Bridging Quantum and Data: Machine Learning Revolutionizes Density Functional Theory for Biomedical Innovation
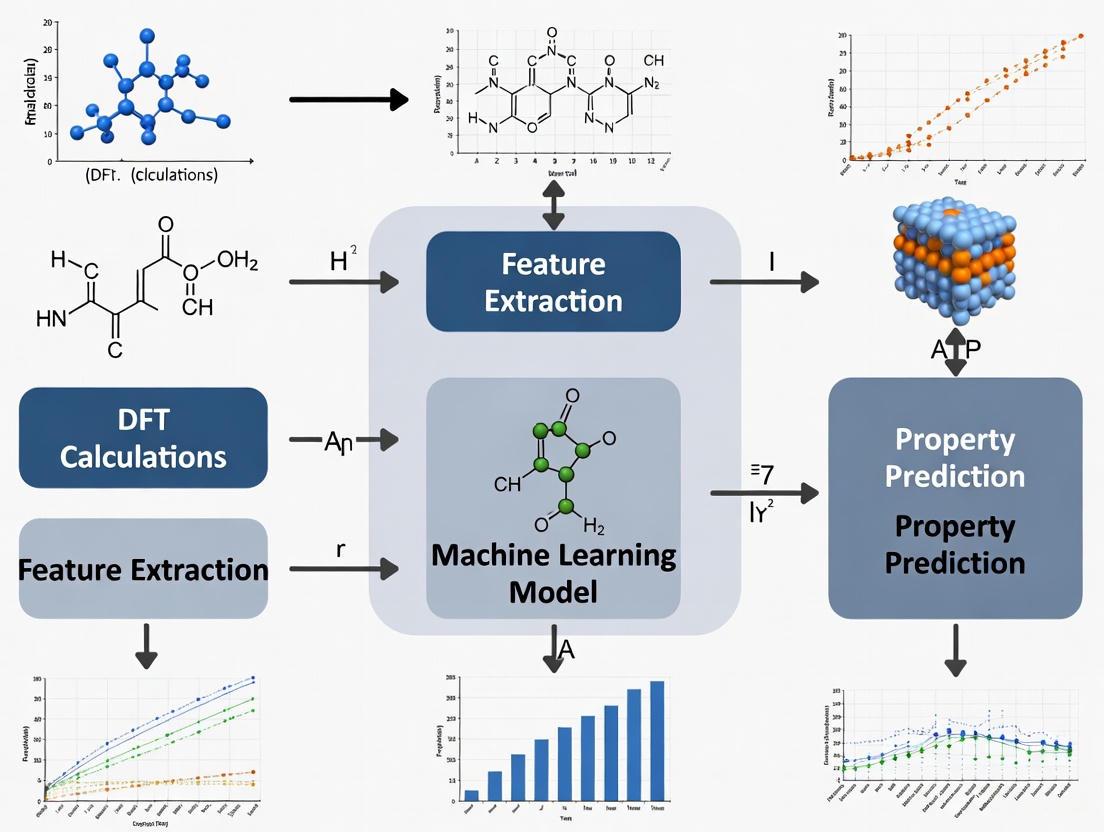
This article explores the transformative integration of Machine Learning (ML) with Density Functional Theory (DFT), a pivotal shift in computational science for biomedical and materials research.
Bridging Quantum and Data: Machine Learning Revolutionizes Density Functional Theory for Biomedical Innovation
Abstract
This article explores the transformative integration of Machine Learning (ML) with Density Functional Theory (DFT), a pivotal shift in computational science for biomedical and materials research. It covers the foundational principles of using ML to bypass the computational bottlenecks of the Kohn-Sham equations, detailed methodologies for emulating electronic properties and learning exchange-correlation functionals, strategies for troubleshooting transferability and data quality, and rigorous validation against high-accuracy benchmarks. Aimed at researchers and drug development professionals, this review synthesizes how these ML-accelerated workflows are enabling the rapid, accurate discovery of novel materials and therapeutic compounds, fundamentally reshaping predictive modeling in the life sciences.
