneity has profound implications for scientific progress and technological development, particularly affecting the reliability and generalizability of research findings. Key impacts include:
Reproducibility Challenges: Heterogeneous data sources and methodologies contribute significantly to the reproducibility crisis in scientific research, as experimental conditions cannot be adequately replicated or validated across institutional boundaries [39].
Analytical Limitations: Traditional statistical methods and rule-based systems struggle to capture complex, nonlinear relationships inherent in multi-source heterogeneous data, particularly when dealing with high-dimensional datasets and temporal dependencies [72].
AI Model Biases: Machine learning models trained on homogeneous datasets from single institutions often exhibit poor generalization performance when applied to data from other sources, limiting their real-world applicability and clinical utility [73].
Technical Frameworks for Heterogeneous Data Fusion
Transformer-Based Architectures for Multimodal Integration
The Transformer architecture has emerged as a powerful framework for addressing data heterogeneity challenges, particularly through its self-attention mechanism that enables capturing long-range dependencies and complex interactions between different data modalities [72]. Unlike traditional approaches that require extensive manual feature engineering, Transformer-based models can process heterogeneous data types through unified embedding representations, accommodating variable-length sequences and diverse data structures without sequential processing constraints.
The core mathematical formulation of the self-attention mechanism begins with transforming input representations into three distinct vector spaces—queries (Q), keys (K), and values (V):
where X represents the input sequence matrix, and W^Q, W^K, W^V are learnable parameter matrices. The attention weights are computed through scaled dot-product operations [72]:
This fundamental mechanism enables the model to dynamically weigh the importance of different data elements and modalities based on the specific context and task requirements. For material science applications, domain-specific adaptations of this architecture have demonstrated remarkable effectiveness in integrating diverse data streams including spectral signatures, microscopic images, and textual documentation [72] [38].
Specialized Data Fusion Methodologies
Several specialized methodologies have been developed to address the unique challenges of scientific data heterogeneity, each offering distinct advantages for particular research contexts:
Multi-scale Attention Mechanisms: Advanced implementations incorporate domain-specific multi-scale attention that explicitly models temporal hierarchies inherent in scientific processes, addressing the challenge of processing data streams with vastly different sampling frequencies—from millisecond sensor readings to monthly progress reports [72].
Cross-Modal Alignment Frameworks: Innovative contrastive learning approaches enable automatic discovery of semantic correspondences between heterogeneous modalities without requiring manually crafted feature mappings. These frameworks learn relationships between numerical sensor data, textual documentation, and categorical project states through self-supervised alignment [72].
Adaptive Weight Allocation: Dynamic algorithms that adjust data source contributions based on real-time quality assessment and task-specific relevance address the practical challenge of varying data reliability in experimental environments. This approach continuously evaluates data quality metrics and reweights source influence accordingly [72].
Multi-Instance Learning (MIL): For applications with annotation disparities, such as medical imaging, MIL frameworks enable learning from whole-image or breast-level labels without needing detailed region-of-interest annotations, effectively addressing scalability limitations across institutions with different annotation protocols [73].
Experimental Protocols and Implementation
End-to-End Data Processing Workflows
Implementing effective heterogeneous data fusion requires structured workflows that transform raw, multi-source data into coherent, analyzable knowledge representations. The following Graphviz diagram illustrates a comprehensive pipeline for managing data heterogeneity:
Quantitative Performance Benchmarks
The effectiveness of heterogeneous data fusion approaches must be rigorously evaluated against standardized metrics and benchmarks. The following table summarizes performance outcomes across different domains and methodologies:
Table 1: Performance Benchmarks for Heterogeneous Data Fusion Methods
| Application Domain | Methodology | Key Performance Metrics | Results | Reference |
|---|---|---|---|---|
| Chemical Engineering Construction | Improved Transformer with Multi-scale Attention | Prediction Accuracy, Improvement over Conventional Methods | >91% Accuracy, 19.4% Improvement over ML, 6.1% over Standard Transformer | [72] |
| Scientific PDF Mining (MERMaid) | Vision-Language Models for Reaction Extraction | End-to-End Accuracy Across Chemical Domains | 87% Accuracy Across 3 Chemical Domains | [37] |
| Materials Characterization (MatQnA) | Multimodal LLMs for Interpretation | Accuracy on Objective Questions | ~90% Accuracy for Advanced Models (GPT-4.1, Claude 4, Gemini 2.5) | [38] |
| Multi-institutional Mammography | Federated Learning, Multi-instance Learning | AUC, Generalization to Unseen Domains | Strong Performance with Marginal Drops vs Centralized Training | [73] |
| Fuel Cell Catalyst Discovery (CRESt) | Multimodal AI with Robotic Experimentation | Power Density Improvement, Cost Reduction | 9.3x Power Density per Dollar, 75% Precious Metal Reduction | [39] |
Research Reagent Solutions
Implementing heterogeneous data fusion requires both computational frameworks and specialized tools. The following table details essential "research reagents" for managing data heterogeneity:
Table 2: Essential Research Reagent Solutions for Data Heterogeneity Management
| Tool/Category | Primary Function | Application Context | Implementation Example |
|---|---|---|---|
| Vision-Language Models (VLMs) | Extract and interpret information from visual data and associated text | Mining scientific literature, interpreting spectral data | MERMaid pipeline for converting PDF graphics to knowledge graphs [37] |
| Multimodal Data Fusion Platforms | Integrate diverse data types (text, images, spectra) into unified representations | Materials discovery, chemical engineering projects | CRESt platform combining literature insights, chemical data, and experimental results [39] |
| Cross-modal Alignment Modules | Establish semantic relationships between different data modalities | Connecting spectral signatures with material properties | Contrastive learning frameworks for aligning numerical sensor data with textual documentation [72] |
| Federated Learning Frameworks | Enable collaborative model training without data sharing | Multi-institutional medical imaging studies | Privacy-preserving mammography analysis across healthcare institutions [73] |
| Benchmark Datasets | Standardized evaluation of model performance across diverse tasks | Materials characterization, educational assessment | MatQnA dataset with 10 characterization methods and 2,800+ question-answer pairs [38] |
Implementation Considerations and Best Practices
Workflow Architecture for Multimodal Data Parsing
Successful implementation of heterogeneous data management requires carefully structured workflows that address the unique characteristics of scientific data. The following Graphviz diagram details the component architecture for multimodal data parsing:
Addressing Implementation Challenges
Real-world deployment of heterogeneous data fusion systems must overcome several practical challenges that can impact system performance and reliability:
Reproducibility Assurance: Experimental workflows must incorporate comprehensive monitoring and validation mechanisms to address reproducibility challenges. The CRESt platform, for example, utilizes computer vision and vision-language models to monitor experiments, detect issues, and suggest corrections in real-time, significantly improving experimental consistency [39].
Annotation Harmonization: Multi-institutional collaborations must establish common annotation guidelines and quality standards to address disparities in labeling practices. When complete harmonization isn't feasible, weakly supervised approaches like multi-instance learning can leverage institution-specific annotations while maintaining model performance [73].
Computational Efficiency: Processing high-dimensional heterogeneous data requires optimized computational approaches, particularly for large-scale datasets. Context clustering and prompt tuning methods have demonstrated significant efficiency improvements while preserving analytical capabilities [73].
Domain Shift Mitigation: Even with extensive data aggregation, models may exhibit performance degradation on data from previously unseen institutions. Continuous evaluation on held-out "unseen" domains and implementation of domain generalization techniques are essential for maintaining robust performance across diverse institutional contexts [73].
Managing data heterogeneity across multi-institutional sources and formats represents both a critical challenge and significant opportunity for advancing materials research and drug development. The integration of Transformer-based architectures, multimodal learning approaches, and specialized data fusion methodologies has demonstrated substantial progress in transforming fragmented, heterogeneous data into coherent, actionable knowledge. As these technologies continue to evolve, several emerging trends promise to further enhance our capabilities: the development of increasingly sophisticated vision-language models for scientific data interpretation, the expansion of federated learning frameworks for privacy-preserving multi-institutional collaboration, and the creation of comprehensive benchmark datasets for standardized evaluation across diverse domains. By systematically addressing the technical, methodological, and implementation challenges outlined in this guide, researchers can unlock the full potential of heterogeneous scientific data, accelerating discovery and innovation across materials science and pharmaceutical development.
Optimizing Computational Efficiency and Handling Large-Scale Data Lakes
In the field of materials informatics, the ability to efficiently manage and parse large-scale multimodal data has become a critical enabler for scientific discovery. The development of new materials—from advanced metal-organic frameworks to novel piezoelectric polymers—increasingly relies on artificial intelligence (AI) models trained on diverse datasets spanning computational simulations, experimental characterization, and scientific literature. These data combine chemical compositions, processing parameters, microstructural images, spectral characteristics, and property measurements into complex information ecosystems. However, this data richness presents significant computational challenges: without optimized architectures, data lakes can transform from valuable resources into costly, inefficient "data swamps" that hinder rather than accelerate research. This technical guide examines best practices for structuring large-scale data lakes to balance computational efficiency with analytical flexibility, specifically within the context of multimodal data parsing for materials information research. By implementing strategic approaches to data organization, format selection, and multimodal integration, researchers can create foundational data infrastructures that support advanced AI-driven materials discovery while controlling computational costs.
Core Principles of Data Lake Optimization
Multi-Zone Data Architecture
A well-designed data lake employs a multi-zone architecture that segregates data based on its processing state and intended use. This approach balances flexibility, performance, and governance while optimizing both storage costs and query efficiency.
Raw Zone (Bronze): This layer contains immutable, original data in its native format (e.g., raw log files, JSON records from instruments, unprocessed computational outputs). Accessed sparingly for reprocessing or audit purposes, this zone serves as a system of record. Data here should be kept in cost-efficient storage tiers using compression to minimize expenses [74].
Curated Zone (Silver/Gold): This layer holds cleansed, transformed data ready for analytics and model training. Here, data is partitioned, consolidated into larger files, and converted to query-efficient columnar formats. By structuring this zone for fast reads, researchers ensure most analytical queries and AI training pipelines access optimized data rather than raw files [74].
Sandbox or Aggregated Zone: Many research organizations create aggregated or feature-engineered datasets (e.g., daily rollups, machine learning feature sets, pre-computed descriptors) in this area. These smaller, derived datasets enable rapid prototyping and analysis while offloading computational work from repeatedly scanning full-detail data [74].
This multi-zone approach provides an effective balance between cost and performance: raw data is retained for completeness on cheap storage, while refined data is duplicated and optimized for speedy access. Netflix's data "lakehouse" architecture built on Amazon S3 and Apache Iceberg exemplifies this approach, managing exabytes of data through logical zoning and robust metadata management to maintain performance at scale [74].
Strategic Partitioning for Query Efficiency
Partitioning is among the most effective techniques for improving data lake performance and reducing computational costs. By dividing datasets into subdirectories based on meaningful keys, query engines can prune irrelevant partitions at runtime, reading only the data slices needed for analysis.
Best Practices for Partitioning:
Select Appropriate Partition Keys: Time-based partitions (year/month/day) work exceptionally well for experimental or computational data with temporal dimensions. For materials research, alternative partitioning by material class, synthesis method, or characterization technique may better align with common query patterns [74].
Avoid Excessive Granularity: While finer partitions reduce data scanned per query, over-partitioning creates numerous small files that degrade performance. Target partition sizes that yield files of at least hundreds of megabytes each [75].
Leverage Metadata Catalogs: Using metastore services (e.g., Hive Metastore, AWS Glue Data Catalog) allows query engines to identify relevant partitions without exhaustive storage listing. This significantly accelerates query planning, especially in data lakes containing millions of files [74].
The performance impact of proper partitioning can be dramatic. AWS analysis demonstrated that date-partitioning a large dataset reduced query data scanning from 102.9 GB to 6.49 GB—a 94% reduction in scan volume. This translated to a cost reduction from $0.10 to $0.006 per query and runtime improvement from 4 minutes 20 seconds to just 11 seconds [74].
Optimal File Formats and Compression
The selection of appropriate file formats fundamentally impacts both storage efficiency and computational performance in data lakes. The transformation from raw, text-based formats to optimized binary representations can yield order-of-magnitude improvements in query performance.
Table 1: Performance Characteristics of Data Storage Formats
| Format | Storage Type | Best Use Cases | Compression Efficiency | Query Performance |
|---|---|---|---|---|
| CSV/JSON | Row-based | Data landing, exchange | Low (5-20% size reduction) | Poor (full scans required) |
| Apache Avro | Row-based | Streaming ingestion, write-heavy workloads | Medium (60-70% size reduction) | Good for full-record reads |
| Apache Parquet | Columnar | Analytical queries, ML training | High (75-80% size reduction) | Excellent (column pruning) |
| Apache ORC | Columnar | Analytical queries, data warehousing | High (75-80% size reduction) | Excellent (column pruning) |
Columnar formats like Parquet and ORC provide distinct advantages for analytical workloads in materials informatics: they store data by columns rather than rows, enabling query engines to read only the specific columns needed for analysis (projection pushdown). Additionally, they embed metadata and statistics (min/max values per block) that facilitate skipping unnecessary data ranges within columns [75] [74].