The Quantum-Mechanical Bridge: How ML is Redefining the Foundations of DFT
Density Functional Theory (DFT) has established itself as a cornerstone of computational materials science and drug discovery, enabling the study of electronic structures from first principles. At its core lies the Kohn-Sham (KS) equation, which transforms the intractable many-electron problem into an effective single-electron problem. While this reformulation makes calculations feasible, the computational process of solving these equations—typically through an iterative self-consistent field (SCF) procedure—creates a fundamental bottleneck. This "Kohn-Sham bottleneck" manifests as the high computational cost required to: (1) determine the KS wavefunctions, (2) construct the associated electron density, and (3) solve for the eigenvalues that describe the system's electronic states. The challenge escalates dramatically with system size and complexity, limiting the practical application of DFT to large molecular systems or long-time-scale molecular dynamics simulations relevant to drug development.
Machine learning (ML) presents a paradigm shift for overcoming this bottleneck. By learning the complex mappings between atomic structure and electronic properties directly from data, ML models can emulate key parts of the DFT workflow, bypassing the need for computationally expensive iterative solutions. This article details the latest ML methodologies and protocols designed to overcome the KS bottleneck, enabling accurate and efficient electronic structure calculations for research and development.
Machine Learning Approaches to the Kohn-Sham Challenge
Several distinct ML strategies have emerged to address different aspects of the KS bottleneck. The table below summarizes the primary approaches, their specific targets, and their performance.
Table 1: Machine Learning Approaches for Overcoming the Kohn-Sham Bottleneck
| ML Approach | Computational Target | Key Innovation | Reported Performance & System |
|---|---|---|---|
| Unsupervised Representation Learning [1] | KS Wavefunctions | Uses a Variational Autoencoder (VAE) to compress high-dimensional KS wavefunctions into a low-dimensional latent space (${10}^{3}-{10}^{4}$ times smaller) [1]. | MAE of 0.11 eV for GW quasiparticle energies of 2D metals/semiconductors [1]. |
| End-to-End DFT Emulation [2] | Entire DFT Workflow | Maps atomic structure directly to electron density, then predicts energies, forces, and other properties, bypassing the explicit KS solution [2]. | Chemical accuracy achieved for organic molecules and polymers; orders of magnitude speedup [2]. |
| Learned Exchange-Correlation (XC) Functional [3] [4] | XC Functional | Employs deep learning to create a non-local XC functional (e.g., Skala) trained on high-accuracy quantum data [3]. | Reaches chemical accuracy (<1 kcal/mol) for atomization energies at semi-local DFT cost [3]. |
| On-the-Fly Machine-Learned Force Fields (MLFF) [5] | Forces and Energies for MD | Uses a Gaussian Multipole (GMP) descriptor for efficient, element-count-independent force field learning during molecular dynamics [5]. | Stable, >20 ps MD simulations for multi-element alloys (up to 6 elements) [5]. |
Detailed Experimental Protocols
This section provides detailed methodologies for implementing the key ML approaches described above, providing a practical guide for researchers.
Protocol: Unsupervised Learning of Kohn-Sham States with a VAE
This protocol outlines the procedure for compressing KS wavefunctions using a Variational Autoencoder (VAE) as described in Nature Communications (2024) [1]. The primary goal is to learn a low-dimensional representation that retains the essential physical information of the original, high-dimensional wavefunctions.
Primary Research Application: Creating a compressed, generative representation of electronic structure for use in downstream tasks, such as predicting quasiparticle band structures within the GW formalism.
Materials/Software Requirements:
- Input Data: Kohn-Sham wavefunction moduli, ( {|}{\phi}{n{{\bf{k}}}}\left({{\bf{r}}}\right){|} ), represented on a real-space grid of dimensions ( Rx \times Ry \times Rz ), indexed by band (( n )) and k-point (( {{\bf{k}} )).
- Computational Framework: A deep learning environment (e.g., Python with TensorFlow or PyTorch) capable of building and training convolutional neural networks (CNNs) and VAEs.