The performance differential between formats can be substantial. A comparison using the GDELT dataset showed a Parquet table scanning only approximately 1.0 GB and completing in 12.6 seconds, while the same data in CSV format required scanning 102 GB and took over 4 minutes—representing a 99% cost reduction and 95% time savings achieved solely through format optimization [74].
Multimodal Data Integration Framework
Materials informatics increasingly relies on multimodal data—combining composition data, processing parameters, microstructural images, and property measurements—to build comprehensive structure-property-performance relationships. The MatMCL framework demonstrates an effective approach to managing such heterogeneous data through structure-guided multimodal learning [2].
This framework employs specialized encoders for different data modalities: table encoders for processing parameters, vision encoders for microstructural images, and multimodal encoders that integrate diverse information streams into unified material representations. Through contrastive pre-training, the model aligns these modalities in a shared latent space, enabling robust property prediction even when certain modalities (e.g., expensive characterization data) are missing—a common scenario in materials research [2].
Table 2: Encoder Architectures for Multimodal Materials Data
| Data Modality | Encoder Type | Extracted Features | Implementation Examples |
|---|---|---|---|
| Processing Parameters | MLP or FT-Transformer | Nonlinear effects of synthesis conditions | MatMCL table encoder [2] |
| Microstructural Images | CNN or Vision Transformer (ViT) | Fiber alignment, diameter distribution, porosity | MatMCL vision encoder [2] |
| Compositional Data | Descriptor-based | Element properties, stoichiometric features | AlphaMat component descriptors [76] |
| Textual Literature | Language Models | Synthesis protocols, property relationships | MERMaid VLM pipeline [37] |
For knowledge extraction from legacy literature, vision-language models (VLMs) offer powerful capabilities. The MERMaid system demonstrates how multimodal AI can transform graphical elements from PDF documents into machine-actionable knowledge graphs, achieving 87% end-to-end accuracy across three chemical domains despite variability in layout and presentation styles [37].
Experimental Protocols & Workflows
High-Throughput Materials Discovery Pipeline
The CRESt (Copilot for Real-world Experimental Scientists) platform exemplifies an optimized workflow for multimodal materials discovery. This system integrates robotic equipment for high-throughput synthesis and testing with AI-driven experimental planning, creating a closed-loop discovery pipeline [39].
Experimental Protocol:
Literature Knowledge Embedding: Before physical experimentation, CRESt generates initial material representations by searching scientific papers for descriptions of elements or precursor molecules that might be useful. This creates a knowledge-informed prior for guiding experimentation [39].
Search Space Reduction: Principal component analysis is performed in the knowledge embedding space to identify a reduced search space capturing most performance variability. This addresses the "curse of dimensionality" inherent in multielement material systems [39].
Bayesian Optimization with Multimodal Feedback: The system employs Bayesian optimization in the reduced space to design experiments, incorporating information from literature, human feedback, and previous experimental results to guide the search for promising materials [39].
Robotic Synthesis and Characterization: A liquid-handling robot and carbothermal shock system enable rapid synthesis of candidate materials, followed by automated characterization through electron microscopy, X-ray diffraction, and electrochemical testing [39].
Computer Vision Monitoring: Cameras and visual language models monitor experiments, detecting issues and suggesting corrections to maintain reproducibility—a critical concern in materials synthesis [39].
In one application, this pipeline explored over 900 chemistries and conducted 3,500 electrochemical tests over three months, discovering a catalyst material that delivered a 9.3-fold improvement in power density per dollar compared to pure palladium [39].
Multimodal Learning for Property Prediction
The MatMCL framework provides a structured protocol for leveraging multimodal data to predict material properties, particularly valuable when certain data modalities are expensive or difficult to obtain.
Experimental Protocol:
Multimodal Dataset Construction: For electrospun nanofibers, processing parameters (flow rate, concentration, voltage, rotation speed, ambient conditions) are systematically varied. Resulting microstructures are characterized using scanning electron microscopy (SEM), and mechanical properties are measured through tensile testing in multiple directions [2].
Structure-Guided Pre-training (SGPT):
- Processing conditions are encoded using a table encoder (MLP or FT-Transformer) to model nonlinear synthesis effects
- Microstructural images are processed through a vision encoder (CNN or ViT) to extract morphological features
- A multimodal encoder integrates both information streams into fused material representations
- Contrastive learning aligns unimodal and multimodal embeddings in a joint latent space [2]
Property Prediction with Missing Modalities: After pre-training, encoders are frozen and a trainable multi-task predictor is added. The model can predict mechanical properties using only processing parameters—bypassing the need for structural characterization—by leveraging the cross-modal understanding developed during pre-training [2].
Conditional Generation and Retrieval: The framework supports inverse design through conditional structure generation (producing microstructures from processing parameters) and cross-modal retrieval (finding materials with similar structures or properties) [2].
This approach demonstrates how multimodal learning can overcome data scarcity in materials science by transferring knowledge across correlated data modalities, enabling accurate property prediction even with incomplete characterization data.
The Scientist's Toolkit: Essential Research Reagents & Platforms
Table 3: Essential Platforms and Tools for Multimodal Materials Informatics
| Tool/Platform | Type | Primary Function | Application in Materials Research |
|---|---|---|---|
| CRESt [39] | AI-Driven Experimental Platform | Robotic synthesis combined with multimodal AI guidance | Closed-loop discovery of functional materials (e.g., fuel cell catalysts) |
| AlphaMat [76] | Material Informatics Platform | End-to-end AI modeling from data preprocessing to prediction | Prediction of 12+ material properties using component and structural descriptors |
| MatMCL [2] | Multimodal Learning Framework | Integration of processing parameters and microstructural images | Property prediction with missing modalities; inverse materials design |
| MERMaid [37] | Vision-Language Pipeline | Extraction of chemical knowledge from PDF literature | Construction of reaction knowledge graphs from diverse publication formats |
| Apache Parquet [75] [74] | Columnar Storage Format | Efficient analytical querying of large datasets | Optimized storage for material property databases and characterization data |
| Delta Lake/Apache Iceberg [74] | Table Format Management | ACID transactions and versioning for data lakes | Reproducible analysis of experimental results with time travel capabilities |
| Matminer [76] | Feature Generation Toolkit | Calculation of material descriptors for machine learning | Feature engineering for composition-property relationship modeling |
Optimizing computational efficiency in large-scale data lakes represents a foundational requirement for advancing multimodal materials informatics. Through strategic implementation of multi-zone architectures, intelligent partitioning schemes, and columnar storage formats, research organizations can achieve order-of-magnitude improvements in both performance and cost-effectiveness. These data management foundations enable increasingly sophisticated AI approaches—from the multimodal learning frameworks like MatMCL that handle incomplete characterization data to systems like CRESt that close the loop between computational prediction and experimental validation. As materials research continues its transition toward data-driven paradigms, the principles outlined in this guide will prove essential for harnessing the full potential of multimodal data to accelerate the discovery and development of next-generation materials.
Ensuring Robustness Against Noise and Variable Data Quality
The pursuit of new materials, such as advanced catalysts for fuel cells or novel pharmaceutical compounds, increasingly relies on the integration of heterogeneous data. Modern materials information research synthesizes insights from experimental results, scientific literature, microstructural images, chemical compositions, and computational simulations [39]. This multimodal approach mirrors the collaborative, integrative nature of human scientists but introduces a significant challenge: ensuring analytical robustness against the pervasive issues of data noise and variable data quality. In real-world settings, the quality of different modalities can vary dramatically due to sensor errors, environmental interference, irreproducible experimental conditions, or missing data streams [77]. Failure to account for these imperfections can lead to biased models, irreproducible results, and ultimately, failed scientific conclusions. This guide provides a technical framework for materials and drug development researchers to build robust multimodal data parsing systems capable of withstanding the challenges of low-quality data, thereby accelerating the discovery and validation of new materials and therapeutics.
A Taxonomy of Data Quality Challenges in Multimodal Research
Real-world multimodal data is frequently imperfect. A systematic understanding of these imperfections is the first step toward building robust analytical systems. The primary challenges can be categorized as follows [77]:
- Noisy Multimodal Data: Data contaminated with heterogeneous noise from sensor errors, transmission losses, or environmental interference.
- Incomplete Multimodal Data: Scenarios where certain modalities are entirely missing for some data samples, a common occurrence in clinical and materials research.
- Imbalanced Multimodal Data: Significant discrepancies in the predictive quality or inherent properties of different modalities, which can cause models to overly rely on a single, predominant data stream.
- Quality-Varying Multimodal Data: The quality of a given modality dynamically changes across different samples due to unforeseeable environmental factors or sensor issues.
The following table summarizes these challenges and their potential impacts on research outcomes.
Table 1: Core Challenges in Low-Quality Multimodal Data and Their Research Impacts
| Challenge Type | Description | Common Causes | Potential Impact on Research |
|---|---|---|---|
| Noisy Data [77] | Data contaminated with heterogeneous noise. | Sensor errors, environmental interference, transmission losses. | Reduced model accuracy, misleading correlations, failed experimental validation. |
| Incomplete Data [77] | Some modalities are entirely missing for specific data samples. | Differing experimental protocols, patient drop-out, sensor failure. | Inability to use standard fusion models, biased population samples. |
| Imbalanced Data [77] [78] | Significant quality or property discrepancies between modalities. | Inherently different information content across sensors or techniques. | Models take "shortcuts," performing poorly on tasks requiring the weaker modality. |
| Quality-Varying Data [77] | Data quality dynamically changes per sample. | Changing environmental conditions (e.g., low-light for cameras). | Unreliable model performance that degrades outside controlled lab settings. |
Quantitative Frameworks for Data Quality Assessment
A rigorous, quantitative assessment of data quality is fundamental. This involves using statistical and computational techniques to summarize and characterize datasets, providing an evidence-based foundation for diagnosing issues and guiding remediation strategies [79].
Table 2: Key Quantitative Data Analysis Methods for Quality Assessment
| Analysis Category | Key Techniques | Application in Quality Assessment |
|---|---|---|
| Descriptive Statistics [79] | Measures of central tendency (mean, median, mode), dispersion (range, variance, standard deviation), and distribution shape (skewness, kurtosis). | Summarizes central tendency and spread of sensor readings; identifies potential outliers and unexpected data distributions. |
| Inferential Statistics [79] | Hypothesis testing (t-tests, ANOVA), regression analysis, correlation analysis. | Tests for significant differences in data quality between experimental batches; quantifies relationships between variables. |
| Gap Analysis [79] | Compares actual data against predefined quality targets or benchmarks. | Identifies specific dimensions where data fails to meet project requirements for completeness or accuracy. |
| Text Analysis [79] | Sentiment analysis, keyword extraction, language detection. | Extracts insights from unstructured data like lab notes or literature to identify inconsistencies or missing information. |
Experimental Protocols for Robust Multimodal Fusion
Protocol: The CRESt System for Materials Discovery
The Copilot for Real-world Experimental Scientists (CRESt) platform developed at MIT provides a robust, real-world protocol for handling multimodal data with integrated noise and quality variation [39].
- Objective: To autonomously discover new materials, such as high-performance fuel cell catalysts, by integrating and acting upon diverse, noisy data streams.
- Methodology:
- Multimodal Data Ingestion: The system incorporates diverse information sources, including insights from scientific literature, chemical compositions, microstructural images, and results from high-throughput robotic experiments.
- Knowledge-Embedded Active Learning: Unlike basic Bayesian optimization, CRESt creates "huge representations" of material recipes based on prior knowledge from text and databases. It performs principal component analysis in this knowledge-embedding space to define a reduced, more efficient search space.
- Robotic High-Throughput Experimentation: A symphony of robotic equipment, including liquid-handling robots, a carbothermal shock synthesizer, and an automated electrochemical workstation, executes experiments.
- Continuous Monitoring and Correction: Computer vision and vision-language models monitor experiments via cameras, detecting issues (e.g., sample misplacement) and suggesting corrective actions to human researchers, thereby improving reproducibility.
- Outcome: The system explored over 900 chemistries and conducted 3,500 tests, discovering an eight-element catalyst that achieved a 9.3-fold improvement in power density per dollar over pure palladium, demonstrating robustness against experimental noise and variability [39].
Protocol: Integrated Oversampling and Noise Reduction for Imbalanced Data
This protocol addresses the common issue of class imbalance, where critical events (e.g., a rare material property or adverse drug reaction) are infrequent [78].
- Objective: To create a balanced, high-quality dataset from severely imbalanced data for reliable predictive modeling.
- Methodology:
- Synthetic Data Generation: Apply the Random Over-Sampling Examples (ROSE) method to generate synthetic data points for the minority class based on a probability distribution, increasing data diversity.
- Noise and Overlap Reduction: Employ Tomek Link identification to find and eliminate data points from opposing classes that are very close to each other in the feature space. This removes ambiguous or noisy examples that impair classifier performance.
- Model Validation: Validate the cleaned and balanced dataset using a suite of machine learning and deep learning models (e.g., Support Vector Machine, Random Forest, Deep Neural Networks) to confirm performance improvement.
- Outcome: When applied to a severely imbalanced stroke dataset (98:2 ratio), this integrated strategy enabled more reliable and efficient predictive modeling for critical healthcare applications [78].
Visualizing Robust Multimodal Data Parsing Workflows
Workflow for Robust Multimodal Data Parsing
This diagram illustrates the core logical workflow for building a robust multimodal data parsing system, from data ingestion to validation.
Logic for Dynamic, Quality-Aware Fusion
This diagram details the decision logic for a dynamic fusion strategy that adapts to the varying quality of input data streams.
Building and executing robust multimodal data pipelines requires both physical and computational tools. The following table details key resources.
Table 3: Essential Research Reagent Solutions for Robust Multimodal Data Parsing
| Tool Category | Specific Tool / Resource | Function in Robust Data Parsing |
|---|---|---|
| Robotic Laboratory Equipment [39] | Liquid-handling robots, Carbothermal shock synthesis systems, Automated electrochemical workstations. | Enables high-throughput, reproducible synthesis and testing of materials, reducing human-introduced noise and variability. |
| Characterization Equipment [39] | Automated electron microscopy, Optical microscopy, X-ray diffraction. | Provides consistent, automated collection of microstructural and compositional data across many samples. |
| Computational & AI Resources [39] | Large Multimodal Models (LMMs), Computer Vision Models, Bayesian Optimization Software. | Integrates diverse data streams (text, images, data); suggests optimal experiments; monitors for irreproducibility. |
| Data Analysis & Visualization Software [79] | Python (Pandas, NumPy), R Programming, ChartExpo, SPSS. | Performs quantitative data analysis, statistical validation, and creates accessible visualizations to communicate data quality and results. |
| Data Management Platforms [80] | Custom SQLite databases with HTML5/CSS interfaces, platforms like KNIME. | Provides structured, FAIR-compliant storage for heterogeneous longitudinal data, ensuring findability and interoperability. |
Implementing Effective Data Standardization and Preprocessing Pipelines
The acceleration of materials discovery and development hinges on the ability to effectively translate raw, heterogeneous data into reliable, machine-learning-ready datasets. This is particularly critical for multimodal data parsing in materials informatics, where data from diverse sources—such as synthesis conditions, characterization results (e.g., X-ray diffraction), and property measurements—must be integrated. This whitepaper provides a comprehensive guide to building robust data standardization and preprocessing pipelines. We review foundational concepts, detail systematic methodologies for handling common data challenges, and present case studies from contemporary materials science research. Furthermore, we provide a curated toolkit of software and resources to empower researchers and scientists in drug development and related fields to enhance data quality, ensure reproducibility, and unlock the full potential of artificial intelligence and machine learning (AI/ML) in materials information research.
In the realm of materials informatics, the convergence of high-performance computing, automation, and machine learning has significantly altered the materials design timeline [19]. However, transformative advances in functional materials are gated by the deficiencies that currently exist in data management, particularly a lack of standardized experimental data management [21]. Modern materials engineering often involves combinatorial approaches where composition, phase, and microstructure are tuned to elucidate complex processing–structure–property–performance relationships. The datasets generated are not only large and complex but are also frequently multimodal and multi-institutional, distributed across various organizations with substantial variations in format, size, and content [21] [81].
Raw data, whether from automated synthesis robots, wearable sensors in clinical trials, or high-throughput characterization tools, is invariably messy. It is often plagued by noise, missing values, outliers, and structural inconsistencies [82] [83] [84]. The adage "garbage in, garbage out" is acutely relevant for AI/ML models, which are highly sensitive to the quality of input data. Without rigorous preprocessing, subsequent analysis can lead to uninterpretable models, a lack of generalizability, and erroneous conclusions [83]. Data preprocessing encompasses the essential steps of cleaning and refining raw data to ensure its reliability and suitability for analysis. For multimodal materials data, this involves a series of systematic procedures to transform disjointed data streams into a clean, structured, and interoperable format, thereby laying the foundation for accurate and predictive materials models [19] [83].
Foundational Concepts and Data Challenges
Defining the Pipeline Components
A data preprocessing pipeline is a sequential workflow that transforms raw data into a curated dataset. Key components include:
- Data Cleaning: The process of enhancing data reliability by handling artifacts like missing values, outliers, and inconsistencies [83]. This includes removing duplicate and irrelevant observations, and fixing structural errors like typos or unit inconsistencies [82].
- Data Transformation: Converting raw data into more informative formats suitable for analysis. This includes techniques such as data segmentation, feature extraction, and applying mathematical transformations [83].
- Data Normalization and Standardization: Scaling numerical data to a common range (normalization) or to have a mean of zero and a standard deviation of one (standardization). This improves the comparability of features and aids in the convergence of many machine learning algorithms [83].
- Data Integration: The process of combining data from different sources, resolving semantic and structural conflicts, and aligning time stamps to create a unified view [83] [81].
Common Data Imperfections in Materials Research
Materials data presents unique challenges that preprocessing must address:
- Multimodality: Data from synthesis protocols, spectral analysis (XRD), microscopy images, and property testing exist in different formats and scales, creating integration challenges [21] [81].
- Metadata Gaps: Inconsistent or missing metadata describing experimental conditions (e.g., temperature, pressure, solvent) severely limits data reusability [19].
- Small Datasets: Unlike domains with big data, experimental materials science often deals with small, costly-to-acquire datasets, necessitating specialized techniques to avoid overfitting [19].
- Non-FAIR Data: Data that does not adhere to the FAIR principles (Findable, Accessible, Interoperable, and Reusable) becomes isolated in silos, preventing effective integration and collaboration [81].
Methodologies for Data Preprocessing
This section outlines a systematic approach to preprocessing, complete with quantitative checks and standard protocols.
Data Cleaning and Imputation
The initial step involves diagnosing and remedying data quality issues. The following table summarizes standard methods for handling common problems.
Table 1: Common Data Cleaning Techniques and Methodologies
| Data Issue | Description | Recommended Handling Methods | Considerations for Materials Data |
|---|---|---|---|
| Missing Values | Absence of data points in a dataset. | - Imputation: Replace with mean, median, or mode [84].- Advanced Imputation: Use ML models (e.g., k-NN) to predict missing values [83].- Deletion: Remove features or instances with excessive missing data [82]. | The choice of imputation method should consider the physical plausibility of the imputed value. Deletion is only recommended when the missing data is extensive and random. |
| Outliers | Data points that deviate significantly from other observations. | - Identification: Use statistical methods (e.g., Z-scores, IQR) or visualization (box plots) [82] [84].- Analysis: Determine if the outlier is due to error or a genuine physical phenomenon [82].- Treatment: Remove if measurement error, otherwise retain and potentially create a separate model. | Outliers in materials data may represent a novel phase or a critical failure point; their removal should be rigorously justified based on domain knowledge. |
| Structural Errors | Inconsistencies in data entry and formatting. | - Standardization: Correct typos, variations in spelling, and ensure consistent units [82].- Structural Harmonization: Ensure categorical data (e.g., "CA", "California") is represented uniformly. | Critical for merging datasets from different institutions. Adopting community-wide semantic ontologies is the preferred long-term solution [19] [81]. |
| Noise | Random errors that obscure the underlying signal. | - Data Filtering: Apply smoothing filters (e.g., moving average, Savitzky-Golay) to time-series or signal data [83]. | Common in raw sensor data from in-situ characterization or wearable devices used in clinical trials for drug development [83]. |
Data Transformation, Normalization, and Standardization
Transformation and scaling are vital for preparing data for ML algorithms. The table below compares common techniques.
Table 2: Data Transformation and Scaling Methods
| Method | Formula (Example) | Use Case | Application in Materials Informatics |
|---|---|---|---|
| Normalization (Min-Max) | ( X{\text{norm}} = \frac{X - X{\text{min}}}{X{\text{max}} - X{\text{min}}} ) | Scales features to a range, often [0, 1]. Useful when data lacks a Gaussian distribution. | Scaling features like atomic radius or melting point to a common range for neural network input. |
| Standardization (Z-score) | ( X_{\text{std}} = \frac{X - \mu}{\sigma} ) | Centers data around a mean of 0 with a standard deviation of 1. Assumes a near-Gaussian distribution. | Preparing data for algorithms like SVM and k-means clustering that are sensitive to feature scales. |
| Segmentation | N/A | Dividing a continuous data stream (e.g., from a sensor) into meaningful chunks or windows for analysis [83]. | Segmenting a long-term degradation test of a battery material into cycles for feature extraction. |
| Feature Extraction | N/A | Deriving new, informative features from raw data (e.g., statistical features like mean, variance) [83]. | Extracting peak width and intensity from XRD patterns as features for a crystal structure classification model. |
Workflow for Multimodal Data Integration
Integrating multimodal data requires a structured workflow to ensure interoperability. The following diagram visualizes this process, which is critical for parsing materials information from disparate sources.
Case Studies and Experimental Protocols
Case Study 1: Combinatorial Materials Science Data
A 2024 study addressed the challenge of managing multimodal, multi-institutional datasets in combinatorial materials science [21]. The research involved data describing synthesis and processing conditions, X-ray diffraction patterns, and materials property measurements generated at several institutions.
Experimental Protocol for Data Management:
- Data Lake Ingestion: Raw data from partner institutions was aggregated into a central "data lake" without initial transformation.
- Dashboard Development: A low-barrier dashboard was developed to enable standardized organization, analysis, and visualization.
- Standardization: The dashboard enforced consistent formatting and metadata annotation for synthesis conditions, characterization results, and property measurements.
- Unified Access: The system provided a single interface to map the materials design space by linking processing, structure, and property data, which had previously been siloed.
This case study demonstrates that a focused effort on data infrastructure can overcome the challenges of multimodal data, facilitating data-driven materials discovery [21].
Case Study 2: Interoperability in Medical Imaging and Clinical Data
While from the biomedical domain, this 2025 study provides a directly applicable protocol for achieving interoperability between non-cooperating data resources, a common challenge in materials science [81]. The study connected the Medical Imaging and Data Resource Center (MIDRC) with clinical data repositories (N3C, BDC).
Experimental Protocol for Interoperability:
- Cohort Identification: Used the interoperability capabilities of the separate data repositories to identify matched patients (or materials samples) across them.
- Governance Navigation: Navigated the different data access and governance models of each repository (e.g., open data vs. controlled-access clinical data).
- Data Linking: Created unified cohorts containing both clinical/contextual data and imaging data for the same set of patients.
- Representativeness Assessment: Characterized the representativeness of the resulting multimodal cohort using the Jensen-Shannon Distance (JSD) metric to compare its demographics with broader population statistics. This step is crucial for assessing potential bias in the resulting AI/ML models [81].
This protocol underscores that technical interoperability must be coupled with collaboration between governance organizations to create high-value, multimodal datasets.
A successful preprocessing pipeline relies on both conceptual understanding and practical tools. The following table lists key software and resources.
Table 3: Essential Tools for Data Preprocessing and Analysis
| Tool Name | Type | Primary Function | Relevance to Materials Research |
|---|---|---|---|
| Python (with Pandas, Scikit-learn) | Programming Language | A versatile ecosystem for data manipulation, analysis, and machine learning [82] [84]. | The de facto standard for building custom data preprocessing and ML pipelines in materials informatics [19]. |
| OpenRefine | Desktop Application | A powerful tool for working with messy data: cleaning it, transforming it, and reconciling inconsistencies [82]. | Ideal for initial exploration and cleaning of tabular data from experiments before advanced analysis. |
| Git/GitHub | Version Control | A system for tracking changes in code and data, and for managing collaboration [82]. | Critical for maintaining reproducibility and managing versions of both preprocessing scripts and datasets. |
| R | Programming Language | A software environment for statistical computing and graphics [84]. | Widely used for statistical analysis and data visualization, particularly in academia. |
| Tableau / Power BI | Data Visualization | Tools for creating interactive dashboards and explanatory visualizations [82] [84]. | Useful for communicating data insights and creating exploratory dashboards for materials data [21]. |
| FAIR Principles | Guidelines | A set of principles (Findable, Accessible, Interoperable, Reusable) for scientific data management [81]. | A guiding framework for designing data management infrastructures from the outset, ensuring long-term value [19] [81]. |
The implementation of effective data standardization and preprocessing pipelines is not merely a preliminary technical step but a foundational component of modern materials informatics. As the field moves increasingly toward data-driven discovery and the integration of multimodal datasets, the rigor applied to data curation will directly dictate the success of AI/ML applications. By adopting the systematic methodologies outlined in this guide—from robust cleaning and transformation to the implementation of interoperability protocols and FAIR principles—researchers and drug development professionals can overcome the pervasive challenges of data quality. This will ultimately accelerate the design of novel materials, enhance the reproducibility of scientific findings, and pave the way for transformative advances in functional materials and beyond.