- Reference Data: For downstream supervised learning (e.g., GW bandstructures), corresponding quasiparticle energies are required.
Step-by-Step Procedure:
- Data Preparation: Collect the KS wavefunctions from a set of DFT calculations for a range of materials of interest. The wavefunctions should be formatted as 2D or 3D arrays (depending on the system dimensionality).
- VAE Architecture Definition:
- Encoder (( e{\theta} )): Design a network that maps the input wavefunction to a latent distribution.
- Layers 1-2: Implement 2D/3D Convolutional Neural Networks (CNNs) to capture local spatial patterns in the wavefunction.
- Symmetry Enforcement: Use circular padding in CNNs to respect the periodic boundary conditions of the crystal [1].
- Invariance Layer: Insert a Global Average Pooling (GAP) layer after the CNNs to enforce translational invariance with respect to the unit cell's origin [1].
- Output: The encoder outputs two vectors in the latent space: the mean (( {{\mathbf{\mu}}} \in {{\mathbb{R}}}^{p} )) and the logarithm of the variance (( \log {{\mathbf{\sigma}}}^2 )), which define the posterior distribution.
- Decoder (( d{\theta} )): Mirror the encoder structure to reconstruct the input wavefunction from a latent vector sampled from the distribution defined by ( {{\mathbf{\mu}}} ) and ( {{\mathbf{\sigma}}} ).
- Encoder (( e{\theta} )): Design a network that maps the input wavefunction to a latent distribution.
- Loss Function and Training:
- Define the total loss function, ( {\mathscr{L}} ), as a weighted sum of a reconstruction loss and a regularization term:
$${{\mathscr{L}}}=\frac{1}{T}{\sum }_{n{{\bf{k}}}}^{T}{{|||}}{\phi }_{n{{\bf{k}}}}{{|}}-{d}_{\theta }({e}_{\theta }({{|}}{\phi }_{n{{\bf{k}}}}{{|}})){{|}}{{{|}}}^{2} + \beta \cdot D_{KL}(q(z | |{\phi }_{n{{\bf{k}}}})|) | N(0,{{\rm I}}))$$ - The first term is the Mean Squared Error (MSE) between the input and reconstructed wavefunction.
- The second term is the Kullback-Leibler (KL) Divergence, which forces the latent space distribution to approximate a standard normal distribution. The parameter ( \beta ) controls the strength of this regularization.
- Train the VAE using an optimizer (e.g., Adam) by minimizing ( {\mathscr{L}} ) over the training set of KS states.
- Define the total loss function, ( {\mathscr{L}} ), as a weighted sum of a reconstruction loss and a regularization term:
Downstream Application: The trained encoder can be used to convert new KS wavefunctions into their latent representations. These compact vectors can then serve as input to a separate, supervised neural network trained to predict properties like GW quasiparticle energies.
Troubleshooting Tips:
- If reconstruction quality is poor, consider increasing the dimensionality of the latent space (( p )) or adjusting the ( \beta ) parameter.
- To improve the physical smoothness of the latent space, ensure the training data includes KS states from closely spaced k-points.
Protocol: On-the-Fly Machine-Learned Force Field Molecular Dynamics
This protocol describes the implementation of an on-the-fly ML force field based on the Normalized Gaussian Multipole (GMP) descriptor, as published in Journal of Chemical Theory and Computation (2024) [5]. This method is particularly powerful for molecular dynamics (MD) simulations of systems with high chemical complexity.
Primary Research Application: Performing stable, long-time-scale MD simulations without the need for pre-training, automatically generating a force field that scales efficiently with the number of chemical elements.
Materials/Software Requirements:
Step-by-Step Procedure:
- Initialization: Start an MD simulation (e.g., in the NVT or NVK ensemble) using the DFT code. The on-the-fly MLFF will be inactive at this stage.
- Descriptor Calculation (GMP): For each new atomic configuration encountered during the MD trajectory, compute the GMP descriptor for every atom.
- The GMP descriptor represents the atomic environment using a fixed-length vector based on a Gaussian representation of atomic valence densities, independent of the number of chemical elements [5].