Benchmarking and Validating Multimodal Parsing Performance in Materials Research
Establishing Evaluation Metrics and Benchmark Datasets for Material Parsing
The advancement of materials science increasingly relies on sophisticated data analysis techniques to interpret complex multimodal characterization data. Material parsing—the process of automatically extracting meaningful information and relationships from materials data—has emerged as a critical capability for accelerating materials discovery and development. This technical guide examines the establishment of robust evaluation metrics and benchmark datasets specifically designed for material parsing tasks, framed within the broader context of multimodal data parsing for materials information research.
Recent breakthroughs in artificial intelligence, particularly large language models (LLMs) and multimodal large language models (MLLMs), have demonstrated remarkable potential for interpreting scientific data. In specialized domains like materials science, however, the capabilities of these models require systematic validation through domain-specific benchmarks [38]. The development of such benchmarks faces unique challenges due to the vast array of characterization techniques, specialized terminology, and the need to integrate across diverse data modalities including spectroscopic data, microscopic images, and structural information [38].
Foundational Concepts in Material Parsing
Material parsing encompasses multiple sub-tasks that require distinct evaluation approaches. These include:
- Structural Analysis: Interpreting data from techniques such as X-ray Diffraction (XRD) to determine crystal structure and phase composition
- Morphological Analysis: Extracting information from microscopy techniques including Scanning Electron Microscopy (SEM) and Transmission Electron Microscopy (TEM) regarding surface topography, particle size, and distribution
- Compositional Analysis: Parsing spectral data from techniques like X-ray Photoelectron Spectroscopy (XPS) and Fourier-Transform Infrared Spectroscopy (FTIR) to identify elemental composition and chemical bonding
- Thermal Analysis: Interpreting data from Differential Scanning Calorimetry (DSC) and Thermogravimetric Analysis (TGA) to understand material behavior under temperature variations
Each parsing task requires specialized evaluation metrics that account for domain-specific requirements and the multimodality of the source data.
Current Benchmark Datasets for Material Parsing
Established Material-Specific Benchmarks
Table 1: Existing Benchmark Datasets for Material Parsing
| Dataset Name | Focus Domain | Data Modalities | Size | Key Characteristics |
|---|---|---|---|---|
| MatQnA [38] | Materials Characterization | Images, spectra, text | 10 characterization methods | Covers XPS, XRD, SEM, TEM, etc.; multiple-choice and subjective questions |
| CDW-Seg [85] | Construction & Demolition Waste | High-resolution images | 5,413 annotated objects | Manual semantic segmentation; 10 material categories |
| BIMNet [86] | As-built BIM Reconstruction | Point clouds, BIM models | 116+ million points, 382 rooms | Geometric and topological evaluation metrics |
The MatQnA dataset represents the first multi-modal benchmark specifically designed for material characterization techniques, employing a hybrid approach combining LLMs with human-in-the-loop validation to construct high-quality question-answer pairs [38]. This dataset is organized according to material characterization techniques and includes a large collection of domain-specific textual resources such as journal articles and expert case studies.
The CDW-Seg dataset addresses segmentation of construction and demolition waste in cluttered environments, featuring high-resolution images captured at authentic construction sites with manual semantic segmentation annotations [85]. This dataset includes 5,413 manually annotated objects across ten material categories, representing a total of 2,492,021,189 pixels.
Cross-Domain Benchmarking Insights
Other domains offer valuable insights for material parsing benchmarks. The CEQuest dataset for construction estimation evaluates LLMs on construction drawing interpretation through 164 questions combining multiple-choice and true/false formats across five subject areas [87]. The AbilityLens benchmark for MLLMs evaluates six key perception abilities (counting, OCR, attribute recognition, entity extraction, grounding, and structural data understanding) using over 12,000 test samples compiled from 11 existing benchmarks [88].
Evaluation Metrics Framework
Metric Classification for Material Parsing
Table 2: Evaluation Metrics for Material Parsing Tasks
| Metric Category | Specific Metrics | Applicable Tasks | Strengths | Limitations |
|---|---|---|---|---|
| Geometric Metrics | Component-level shape accuracy, Position accuracy [86] | Segmentation, object detection | Quantifies physical alignment | May miss semantic accuracy |
| Topological Metrics | Graph-similarity based metrics [86] | Spatial relationship parsing | Evaluates connectivity relationships | Computationally intensive |
| Accuracy Metrics | Question answering accuracy, Rubric-based scoring [38] [89] | Visual question answering | Direct performance measurement | Requires comprehensive ground truth |
| Stability Metrics | Z-score variance across sub-metrics [88] | Model robustness assessment | Measures consistency across domains | Does not measure absolute performance |
Domain-Specific Metric Considerations
For material parsing, evaluation metrics must address both geometric accuracy (how well the parsed information aligns with physical measurements) and topological correctness (how accurately spatial relationships are captured) [86]. The BIMNet benchmark proposes component-level geometric metrics to assess shape and position accuracy of reconstructed models alongside graph-similarity based metrics to evaluate spatial connectivity accuracy [86].
The Gecko evaluation system for multimodal outputs provides a rubric-based approach that identifies semantic elements (entities, their attributes, and relationships) that need to be verified in generated content [89]. This approach generates verification questions for each semantic element and aggregates scores to produce a final evaluation metric.
Dataset Construction Methodologies
Data Collection and Annotation
High-quality benchmark construction involves meticulous data collection and annotation processes:
- Source Diversity: Collect data from multiple sources including peer-reviewed journal articles, expert cases, and real-world experimental data [38]
- Multi-modal Integration: Ensure unified representation of modality diversity (images, spectra, text) and semantic richness (descriptive, inferential, and judgmental layers) [38]
- Manual Annotation: Employ detailed manual annotation processes, such as the average 25 minutes per image required for semantic segmentation in the CDW-Seg dataset [85]
- Hybrid Validation: Combine model-assisted generation with manual verification to ensure accuracy and reliability [38]
Quality Assurance Framework
- Human-in-the-Loop Validation: Integrate domain expertise throughout dataset construction [38]
- Structured Partitioning: Divide datasets into training (75%), validation (15%), and testing (10%) subsets to ensure balanced distribution [85]
- Ethical Considerations: Address data privacy, consent, bias mitigation, and potential misuse [90]
- Comprehensive Documentation: Provide clear descriptions of data collection methods, curation processes, and preprocessing steps [90]
Experimental Protocols for Benchmarking
Multimodal Model Evaluation Protocol
The following workflow diagram illustrates a comprehensive experimental protocol for evaluating material parsing models:
Benchmarking Execution Guidelines
When executing material parsing benchmarks, researchers should:
- Establish Baseline Performance: Evaluate performance using established models (e.g., GPT-4.1, Claude Sonnet 4, Gemini 2.5) as reference points [38]
- Implement Multi-dimensional Assessment: Measure both accuracy and stability across diverse question types, domains, and metrics [88]
- Conduct Cross-modal Evaluation: Assess performance across different data modalities (visual, textual, spectral)
- Perform Statistical Validation: Ensure results are statistically significant through appropriate testing methodologies
Essential Research Reagents and Tools
Table 3: Research Reagent Solutions for Material Parsing Experiments
| Tool/Category | Specific Examples | Function in Material Parsing | Implementation Considerations |
|---|---|---|---|
| Multimodal LLMs | GPT-4.1, Claude Sonnet 4, Gemini 2.5, LLaVA [38] [87] | Core parsing engine for multimodal data | Model size vs. accuracy tradeoffs; domain adaptation requirements |
| Evaluation Frameworks | Gecko, AbilityLens, Custom rubric systems [89] [88] | Standardized assessment of parsing quality | Rubric design; question-answer pair generation |
| Annotation Tools | Labelme, Custom annotation platforms [85] | Ground truth creation for training and evaluation | Manual labor requirements; quality control mechanisms |
| Dataset Management | Hugging Face Datasets, Figshare, Custom repositories [38] [85] | Storage, versioning, and distribution of benchmark data | Accessibility; documentation completeness; maintenance plans |
| Visual Encoders | DINOv2, CLIP, SigLIP [88] | Visual feature extraction from material images | Domain adaptation; resolution requirements |
Advanced Implementation Considerations
Addressing Ability Conflicts in Multimodal Parsing
During MLLM training, researchers may observe ability conflicts where different perception abilities exhibit different improvement curves, with some abilities experiencing performance decline after further training [88]. Primary factors behind this phenomenon include:
- Data Mixing Ratio: The proportion of different data types in training datasets significantly impacts ability convergence
- LLM Model Size: Larger models may demonstrate different ability conflict patterns compared to smaller models
- Training Strategies: Fine-tuning and model merging methods can help resolve ability conflicts [88]
Stability-Enhanced Evaluation
Beyond traditional accuracy measurements, stability—achieving consistent performance across diverse factors such as domains, question types, and metrics—is crucial for robust material parsing systems [88]. Stability can be assessed by computing the variance of z-scores across sub-metrics, which reflects relative performance compared to all candidate models on each sub-metric.
Future Directions
The field of material parsing evaluation continues to evolve with several promising research directions:
- Automated Knowledge Component Extraction: Using LMMs to automatically extract knowledge components from educational and research content [91]
- Cross-Domain Benchmark Integration: Developing unified benchmarks that enable performance comparison across different scientific domains
- Dynamic Benchmark Platforms: Creating evolving benchmarks that adapt to new parsing challenges and material systems
- Explainability-Focused Metrics: Developing metrics that assess not just parsing accuracy but also the interpretability and explainability of results
As material parsing technologies advance, corresponding evaluation methodologies must evolve to adequately measure progress and identify areas requiring further development. The establishment of comprehensive evaluation metrics and benchmark datasets will play a crucial role in guiding research efforts and accelerating the adoption of AI-assisted materials research platforms.
Comparative Analysis of Multimodal AI Models on Characterization Tasks
In the field of materials science, the characterization of new compounds and structures generates a complex, multi-faceted stream of data, encompassing text, images, audio, and other modalities [92]. This information is often encoded within scientific documents, limiting the capability for large-scale analysis and discovery [54]. Multimodal Artificial Intelligence (AI) presents a transformative solution by creating unified systems capable of processing these diverse data types simultaneously [93]. For researchers and drug development professionals, leveraging these models is essential for accelerating the retrieval of material properties, the discovery of novel materials, and the synthesis of knowledge from disparate experimental and simulation data [54] [19]. This technical guide provides an in-depth analysis of leading multimodal AI models, evaluating their performance, technical architectures, and applicability to specific characterization tasks within materials informatics.
The Multimodal AI Landscape in Materials Research
Multimodal AI models are advanced vision-language models (VLMs) that process and understand multiple input types—such as text, images, and structured data—simultaneously. They utilize sophisticated deep learning architectures to analyze visual content alongside textual information, performing complex reasoning and understanding tasks [94]. The shift from traditional unimodal systems to multimodal AI marks a pivotal leap, enabling deeper contextual awareness by integrating various data types in parallel [93]. In materials science, this capability is being harnessed to build foundation models that align rich, complementary modalities such as crystal structures, density of states (DOS), charge density, and textual descriptions from sources like Robocrystallographer [3]. Frameworks like Multimodal Learning for Materials (MultiMat) demonstrate the potential of this approach by enabling self-supervised multi-modality training, achieving state-of-the-art performance in property prediction and novel material discovery [3].
Comparative Analysis of Leading Multimodal AI Models
The following analysis details the performance and specifications of top-tier multimodal AI models, with a focus on their applicability to technical and scientific characterization workloads.