- Uncertainty Quantification & Active Learning:
- The ML model (e.g., Bayesian linear regression) uses the GMP features to predict atomic forces and energy and provides a Bayesian uncertainty estimate for its predictions.
- Decision Point: For each MD step, compare the model's uncertainty to a pre-defined threshold.
- Low Uncertainty: Use the MLFF-predicted forces to propagate the dynamics. This is the "fast" path that bypasses the KS bottleneck.
- High Uncertainty: Trigger a full DFT calculation on the current atomic configuration. This new data point is added to the training set, and the ML model is updated. The DFT-calculated forces are used for the MD step.
- Model Update: Retrain the regression model on the updated training set after incorporating new data from high-uncertainty steps.
Validation and Analysis:
- Stability Check: Monitor the total variation distance (TVD) of the pair correlation functions (PCFs) between the MLFF-MD and a reference AIMD trajectory to ensure the model's stability and accuracy [5].
- Property Calculation: Once a stable trajectory is obtained, compute the desired thermodynamic or dynamic properties (e.g., diffusion coefficients, free energies).
Troubleshooting Tips:
- If the number of DFT calls does not decrease over time, the uncertainty threshold may be set too low, or the system may be exploring many novel configurations.
- For systems with severe element segregation, the model may require more data to learn all the unique chemical interactions.
The logical workflow and decision points for this on-the-fly protocol are summarized in the diagram below.
On-the-Fly MLFF Workflow: This diagram illustrates the decision-making process during an on-the-fly machine-learned force field molecular dynamics simulation, showing how the model selectively invokes DFT calculations based on uncertainty.
The Scientist's Toolkit: Research Reagent Solutions
The following table lists key computational "reagents"—software, descriptors, and datasets—essential for implementing the ML-driven DFT workflows discussed in this article.
Table 2: Essential "Research Reagents" for Machine Learning-Enhanced DFT
| Reagent Name/Type | Function/Purpose | Key Features / Relevance to KS Bottleneck |
|---|---|---|
| Variational Autoencoder (VAE) [1] | Compresses high-dimensional KS wavefunctions into a low-dimensional, smooth latent space. | Enables generative modeling of electronic structure; representation is ${10}^{3}-{10}^{4}$ times smaller than input [1]. |
| Gaussian Multipole (GMP) Descriptor [5] | Describes an atom's chemical environment for force prediction. | Fixed-length descriptor; scales efficiently with number of chemical elements, unlike SOAP [5]. |
| Skala Functional [3] | A deep learning-based exchange-correlation (XC) functional. | Learns non-local representations; targets chemical accuracy at the computational cost of semi-local DFT [3]. |
| AGNI Fingerprints [2] | Atomic descriptors representing the structural and chemical environment. | Translation, permutation, and rotation invariant; used as input for predicting charge density and other properties [2]. |
| MSR-ACC/TAE25 Dataset [3] | A high-accuracy dataset of total atomization energies. | Used for training and validating ML-based XC functionals like Skala on chemically accurate data [3]. |
| SPARC / VASP / CASTEP [2] [5] | DFT software packages. | Provide the electronic structure engine for generating reference data and are often the platform for integrating on-the-fly MLFFs [2] [5]. |
The Kohn-Sham bottleneck, long a fundamental constraint in computational chemistry and materials physics, is now being decisively addressed by a new generation of machine learning workflows. The protocols detailed herein—ranging from unsupervised wavefunction learning and end-to-end DFT emulation to robust on-the-fly force fields—demonstrate that it is possible to achieve the accuracy of high-level electronic structure theory at a fraction of the computational cost. For researchers and drug development professionals, these tools unlock new possibilities: screening vast libraries of molecular candidates with chemical accuracy, simulating complex biological processes at atomic resolution, and exploring reaction mechanisms that were previously beyond computational reach. As these ML-driven methodologies continue to mature and integrate, they promise to make predictive, first-principles modeling a routine tool in the quest for new drugs and advanced materials.