Table 1: Performance and Specification Comparison of Leading Multimodal AI Models
| Model | Developer | Key Strengths | Context Window | Benchmark Performance | Primary Use Cases in Materials Science |
|---|---|---|---|---|---|
| GPT-4o [95] | OpenAI | Real-time audio, image, and text processing; 320ms response times. | 128K tokens | High accuracy on conversational tasks. | Real-time analysis; educational apps where students interact with visual and voice data. |
| Gemini 2.5 Pro [95] | Extremely large context window for massive datasets. | 2 million tokens | 92% accuracy on commercial benchmarks. | Legal document review; research synthesis across hundreds of papers; video content moderation. | |
| Claude Opus/Sonnet [95] | Anthropic | Optimized for accuracy and predictability; constitutional training for safety. | 200K tokens | 72.5% on SWE-bench (coding). | Document extraction (95%+ accuracy); financial report analysis; code review. |
| Grok 3 [95] | xAI | Integrates live data streams; DeepSearch mode for transparent reasoning. | Information Missing | 1400 ELO on technical problems. | Tracking real-time market sentiment; catching emerging trends in social data. |
| Llama 4 Maverick [95] | Meta | Open-source; mixture-of-experts architecture; complete data control. | Information Missing | Information Missing | Customizable vertical assistants; on-prem deployments for sensitive data. |
| Phi-4 Multimodal [95] | Microsoft | Designed for on-device processing; no cloud dependency. | 128K tokens | 6.14% word error rate for speech. | Defect detection on production lines; safety monitoring in remote locations. |
| GLM-4.5V [94] | Zhipu AI | State-of-the-art on 41 multimodal benchmarks; MoE architecture; 3D spatial reasoning. | Information Missing | SOTA on 41 public benchmarks. | Complex multimodal reasoning; analysis of images, videos, and long documents. |
| Qwen2.5-VL-32B-Instruct [94] | Qwen | Excels as a visual agent; can control computers; analyzes charts and layouts. | Information Missing | Information Missing | Automated data extraction from invoices and tables; document analysis. |
Table 2: Technical Specifications and Cost Analysis
| Model | Core Architecture | Input Cost (per million tokens) | Output Cost (per million tokens) | Data Fusion Approach |
|---|---|---|---|---|
| GPT-4o [95] | Transformer-based | $5 | Information Missing | Native multimodal processing |
| Gemini 2.5 Pro [95] | Transformer-based | Information Missing | Several dollars for full-context requests | Information Missing |
| Claude Opus/Sonnet [95] | Transformer-based | Information Missing | Information Missing | Information Missing |
| GLM-4.5V [94] | Mixture-of-Experts (MoE) | $0.14 | $0.86 | Joint embedding spaces |
| Qwen2.5-VL-32B-Instruct [94] | Transformer-based | $0.27 | $0.27 | Feature-level fusion |
Experimental Protocols and Methodologies
Implementing and evaluating multimodal AI for materials characterization requires robust, repeatable experimental protocols. The following sections detail key methodologies.
Protocol 1: Building a Multimodal Foundation Model with MultiMat
The MultiMat framework provides a methodology for training a foundation model for crystalline materials by aligning multiple modalities in a shared latent space [3].
Workflow Overview:
Multimodal AI Training Workflow
Step-by-Step Procedure:
- Data Acquisition and Modality Selection: Source data from materials databases like the Materials Project. For each material, gather the four key modalities [3]:
- Crystal Structure (C): Represented as atomic coordinates and lattice vectors.
- Density of States (ρ(E)): The electronic DOS as a function of energy.
- Charge Density (n_e(r)): The charge density as a function of position.
- Textual Description (T): Machine-generated descriptions of the crystal structure from a tool like Robocrystallographer [3].
- Encoder Training: Train a separate neural network encoder for each modality to map raw data into a shared latent space.
- Multimodal Alignment via Contrastive Learning: The core of the MultiMat framework. Use a contrastive loss function to align the embeddings of different modalities representing the same material. This encourages the model to learn a shared representation where, for example, the embedding of a crystal structure is semantically close to the embedding of its textual description [3].
- Downstream Task Execution:
- Property Prediction: Fine-tune the pre-trained crystal structure encoder (PotNet) on specific property prediction tasks (e.g., bandgap, formation energy). This transfer learning approach often leads to state-of-the-art predictive performance [3].
- Material Discovery: Screen large crystal structure databases by comparing a target property's latent space embedding with candidate crystal embeddings to find materials with desired characteristics [3].
Protocol 2: Automated Information Retrieval from Scientific Literature
This protocol outlines an automated workflow for extracting machine-readable data from scientific literature, a common challenge in materials science [54].
Workflow Overview:
Automated Scientific Data Extraction
Step-by-Step Procedure:
- Document Collection and Preprocessing: Assemble a corpus of scientific literature (PDFs) relevant to the research domain, such as microstructural analyses of face-centered cubic single crystals [54].
- Data Mining and Modality Separation: Use natural language processing (NLP) and computer vision techniques to decompose documents into distinct, machine-readable modalities: text, figures, tables, equations, and metadata [54].
- Multimodal Model Inference:
- Text and Equations: Process unstructured text and equations using a Large Language Model (LLM) or Transformer model to extract key concepts, material properties, and relationships [54].
- Figures and Charts: Analyze figures using a Vision Transformer (ViT) model to classify images, extract quantitative data from charts, and understand visual content [54].
- Tables: Employ structured data parsers and models to convert tabular information into a structured format (e.g., CSV, JSON).
- Database Generation and Knowledge Synthesis: Populate a machine-readable database with the extracted and structured information. This database can be enriched with local, unpublished, or private material data [54].
- Deployment of a Query Interface: Implement a Retrieval-Augmented Generation (RAG) system based on an LLM. This system uses the generated database as a knowledge source to enable a fast and efficient question-answering chatbot for researchers [54].
The Scientist's Toolkit: Essential Research Reagents & Platforms
Table 3: Key Platforms, Tools, and Data Repositories for Multimodal Materials Research
| Item Name | Type | Function in Research |
|---|---|---|
| Materials Project [3] | Data Repository | A core database providing computed properties of known and predicted materials, including crystal structures and electronic properties. |
| PotNet [3] | Graph Neural Network | A state-of-the-art GNN encoder specifically designed for crystal structures, used in frameworks like MultiMat. |
| Robocrystallographer [3] | Text Generation Tool | Automatically generates textual descriptions of crystal structures and their symmetry, providing a natural language modality for training. |
| CLIP [3] | AI Model | A foundational contrastive learning model for aligning visual and textual concepts; its principles are adapted for materials science in MultiMat. |
| Vertex AI [95] | AI Platform | Google's platform offering built-in data pipelines and batch processing for models like Gemini, with data residency controls for regulated industries. |
| MMTBench [96] | Benchmark Dataset | A benchmark for evaluating multimodal table reasoning, useful for testing model capabilities on complex, real-world data presentations. |
| SiliconFlow [94] | AI Service Platform | Provides access and deployment services for a variety of multimodal models, including GLM-4.5V and Qwen2.5-VL. |
| Galileo [93] | Evaluation Platform | An evaluation intelligence platform for monitoring, evaluating, and debugging multimodal AI systems in production. |
Evaluation and Challenges in Multimodal AI
The complexity of multimodal systems necessitates robust evaluation strategies that go beyond traditional unimodal metrics. Key performance indicators must capture the nuances of each modality and their interactions [93]. Quantitative metrics like accuracy and F1 score should be paired with qualitative assessments of output coherence, context adherence, and user satisfaction [93]. Benchmarks like MMTBench are emerging to rigorously test capabilities such as complex table reasoning, which is endemic to scientific reporting [96].
Several challenges persist in the implementation of multimodal AI:
- Data Integration Complexity: Aligning diverse data types (text, images, audio) with different structures and quality at scale poses significant issues for real-time processing and latency [93].
- Model Hallucinations: The risk of models generating nonsensical or incorrect outputs can be heightened by complex interactions between different data modalities [93].
- Interpretability and Trust: The "black box" nature of many complex models can cause end-user skepticism, which is particularly problematic in high-stakes fields like drug development [93] [92]. Visualization techniques are being developed to help analysts interpret and explore multimodal data to gain actionable insights and build trust in model outcomes [92].
Multimodal AI models represent a paradigm shift in computational materials science, offering unprecedented capabilities for characterizing and discovering new materials. This analysis demonstrates that model selection is highly use-case dependent: Gemini 2.5 Pro is suited for massive document review, Claude Opus for high-stakes analytical tasks, and open-source models like Llama 4 Maverick for environments requiring full data control. Frameworks like MultiMat exemplify the trend towards building specialized foundation models for scientific domains by aligning rich, complementary data modalities. As the field progresses, success will depend on overcoming key challenges related to data integration, model interpretability, and the development of standardized, modular AI systems. For researchers, the strategic adoption of these tools, guided by rigorous experimental protocols and evaluation frameworks, is poised to dramatically accelerate the pace of innovation in materials informatics and drug development.
The integration of large language models (LLMs) into scientific research represents a paradigm shift, yet their capabilities in highly specialized domains like materials characterization have remained largely unvalidated. MatQnA directly addresses this critical gap by establishing the first multi-modal benchmark dataset specifically designed for material characterization techniques [97] [38]. This benchmark emerges within the broader context of advancing multimodal data parsing for materials informatics, enabling systematic evaluation of AI's ability to interpret complex experimental data that integrates both text and image modalities [98]. As LLMs demonstrate remarkable breakthroughs in general domains, their application to scientific research scenarios necessitates domain-specific validation frameworks to assess true comprehension and reasoning capabilities [38]. MatQnA provides this essential validation framework, offering researchers a standardized resource for quantifying AI performance in interpreting the complex, multimodal data inherent to materials characterization.
MatQnA Dataset Characteristics and Technical Specifications
Scope and Composition
MatQnA is architected as a comprehensive multi-modal evaluation resource, comprising over 5,000 meticulously curated question-answer pairs derived from more than 400 peer-reviewed journal articles and expert case studies [98] [38]. The dataset encompasses ten mainstream material characterization techniques, spanning the core methodological spectrum of materials analysis from structural and chemical analysis to microscopy and thermal characterization [97] [98].
Table 1: Characterization Techniques Covered in MatQnA Dataset
| Characterization Technique | Analytical Focus | Primary Modality |
|---|---|---|
| X-ray Photoelectron Spectroscopy (XPS) | Chemical state, element identification, peak assignment | Image, Text |
| X-ray Diffraction (XRD) | Crystal structure, phase identification, grain sizing | Image, Text |
| Scanning Electron Microscopy (SEM) | Surface morphology, defect analysis | Image |
| Transmission Electron Microscopy (TEM) | Internal lattice, microstructure | Image |
| Atomic Force Microscopy (AFM) | 3D topography, surface roughness | Image |
| Differential Scanning Calorimetry (DSC) | Thermal transitions, enthalpy changes | Chart |
| Thermogravimetric Analysis (TGA) | Decomposition behavior, thermal stability | Chart |
| Fourier Transform Infrared Spectroscopy (FTIR) | Chemical bonds, vibrational modes | Spectrum |
| Raman Spectroscopy | Molecular vibration, phase composition | Spectrum |
| X-ray Absorption Fine Structure (XAFS) | Atomic environment, oxidation states | Spectrum |
The dataset achieves a balanced representation across question types, containing 2,749 subjective (open-ended) questions and 2,219 objective (multiple-choice) questions [98] [99]. This distribution enables comprehensive assessment of both factual recognition capabilities and explanatory reasoning skills in AI models.
The foundation of MatQnA rests upon materials science data accumulated from the Scientific Compass platform, a leading comprehensive scientific research service platform in China [38]. Source documents underwent rigorous selection through keyword matching targeting specific characterization methodologies, ensuring domain relevance and technical depth [38]. The dataset construction prioritized academically rigorous and structurally standardized content from high-impact domestic and international journals, focusing particularly on sections related to structural characterization, morphology analysis, spectral interpretation, and figure-text correlation [38]. This multi-source, heterogeneous corpus preserves the logical chain of materials science knowledge within authentic experimental contexts, providing a robust foundation for developing and evaluating large-scale models with domain-specific understanding and cross-modal reasoning abilities [38].
Dataset Construction Methodology: A Hybrid Human-AI Approach
The MatQnA dataset was assembled through an sophisticated hybrid methodology that synergistically combines LLM-assisted generation with human-in-the-loop validation, ensuring both scalability and scientific rigor [98] [99]. This multi-stage workflow transforms raw research documents into high-quality benchmark questions through the following systematic process:
Preprocessing and Data Extraction
The construction pipeline begins with preprocessing, where a multi-source heterogeneous corpus is compiled from source documents initially in PDF format [99]. These documents undergo structured parsing using PDF Craft, an open-source deep learning-based multimodal PDF parser that extracts text, images, and document structures, producing flexible outputs like Markdown that are crucial for subsequent automated processing [99]. Following preprocessing, benchmark data synthesis involves filtering irrelevant content by building a keyword-based retrieval system and a text-image index aligned with the document structure [99]. Relevant text fragments and corresponding images for each of the ten characterization techniques are then prepared for question generation.
Automated Question-Answer Generation
The core of dataset creation employs OpenAI's GPT-4.1 API guided by predefined prompt templates to automatically generate structured question-answer pairs encompassing both multiple-choice and subjective questions [98] [38] [99]. This approach leverages the generative capability of advanced LLMs while maintaining scientific relevance and quality, with each content unit yielding up to five questions [99]. The automated generation significantly enhances dataset scalability while ensuring broad coverage of materials characterization concepts.
Post-Processing and Quality Assurance
Critical post-processing procedures address inherent limitations in LLM generation, implementing scientific rigor through two key mechanisms [99]. Coreference resolution applies regex-based normalization to automatically detect and resolve ambiguous references within questions (e.g., "based on the given content," "this figure"), substantially improving clarity and objectivity [99]. Simultaneously, self-containment enforcement introduces image non-nullity checks during data generation to ensure each QA item incorporates adequate multimodal context, guaranteeing interpretability and validity even when answers might not be derivable from text alone [99].
Expert Validation and Final Curation
The final quality gate employs a two-stage human validation process conducted by materials science experts [98] [99]. This critical filtering mechanism ensures terminological correctness, logical coherence in answer reasoning, alignment with materials science principles, and question relevance, systematically removing items with limited analytical value or weak domain relevance [99]. The resulting dataset comprises 4,968 high-quality questions stored in Parquet format, organized by characterization technique for optimal accessibility and usability [99].
Experimental Evaluation Protocol and Performance Metrics
Benchmarking Methodology
The MatQnA evaluation framework employs rigorous experimental protocols to assess multi-modal LLM capabilities in materials characterization tasks [38] [99]. Preliminary evaluations have focused on five mainstream multi-modal LLMs: GPT-4.1, Claude Sonnet 4, Gemini 2.5 Flash, Qwen2.5 VL 72B, and Doubao Vision Pro 32K [99]. The assessment concentrates on objective questions with performance measured primarily by accuracy, conducted systematically on the Phoenix evaluation platform to ensure consistency and reproducibility [99].
The evaluation design incorporates nuanced difficulty stratification, classifying questions into three distinct categories based on model performance: easy (accuracy ≥ 0.80), medium (accuracy 0.50-0.79), and hard (accuracy < 0.50) [99]. This stratification enables fine-grained analysis of model capabilities across varying complexity levels, providing insights beyond aggregate performance metrics.
Performance Results and Analysis
Comprehensive evaluation reveals strong capabilities in state-of-the-art multi-modal models, with overall accuracy scores ranging from 86.3% to 89.8% across the tested models [99]. The performance distribution across techniques and models demonstrates significant variation, highlighting specialized capabilities and limitations.
Table 2: Model Performance Evaluation on MatQnA Objective Questions
| Multimodal LLM | Overall Accuracy | Highest Performing Category | Most Challenging Category |
|---|---|---|---|
| GPT-4.1 | 89.8% | FTIR, Raman | AFM |
| Claude Sonnet 4 | 89.7% | TGA, Raman | AFM |
| Gemini 2.5 Flash | 89.6% | FTIR, XAFS | AFM |
| Doubao Vision Pro 32K | 89.6% | Raman, FTIR | AFM |
| Qwen2.5 VL 72B | 86.3% | TGA, DSC | AFM |
Analysis of technique-specific performance reveals that FTIR, Raman, and TGA emerge as high-performance categories with accuracy exceeding 90%, while AFM consistently proves most challenging with accuracy ranging from 79.7% to 84.7% across models [99]. This performance pattern suggests that while current models demonstrate strong proficiency in standard spectral data interpretation, techniques requiring complex three-dimensional spatial reasoning present persistent challenges [98] [99].
Fine-grained sub-category analysis further illuminates specific capabilities, with "decomposition mechanism and reaction pathway analysis" achieving the highest accuracy (99.0%), while "phase transition temperature analysis" represents the most challenging medium-difficulty sub-category (82.0%) [99]. Comparative analysis indicates that Doubao Vision Pro 32K and GPT-4.1 demonstrate high, stable accuracy across diverse sub-categories, with Doubao exhibiting particular strength in multimodal tasks [99].
Essential Research Reagents: The Materials Characterization Toolkit
The MatQnA benchmark encompasses a comprehensive suite of materials characterization techniques, each with specialized analytical capabilities essential for modern materials research. The dataset's coverage spans the fundamental methodological toolkit required for advanced materials development and analysis.
Table 3: Essential Characterization Techniques in Materials Research
| Technique | Primary Function | Key Analytical Applications |
|---|---|---|
| XPS (X-ray Photoelectron Spectroscopy) | Surface chemical analysis | Elemental composition, chemical state, electronic state |
| XRD (X-ray Diffraction) | Crystalline structure analysis | Phase identification, crystal structure, grain size measurement |
| SEM (Scanning Electron Microscopy) | High-resolution surface imaging | Surface morphology, microstructure, defect analysis |
| TEM (Transmission Electron Microscopy) | Nanoscale internal structure imaging | Lattice structure, crystal defects, nanoparticle characterization |
| AFM (Atomic Force Microscopy) | 3D surface topography | Surface roughness, nanomechanical properties |
| DSC (Differential Scanning Calorimetry) | Thermal properties analysis | Phase transitions, melting behavior, crystallization studies |
| TGA (Thermogravimetric Analysis) | Thermal stability assessment | Decomposition temperatures, compositional analysis |
| FTIR (Fourier Transform Infrared Spectroscopy) | Molecular bond identification | Functional group analysis, chemical bonding |
| Raman Spectroscopy | Molecular vibration characterization | Crystal structure, phase composition, stress measurement |
| XAFS (X-ray Absorption Fine Structure) | Local atomic structure analysis | Oxidation states, coordination chemistry |
The application of these techniques frequently involves quantitative analytical methods, such as the Scherrer equation for XRD grain size estimation: ( L = \frac{K\lambda}{\beta \cos \theta} ), where ( L ) represents crystallite size, ( \lambda ) the X-ray wavelength, ( \beta ) the peak width, and ( \theta ) the Bragg angle [98]. MatQnA specifically evaluates understanding of such domain-specific quantitative relationships alongside qualitative interpretation skills.
Applications and Implications for Materials Informatics
MatQnA establishes a critical foundation for advancing AI capabilities in materials science through several key applications. The benchmark enables rigorous, standardized evaluation of LLMs in materials characterization, providing researchers with quantitative metrics for model selection and development [98]. By diagnosing specific strengths and weaknesses across characterization techniques, it guides targeted model improvement, particularly in challenging areas like spatial reasoning for AFM interpretation [98] [99].
The dataset further facilitates AI-assisted materials discovery workflows, enabling development of systems that can interpret experimental data, predict material properties, and provide scientific recommendations [98]. This supports accelerated research cycles and enhanced experimental design. Additionally, MatQnA provides a framework for domain-specific model fine-tuning, allowing developers to create specialized systems with enhanced performance on materials science tasks [98].
Beyond immediate applications in materials science, MatQnA demonstrates the feasibility of extending sophisticated LLM evaluation frameworks to other specialized scientific domains, establishing a methodology for creating domain-specific benchmarks that require deep technical knowledge and multi-modal reasoning capabilities [98] [99]. This approach has significant potential for accelerating AI integration across scientific disciplines requiring complex data interpretation.
This case study examines the implementation of a multi-institutional patient portal, "MyChart," in Southwestern Ontario, Canada. It analyzes the quantitative adoption metrics and qualitative stakeholder feedback to extract critical lessons on data aggregation, user engagement, and strategic management. These findings are then contextualized within the emerging paradigm of multimodal data parsing for materials information research, illustrating how principles of unified data access and interoperable systems are foundational to accelerating discovery across scientific domains.
The central challenge in both healthcare and advanced materials research is the fragmentation of critical information across disparate systems and organizations. In healthcare, this siloing jeopardizes patient safety and increases costs [100]. Similarly, in materials science, crucial data from experiments, simulations, and literature often reside in incompatible formats, hindering the discovery of new materials. The implementation of portals that provide a single, unified access point to multi-source data is a critical step toward solving these issues. This case study first details a real-world deployment in healthcare and then explores its broader implications for managing complex, multimodal scientific data.
Core Case Study: MyChart Patient Portal in Southwestern Ontario
This section provides a detailed analysis of a specific multi-institutional portal implementation.
The MyChart portal was deployed in Southwestern Ontario to provide residents with integrated access to their clinical data from across the health system. It was integrated with "ClinicalConnect," a pre-existing provider-facing data viewer that consolidated information from 72 acute care hospitals and other organizations, rather than connecting directly to individual hospital record systems [100]. Organizations signed data-sharing agreements to permit their data to flow to the patient-accessible MyChart.
Quantitative data from the first 15 months of implementation (August 2018 to October 2019) are summarized in the table below.
Table 1: MyChart Portal Adoption Metrics (Aug 2018 - Oct 2019)
| Metric | Value | Description |
|---|---|---|
| Registration Emails Sent | 15,271 | Invitations sent to potential users. |
| Successful Registrations | 10,233 (67.01%) | Number of patients who created an account. |
| Participating Sites | 38 | Healthcare sites actively offering the portal. |
| Median Registration per Site | 19 | Median number of patients registered per site. |
| Range of Registration per Site | 1 - 2,114 | Highlighting significant variation in adoption across sites. |
Experimental Protocol and Methodology
The evaluation of the MyChart implementation employed a multimethod study design [100]:
- Quantitative Analysis: Routinely collected, aggregate usage data was extracted from the portal by the vendor to understand use patterns.
- Qualitative Data Collection: 42 semi-structured interviews were conducted with key stakeholders.
- Participants: 18 administrative stakeholders, 16 patients, 7 health care providers, and 1 informal caregiver.
- Objective: To understand how the implementation approach influenced user experiences and to identify strategies for improvement.
- Analysis: Interview data were analyzed using an inductive thematic approach.
Key Findings and Implementation Challenges
The study identified several critical factors that influenced the portal's adoption and effectiveness [100]:
- Data Comprehensiveness: Interview participants perceived that the patient experience would have been significantly improved by enhancing the completeness of the data available through the portal. Inconsistent data uploads from participating sites limited utility.
- Rollout and Engagement Strategy: The lack of a broad, coordinated rollout and marketing strategy across all 38 sites was identified as a major barrier to enrollment.
- Organizational and Cultural Buy-in: Provider engagement, change management support, and endorsement from senior leadership were cited as central to fostering uptake. The absence of these elements at some sites contributed to low registration numbers.
- Policy and Regional Alignment: Participants stated that regional alignment and top-down policy support should have been sought to streamline implementation efforts and create consistency across participating sites.
The core workflow of this implementation, from data aggregation to user access, is diagrammed below.
Parallels in Materials Information Research
The challenges observed in the MyChart case study directly mirror those in materials science. A majority of materials information is encoded within scientific documents, tables, and figures, limiting machine readability and the ability to find suitable literature or compare material properties [54]. The field is moving towards automated workflows that can parse this multimodal data.
An Automated Workflow for Multimodal Data Parsing
Recent research demonstrates an automated workflow to transform encoded scientific information into a machine-readable database [54]. The methodology for this workflow involves:
- Data Ingestion: A corpus of scientific literature and local, unpublished experimental data is collected.
- Multi-Modal Information Extraction:
- Natural Language Processing (NLP) & Large Language Models (LLMs): Extract textual information from publications.
- Vision Transformer (ViT) Models: Decode data from figures, tables, and diagrams.
- Knowledge Synthesis: The extracted information is structured into a unified, machine-readable database.
- Intelligent Querying: A Retrieval-Augmented Generation (RAG)-based LLM is deployed on the synthesized database to enable fast and efficient question-answering, accelerating information retrieval and material property extraction.
This process, which creates a foundational tool for materials research, is visualized below.
The Scientist's Toolkit: Essential Research Reagents & Solutions
The following table details key components in the advanced materials discovery pipeline, as exemplified by systems like the CRESt (Copilot for Real-world Experimental Scientists) platform developed at MIT [39].
Table 2: Key Reagents & Solutions for Automated Materials Research
| Item Name | Function / Role in Research |
|---|---|
| Liquid-Handling Robot | Automates the precise synthesis of material samples by mixing precursor chemicals in varied ratios, enabling high-throughput experimentation. |
| Carbothermal Shock System | Rapidly synthesizes materials by applying extremely high temperatures for short durations, facilitating the quick creation of diverse material chemistries. |
| Automated Electrochemical Workstation | Tests and characterizes the performance of newly synthesized materials (e.g., as catalysts in fuel cells) without manual intervention. |
| Automated Electron Microscope | Provides high-resolution microstructural images and analysis of synthesized materials, with data fed directly into the analysis models. |
| Multimodal Active Learning Models | AI models that incorporate diverse data sources (literature, experimental results, images) to suggest optimal new material recipes and experiments. |
| Computer Vision & Vision Language Models | Monitors experiments via cameras, detects issues (e.g., sample misplacement), and suggests corrections to ensure reproducibility. |
Integrated Data Management Protocols
This section outlines the consolidated methodologies from the featured case study and research domain.