Density Functional Theory (DFT) stands as a cornerstone of modern computational chemistry and materials science, enabling the prediction of electronic structure and properties from first principles. The fundamental theorem of DFT states that all ground-state properties of a many-electron system are uniquely determined by its electron density. However, the practical accuracy of DFT calculations hinges on the exchange-correlation (XC) functional, which accounts for quantum mechanical effects not captured in simple electrostatic models. The pursuit of a "universal functional" that delivers chemical accuracy across diverse systems and elements represents a grand challenge in the field.
Traditional approaches to developing XC functionals have followed a heuristics-based paradigm, systematically climbing "Jacob's ladder" by incorporating increasingly complex physical ingredients—from local density (LDA) to generalized gradient approximations (GGA) and hybrid functionals. While this progression has yielded significant improvements, each rung on the ladder introduces greater computational cost without guaranteeing proportional gains in accuracy. Moreover, these functionals still struggle with quantitative prediction of formation enthalpies, band gaps, and reaction barriers, limiting their predictive power for materials discovery and drug development.
The integration of Machine Learning (ML) with DFT has emerged as a transformative pathway to transcend these limitations. By leveraging data-driven approaches, researchers can now develop more accurate and efficient approximations to the universal functional, create machine learning interatomic potentials (MLIPs) that preserve quantum accuracy at reduced computational cost, and establish robust structure-property relationships for accelerated materials design. This Application Note details the protocols and methodologies underpinning these advances, providing researchers with practical frameworks for implementation.
ML-Enhanced XC Functionals: Protocols and Performance
Core Methodology: Energy and Potential Learning
Recent breakthroughs in ML-accelerated DFT have demonstrated that incorporating both energies and quantum potentials during training enables the development of more accurate and transferable XC functionals. This approach, pioneered by Gavini and colleagues, leverages the fact that potentials provide a stronger training foundation than energies alone, as they more sensitively capture subtle electronic variations across chemical systems [4].
Experimental Protocol: ML-XC Functional Development
Data Acquisition: Obtain high-quality quantum many-body (QMB) reference data for small systems (atoms, diatomic molecules) where exact solutions are computationally feasible. The training set should include:
- Total energies from coupled-cluster or quantum Monte Carlo calculations
- Electronic potentials and their spatial variations
- Systems with diverse bonding characteristics (metallic, covalent, ionic)
Feature Engineering: Represent the electron density and its gradients using:
- Local density approximations at grid points
- Density gradient components (∇ρ)
- Kinetic energy density (τ)
- Hartree-Fock exchange fractions for hybrid functionals
Model Architecture: Implement a multi-layer perceptron (MLP) or deep neural network with:
- 3-5 hidden layers with 50-200 neurons per layer
- Swish or ReLU activation functions
- Residual connections to facilitate training depth
- Physical constraints to ensure functional derivatives satisfy sum rules
Training Protocol:
- Use leave-one-out cross-validation to assess transferability
- Apply k-fold cross-validation (k=5-10) to prevent overfitting
- Utilize physically-informed regularization to maintain functional convexity
- Optimize with adaptive moment estimation (Adam) or L-BFGS algorithms
Validation: Benchmark against experimental data for:
- Formation enthalpies of binary and ternary alloys
- Band gaps of semiconductors and insulators
- Reaction energies and barrier heights
- Structural parameters (bond lengths, angles)
Performance Metrics and Comparative Analysis
Table 1: Performance Comparison of Traditional vs. ML-Enhanced DFT Approaches for Formation Enthalpy Prediction
| Method | Training Data | MAE (eV/atom) | Computational Cost | Transferability |
|---|---|---|---|---|
| LDA | N/A | 0.15-0.25 | Low | Moderate |
| GGA (PBE) | N/A | 0.10-0.20 | Low | Moderate |
| Hybrid (HSE06) | N/A | 0.05-0.15 | High | Good |
| ML-XC (Linear Correction) | 50-100 binary alloys | 0.08-0.12 | Low | Limited |
| ML-XC (Neural Network) | 100-200 binary/ternary alloys | 0.03-0.06 | Medium | Good |
| ML-XC (Potential-Enhanced) | 5-10 atoms + simple molecules | 0.02-0.04 | Medium | Excellent |
The ML-enhanced approach demonstrates particular strength in predicting formation enthalpies for ternary systems like Al-Ni-Pd and Al-Ni-Ti, which are crucial for high-temperature applications in aerospace and protective coatings [6]. By learning the systematic errors of traditional DFT, the ML-corrected functionals achieve accuracy接近ing high-level quantum chemistry methods at a fraction of the computational cost.