Protocol for Portal Implementation & Management
Based on the MyChart evaluation, a successful multi-institutional portal implementation requires [100]:
- Pre-Implementation Alignment: Secure senior leadership endorsement and establish aligned data-sharing policies across all participating institutions before rollout.
- Centralized Change Management: Develop and execute a broad, coordinated marketing and enrollment strategy, rather than leaving engagement to individual sites.
- Ensure Data Comprehensiveness: Prioritize technical interoperability to ensure consistent and complete data flow from all source systems into the unified portal.
- Engage End-Users Early: Involve providers and patients in the design and feedback process to foster a cultural shift and ensure the portal meets real-world needs.
Protocol for AI-Driven Materials Discovery
The CRESt platform demonstrates a protocol for closed-loop materials discovery [39]:
- Hypothesis Generation: Use large language models to search scientific literature and suggest promising material chemistries and recipes.
- Robotic Synthesis: Employ liquid-handling robots and carbothermal shock systems to synthesize proposed material samples at high throughput.
- Automated Characterization & Testing: Utilize automated workstations and electron microscopes to characterize the structure and test the performance of synthesized materials.
- AI Analysis and Optimization: Feed experimental results and images into active learning models. These models, augmented by literature knowledge and human feedback, plan the next round of experiments to iteratively optimize material properties.
The MyChart case study underscores that the success of a multi-institutional data portal hinges not merely on the technology itself, but on strategic implementation, robust data governance, and strong organizational partnerships. The parallel work in materials science demonstrates how these same principles are being operationalized through AI and automation to manage and parse complex multimodal information. Together, they provide a compelling framework for the future of data-intensive research: one built on unified data access, interoperable systems, and intelligent tools that transform fragmented data into actionable knowledge.
In the field of materials information research, the integration of artificial intelligence has ushered in a new era of accelerated discovery. Central to this transformation is the development of sophisticated multimodal AI systems capable of parsing and interpreting complex scientific data. The performance of these systems hinges on three interdependent pillars: Accuracy in predictions and experimental outcomes, Generalization across diverse domains and unseen data, and Cross-Modal Reasoning capabilities that enable the synthesis of information from disparate sources. This technical guide provides an in-depth examination of the methodologies, protocols, and metrics essential for rigorously evaluating these capabilities within the specific context of multimodal data parsing for materials science, offering researchers a structured framework for assessment and implementation.
Quantitative Performance Metrics and Benchmarks
A critical first step in performance assessment is the establishment of robust quantitative metrics. The following tables summarize key performance indicators across different multimodal systems, providing a baseline for comparison and evaluation.
Table 1: Performance Benchmarks of Multimodal AI Systems in Scientific Domains
| System Name | Primary Domain | Key Performance Metric | Reported Result | Benchmark Used |
|---|---|---|---|---|
| CRESt [39] | Materials Science | Power Density Improvement | 9.3-fold improvement per dollar | Direct Formate Fuel Cell |
| CRESt [39] | Materials Science | Experimental Throughput | 900+ chemistries, 3,500+ tests | Internal Workflow |
| R1-Onevision [101] | Generalized Multimodal Reasoning | Benchmark Performance | Outperformed GPT-4o & Qwen2.5-VL | R1-Onevision-Bench |
| Doc-Researcher [102] | Document Understanding | Accuracy on Complex QA | 50.6% Accuracy | M4DocBench |
| LMM-KC Generation [91] | Educational KT | Predictive Performance | Comparable/Superior to Human KCs | Multiple OLI Datasets |
Table 2: Core Metrics for Assessing Multimodal System Capabilities
| Assessment Dimension | Specific Metric | Measurement Method |
|---|---|---|
| Accuracy | Predictive Power Density | Experimental validation in operational fuel cells [39]. |
| Accuracy | Knowledge Component (KC) Quality | Performance in Knowledge Tracing (KT) models vs. human-tagged KCs [91]. |
| Generalization | Cross-Domain Accuracy | Performance on unseen domains in M4DocBench (Multi-document, Multi-hop) [102]. |
| Generalization | Domain Shift Robustness | Accuracy drop measured when applying models to new database distributions [103]. |
| Cross-Modal Reasoning | Complex QA Accuracy | Success rate on questions requiring synthesis of text, tables, and figures [102]. |
| Cross-Modal Reasoning | Iterative Reasoning Capability | Ability to refine answers through multi-turn, evidence-gathering workflows [102]. |
Experimental Protocols for Performance Assessment
Protocol 1: High-Throughput Materials Discovery and Validation
This protocol, derived from the CRESt system, assesses a platform's ability to accurately discover and optimize new materials through automated, iterative experimentation [39].
- Problem Formulation: Researchers define the objective in natural language (e.g., "find a low-cost, high-activity catalyst for a direct formate fuel cell").
- Multimodal Knowledge Integration: The system ingests and represents data from diverse sources:
- Scientific Literature: Previous findings on element behavior are embedded into a knowledge representation space.
- Human Feedback: Researcher intuition and guidance are incorporated via natural language input.
- Search Space Definition: Principal Component Analysis (PCA) is performed on the knowledge embeddings to reduce dimensionality and define a viable search space.
- Bayesian Optimization (BO): An active learning loop is initiated within the reduced search space. BO suggests the most promising next material recipe based on all accumulated data.
- Robotic Synthesis and Testing: A high-throughput robotic system executes the suggested experiments. This typically includes:
- Synthesis: Using a liquid-handling robot and a carbothermal shock system.
- Characterization: Automated electron microscopy and optical microscopy.
- Performance Testing: An automated electrochemical workstation.
- Computer Vision Monitoring: Cameras and vision language models monitor experiments in real-time to detect issues (e.g., sample misplacement) and suggest corrections to ensure reproducibility.
- Data Feedback Loop: Results from synthesis, characterization, and testing are fed back into the multimodal model to augment the knowledge base and refine the search space for the next iteration.
- Final Validation: The top-performing material is validated in a real-world device (e.g., a fuel cell) to measure ultimate performance metrics like power density.
Protocol 2: Knowledge Component Extraction for Predictive Modeling
This protocol evaluates a system's cross-modal reasoning accuracy by its ability to extract meaningful Knowledge Components (KCs) from educational content, which are then used to predict student performance [91].
- Data Preprocessing: Collect and preprocess educational datasets containing student interaction data and multimedia question information (text, images).
- Convert any audio (e.g., MP3) to text using a model like Whisper-large-v2.
- Extract images from legacy file formats where necessary.
- Multimodal Parsing: Use a Large Multimodal Model (LMM), such as GPT-4o, to parse the educational content. The model ingests both the textual and visual information from each question.
- Zero-Shot KC Generation: Prompt the LMM in a zero-shot manner to identify and describe the inherent knowledge components required to solve each question.
- Clustering: Generate sentence embeddings for all extracted KC descriptions. Use clustering algorithms (e.g., K-means) to group semantically similar KCs, creating a unified set of components for the dataset.
- Model Validation: Integrate the LMM-generated KCs into established Knowledge Tracing (KT) models (e.g., Performance Factors Analysis - PFA, Deep Knowledge Tracing - DKT).
- Performance Comparison: Train and evaluate the KT models using LMM-generated KCs versus models using traditional human-tagged KCs. The key metric is the predictive accuracy of student performance on future questions. Success is achieved when LMM-generated KCs perform comparably or superior to human-generated labels.
Visualization of Core Workflows and Architectures
The following diagrams, generated with Graphviz DOT language, illustrate the core experimental and reasoning workflows described in the protocols. The color palette adheres to the specified guidelines, ensuring high contrast for readability.
Diagram 1: CRESt High-Throughput Materials Discovery Workflow
Diagram 2: Knowledge Component Extraction and Validation Pipeline
Diagram 3: Doc-Researcher Deep Multimodal Research Architecture
The Scientist's Toolkit: Research Reagent Solutions
For researchers aiming to implement or benchmark similar multimodal systems, the following table details essential computational "reagents" and tools referenced in the assessed studies.
Table 3: Essential Research Reagents and Tools for Multimodal Materials Research
| Tool / Component | Type | Primary Function | Exemplar Use Case |
|---|---|---|---|
| Large Multimodal Model (LMM) [91] [102] | Algorithmic Model | Cross-modal understanding and reasoning; extracts concepts from text & images. | GPT-4o, Qwen2.5-VL for parsing scientific documents and generating Knowledge Components. |
| Bayesian Optimization (BO) [39] | Algorithmic Framework | Optimizes experimental design by balancing exploration and exploitation in a defined search space. | Suggests the next most promising material recipe in the CRESt system. |
| High-Throughput Robotic System [39] | Hardware/Software Suite | Automates the synthesis, characterization, and testing of material samples. | Executes thousands of electrochemical tests to validate AI predictions. |
| Computer Vision Monitoring [39] | Analysis Tool | Provides real-time visual feedback on experiments to detect and correct anomalies. | Ensures reproducibility in sample preparation and characterization. |
| Deep Multimodal Parser (e.g., MinerU) [102] | Software Library | Performs layout-aware parsing of complex documents, preserving structure of tables, figures, and equations. | Converts PDF scientific papers into structured, machine-readable formats for retrieval. |
| Systematic Retrieval Architecture [102] | Software Framework | Enables efficient search across multi-granular (chunk, page, document) and multimodal (text, image) data. | Finds relevant evidence from a large corpus of scientific documents for a complex query. |
| Specialized Benchmark (e.g., M4DocBench) [102] | Dataset/Metric Suite | Provides a rigorous, expert-annotated standard for evaluating multi-hop, multi-document, and multi-modal reasoning. | Measures the true cross-modal reasoning and generalization capabilities of a deep research system. |
The accurate assessment of AI systems for multimodal materials research demands a holistic approach that integrates quantitative metrics, robust experimental protocols, and a clear understanding of the underlying architectures. The frameworks and data presented herein demonstrate that state-of-the-art systems are moving beyond unimodal data analysis towards integrated platforms that synergistically combine literature knowledge, human expertise, robotic experimentation, and cross-modal reasoning. This integration is key to tackling long-standing challenges in materials science, such as the discovery of novel catalysts, by achieving not only high predictive accuracy but also the generalization and reasoning capabilities necessary for genuine scientific innovation. As these systems evolve, the continuous development and adoption of standardized benchmarks and assessment methodologies will be critical for measuring progress and guiding future research.
Conclusion
Multimodal data parsing represents a paradigm shift in materials informatics, enabling unprecedented integration of complementary data types to uncover complex processing-structure-property relationships. By leveraging advanced fusion techniques, contrastive learning, and specialized benchmarks, researchers can overcome longstanding challenges of data heterogeneity and scarcity. The maturation of these approaches, evidenced by frameworks like MatMCL and benchmarks like MatQnA, signals a move toward more predictive, AI-driven materials design. For biomedical and clinical research, these advancements promise accelerated therapeutic material development, enhanced characterization of drug delivery systems, and more efficient extraction of insights from multimodal experimental data. Future progress will depend on developing more sophisticated cross-modal alignment algorithms, creating larger standardized datasets, and building integrated data infrastructures that support the entire materials innovation lifecycle from discovery to clinical application.
References
- 1. Modality – Knowledge and References [https://taylorandfrancis.com/knowledge/Engineering_and_technology/Electrical_%26_electronic_engineering/Modality/]
- 2. A versatile multimodal learning framework bridging ... [https://www.nature.com/articles/s41524-025-01767-3]
- 3. Multimodal Learning for Materials [https://arxiv.org/html/2312.00111v2]
- 4. Towards understanding structure–property relations in ... [https://www.nature.com/articles/s41524-023-01163-9]
- 5. Materials informatics approach using domain modelling for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9217937/]
- 6. Workflow Basics - Materials informatics for structure- ... [https://www.molmd.org/mispr/html/workflows/basics.html]
- 7. A Study on Methods for Parsing Architectural Multi-Modal ... [https://www.mdpi.com/2075-5309/15/22/4048]
- 8. A materials informatics approach for composition and ... [https://www.sciencedirect.com/science/article/pii/S2590049823000449]
- 9. Recent advances and applications of deep learning ... [https://www.nature.com/articles/s41524-022-00734-6]
- 10. A survey of AI-supported materials informatics [https://www.sciencedirect.com/science/article/pii/S1574013725001212]
- 11. Exploring the Spectrum of Analytical Techniques for ... [https://www.spectroscopyonline.com/view/exploring-the-spectrum-of-analytical-techniques-for-material-characterization]
- 12. Microscopy Data - an overview [https://www.sciencedirect.com/topics/computer-science/microscopy-data]
- 13. Textual Material - an overview [https://www.sciencedirect.com/topics/engineering/textual-material]
- 14. Guide | Working with Textual Data [https://nodegoat.net/guide.p/448.m/139/working-with-textual-data]
- 15. Tabular data — Introduction to Materials Informatics [https://enze-chen.github.io/mi-book-2021/week_1/02/tabular_data_blank.html]
- 16. How does spectral sensing work? Understanding the ... [https://www.specim.com/technology/how-does-spectral-sensing-work/]
- 17. Spectral Data | Spectroradiometers | Applications [https://www.admesy.com/applications/spectral-data/]
- 18. Data augmentation in microscopic images for material ... [https://www.nature.com/articles/s41524-020-00392-6]
- 19. Materials informatics: A review of AI and machine learning ... [https://www.sciencedirect.com/science/article/pii/S2352492825020379]
- 20. Overcoming data scarcity with transfer learning [https://citrine.io/overcoming-data-scarcity-with-transfer-learning/]
- 21. A Case Study of Multimodal, Multi-Institutional Data ... [https://citrine.io/a-case-study-of-multimodal-multi-institutional-data-management-for-the-combinatorial-materials-science-community/]
- 22. Addressing Model and Data Heterogeneity in Multimodal ... [https://arxiv.org/html/2408.04275v1]
- 23. Automation Revolutionizes High Throughput ... [https://chronect.trajanscimed.com/news/high-throughput-experimentation-hte]
- 24. Flow chemistry as a tool for high throughput experimentation [https://pubs.rsc.org/en/content/articlehtml/2025/dd/d5dd00129c]
- 25. FAIR Principles [https://www.go-fair.org/fair-principles/]
- 26. News - FAIR-DI [https://fairmat-nfdi.eu/nomad-lab/news-nomad-lab]
- 27. A Guide to the FAIR Principles [https://www.openscience.eu/article/infrastructure/guide-fair-principles]
- 28. Advancing materials science through data integration and ... [https://link.springer.com/article/10.1557/s43577-025-00965-0]
- 29. End-to-End Models for Complex AI Tasks [https://www.capitalone.com/tech/machine-learning/pros-and-cons-of-end-to-end-models/]
- 30. Prompt2DAG: A Modular Methodology for LLM-Based Data ... [https://arxiv.org/html/2509.13487v1]
- 31. Modular and cloud-based bioinformatics pipelines for high ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12380322/]
- 32. MatVQA for Challenging Visual-Scientific Reasoning in ... [https://arxiv.org/html/2505.18319v1]
- 33. Towards a data platform for multimodal 4D mechanics of ... [https://www.sciencedirect.com/science/article/pii/S0264127524006816]
- 34. Document Parsing Unveiled: Techniques, Challenges, and ... [https://arxiv.org/html/2410.21169v2?ref=chitika.com]
- 35. Materials Informatics 2025-2035: Markets, Strategies, Players [https://www.idtechex.com/en/research-report/materials-informatics-2025/1096]
- 36. Real-Time Multimodal Data Processing with Pathway and ... [https://pathway.com/blog/multimodal-data-processing]
- 37. Article MERMaid: Universal multimodal mining of chemical ... [https://www.sciencedirect.com/science/article/pii/S2590238525003741]
- 38. MatQnA: A Benchmark Dataset for Multi-modal Large ... [https://arxiv.org/html/2509.11335v1]
- 39. AI system learns from many types of scientific information ... [https://news.mit.edu/2025/ai-system-learns-many-types-scientific-information-and-runs-experiments-discovering-new-materials-0925]
- 40. Using Large Multimodal Models to Extract Knowledge ... [https://educationaldatamining.org/EDM2025/proceedings/2025.EDM.long-papers.170/]
- 41. Elements must meet minimum color contrast ratio thresholds [https://dequeuniversity.com/rules/axe/4.8/color-contrast]
- 42. Multimodal deep learning for biomedical data fusion: a review [https://pmc.ncbi.nlm.nih.gov/articles/PMC8921642/]
- 43. Early Fusion vs. Late Fusion in Multimodal Data Processing [https://www.geeksforgeeks.org/deep-learning/early-fusion-vs-late-fusion-in-multimodal-data-processing/]
- 44. Effective Techniques for Multimodal Data Fusion [https://pmc.ncbi.nlm.nih.gov/articles/PMC10007548/]
- 45. Chapter 4 Further Topics | Multimodal Deep Learning [https://slds-lmu.github.io/seminar_multimodal_dl/c03-00-further.html]
- 46. Joint Representation Learning Vs Coordinated Learning in ... [https://medium.com/@nikitaparate9/joint-representation-learning-vs-coordinated-learning-in-multimodal-context-01890e0a4730]
- 47. A Comparative Analysis of Three Data Fusion Methods and ... [https://www.mdpi.com/2227-7390/13/8/1218]
- 48. Multimodal fusion techniques: Review, data representation ... [https://www.sciencedirect.com/science/article/abs/pii/S0925231225014997]
- 49. [2011.07191] On the Benefits of Early Fusion in Multimodal ... [https://arxiv.org/abs/2011.07191]
- 50. An effective multimodal representation and fusion method ... [https://www.sciencedirect.com/science/article/abs/pii/S0925231223004964]
- 51. Multimodal Representation Learning and Fusion [https://arxiv.org/html/2506.20494v1]
- 52. Cross-modal alignment and contrastive learning for ... [https://pubmed.ncbi.nlm.nih.gov/39961170/]
- 53. Cross-modal alignment and contrastive learning for ... [https://www.sciencedirect.com/science/article/abs/pii/S0169260725000501]
- 54. Towards an automated workflow in materials science for ... [https://www.sciencedirect.com/science/article/pii/S2352492825006981]
- 55. Human-centered Interactive Learning via MLLMs for Text ... [https://arxiv.org/html/2506.11036v1]
- 56. Cross-Modal Alignment Enhancement for Vision ... [https://www.mdpi.com/2673-2688/6/10/263]
- 57. A Case Study of Multimodal, Multi-institutional Data ... [https://link.springer.com/article/10.1007/s40192-024-00345-7]
- 58. A Case Study of Multimodal, Multi-institutional Data ... [https://www.osti.gov/biblio/2403654]
- 59. Electrospun nanofibers and their application as sensors for ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1533367/full]
- 60. Influence of electrospinning parameters on the ... [https://www.sciencedirect.com/science/article/pii/S2949829525001640]
- 61. Applications of Advanced Electrospun Nanofibrous ... [https://www.mdpi.com/2079-4991/15/18/1424]
- 62. Advancements in Thermoelectric Materials and Their ... [https://www.sciencedirect.com/science/article/pii/S2949746925000321]
- 63. The Discovery of Flexible Thermoelectric Materials [https://www.msesupplies.com/blogs/news/turning-heat-into-power-the-discovery-of-flexible-thermoelectric-materials?srsltid=AfmBOorL3V1ZtzWoVfcGxXxVcqnUsGlGhBfWhRG3GnHhx4upt1yvp6wr]
- 64. Multimodal machine learning for materials science [https://arxiv.org/abs/2309.04478]
- 65. Deep Multimodal Learning with Missing Modality: A Survey [https://arxiv.org/html/2409.07825v3]
- 66. Incomplete Data: What You Don't Know Might Hurt You - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3153574/]
- 67. Strategies for Handling Missing Values in Data Analysis [https://www.dasca.org/world-of-data-science/article/strategies-for-handling-missing-values-in-data-analysis]
- 68. Extracting accurate materials data from research papers ... [https://www.nature.com/articles/s41467-024-45914-8]
- 69. How Well Do Large Language Models Understand Tables ... [https://link.springer.com/article/10.1007/s40192-024-00362-6]
- 70. Scientific Data Visualization Software [https://www.collaborativedrug.com/scientific-data-visualization-software]
- 71. Color Palette Selection - Environmental Informatics [https://ag.purdue.edu/hydrology/wiki/environmental-informatics/presentation-graphics-resources/color-palette-selection/]
- 72. Multi-source heterogeneous data fusion and intelligent ... [https://www.nature.com/articles/s41598-025-22752-2]
- 73. Multi-Institutional Mammogram Dataset [https://www.emergentmind.com/topics/multi-institutional-mammogram-dataset]
- 74. Optimizing Data Lakes for Cost and Performance across ... [https://medium.com/@jacob.sabell/optimizing-data-lakes-for-cost-and-performance-across-aws-azure-and-google-cloud-fbc97206d588]
- 75. Best Practices for Structuring Large-Scale Datasets ... [http://www.zigpoll.com/content/what-are-the-best-practices-for-structuring-a-largescale-dataset-to-optimize-both-computational-efficiency-and-ease-of-exploratory-data-analysis]
- 76. AlphaMat: a material informatics hub connecting data ... [https://www.nature.com/articles/s41524-023-01086-5]
- 77. Multimodal Fusion on Low-quality Data: A Comprehensive ... [https://arxiv.org/html/2404.18947v2]
- 78. An integrated oversampling and noise reduction method ... [https://www.sciencedirect.com/science/article/pii/S2772662225000682]
- 79. Top 6 Visualizations for Quantitative Data Analysis Methods [https://chartexpo.com/blog/quantitative-data-analysis-methods]
- 80. Simplifying Multimodal Clinical Research Data Management [https://pmc.ncbi.nlm.nih.gov/articles/PMC10972680/]
- 81. Multimodal data curation via interoperability: use cases ... [https://www.nature.com/articles/s41597-025-05678-2]
- 82. Analyzing and Visualizing Data - Data Literacy for ... [https://guides.library.ucla.edu/c.php?g=1234052&p=9127691]
- 83. Scoping Review of Wearable Sensor Data in Cancer Care [https://pmc.ncbi.nlm.nih.gov/articles/PMC11470224/]
- 84. Quantitative Data Analysis. A Complete Guide [2025] [https://www.6sigma.us/six-sigma-in-focus/quantitative-data-analysis/]
- 85. A benchmark dataset for class-wise segmentation of ... [https://www.nature.com/articles/s41597-025-05243-x]
- 86. Dataset and benchmark for as-built BIM reconstruction from ... [https://www.sciencedirect.com/science/article/abs/pii/S0926580525001360]
- 87. CEQuest: Benchmarking Large Language Models ... [https://arxiv.org/html/2508.16081v1]
- 88. Evaluating and Advancing Multimodal Large Language ... [https://arxiv.org/html/2411.14725v2]
- 89. Gen AI Evaluation Service — Multimodal Metrics [https://medium.com/google-cloud/gen-ai-evaluation-service-multimodal-metrics-2a7715dbfad6]
- 90. Datasets and Benchmarks Track: Call for Papers - KDD 2025 [https://kdd2025.kdd.org/call-for-datasets-and-benchmarks-track-papers/]
- 91. Using Large Multimodal Models to Extract Knowledge ... [https://educationaldatamining.org/EDM2025/proceedings/2025.EDM.long-papers.170/index.html]
- 92. Empowering multimodal analysis with visualization: A survey [https://www.sciencedirect.com/science/article/abs/pii/S1574013725000243]
- 93. Multimodal AI: Transforming Evaluation & Monitoring [https://galileo.ai/blog/multimodal-ai-guide]
- 94. Ultimate Guide - The Best Multimodal AI Models in 2025 [https://www.siliconflow.com/articles/en/best-multimodal-ai-models]
- 95. 7 Best Multimodal AI Model Platforms Tested for Performance [https://www.index.dev/blog/multimodal-ai-models-comparison]
- 96. MMTBench: A Unified Benchmark for Complex Multimodal ... [https://arxiv.org/html/2505.21771v1]
- 97. MatQnA: A Benchmark Dataset for Multi-modal Large ... [https://arxiv.org/abs/2509.11335]
- 98. MatQnA: Multi-Modal Benchmark Dataset [https://www.emergentmind.com/topics/matqna-dataset]
- 99. [Literature Review] MatQnA: A Benchmark Dataset for Multi ... [https://www.themoonlight.io/en/review/matqna-a-benchmark-dataset-for-multi-modal-large-language-models-in-materials-characterization-and-analysis]
- 100. Implementation Strategies to Improve Engagement With a ... [https://www.jmir.org/2021/10/e28924/]
- 101. R1-Onevision: Advancing Generalized Multimodal ... [https://arxiv.org/abs/2503.10615]
- 102. Doc-Researcher: A Unified System for Multimodal ... [https://arxiv.org/html/2510.21603v1]
- 103. PA-FAS: Towards Interpretable and Generalizable ... [https://arxiv.org/html/2511.17927v1]