Machine Learning Interatomic Potentials: Bridging Accuracy and Efficiency
Development Workflow for General NNPs
Machine Learning Interatomic Potentials (MLIPs) represent a powerful alternative to traditional force fields, offering DFT-level accuracy for molecular dynamics simulations of large systems and extended timescales. The EMFF-2025 potential for C, H, N, O-based high-energy materials exemplifies this approach, achieving accurate predictions of structures, mechanical properties, and decomposition characteristics across 20 different molecular systems [7].
Experimental Protocol: MLIP Development via Transfer Learning
Base Model Preparation:
- Start with a pre-trained neural network potential (e.g., Deep Potential framework)
- Ensure the base model captures diverse bonding environments and elemental interactions
- The DP-CHNO-2024 model serves as an effective foundation for energetic materials
Target System Data Generation:
- Perform targeted DFT calculations on new molecular systems not in the original training set
- Sample diverse configurations including:
- Equilibrium crystal structures
- Perturbed geometries (±0.05-0.1 Å atomic displacements)
- Reaction pathways with bond breaking/forming
- High-temperature configurations from ab initio MD
Transfer Learning Implementation:
- Freeze early layers of the neural network preserving general chemical knowledge
- Retrain final layers with new system-specific data (typically 100-500 configurations)
- Use a reduced learning rate (10-50% of original) to fine-tune parameters
- Employ a combined loss function balancing energy and force accuracy
Validation and Deployment:
- Validate against held-out DFT calculations for energies (target MAE < 0.1 eV/atom) and forces (target MAE < 2 eV/Å)
- Test transferability to related chemical systems not included in training
- Deploy for large-scale molecular dynamics simulations (10,000-1,000,000 atoms)
Performance Benchmarking
The EMFF-2025 framework demonstrates that transfer learning with minimal additional data enables the development of highly accurate potentials. Quantitative assessment shows mean absolute errors predominantly within ±0.1 eV/atom for energies and ±2 eV/Å for forces across a wide temperature range [7]. This accuracy permits reliable investigation of thermal decomposition mechanisms and mechanical properties previously inaccessible through conventional force fields or direct DFT-MD.
Descriptor Strategies for Electrocatalyst Design
Descriptor Classification and Selection Guidelines
In ML-accelerated electrocatalyst discovery, descriptors serve as quantitative representations of material features that determine catalytic performance. Three fundamental descriptor classes enable efficient structure-property mapping across different phases of the discovery pipeline [8].
Table 2: Electrocatalysis Descriptor Classes and Their Applications
| Descriptor Class | Key Examples | Data Requirements | Computational Cost | Primary Use Cases |
|---|---|---|---|---|
| Intrinsic Statistical | Magpie features (132 elemental attributes), atomic number, valence electron count | Low (elemental data only) | Very Low | High-throughput initial screening, binary classification (active/inactive) |
| Electronic Structure | d-band center, orbital occupation, spin magnetic moment, Bader charges | Medium (requires DFT) | Medium | Mechanism interpretation, activity trend analysis, fine screening |
| Geometric/Microenvironment | Coordination number, interatomic distances, local strain, site symmetry | High (requires optimized structures) | High | Complex environments (alloys, SACs, DACs), support effect quantification |
Experimental Protocol: Hierarchical Descriptor Implementation
Phase 1: Initial Screening with Intrinsic Descriptors
- Compute 132 Magpie features for candidate elements
- Apply feature selection (variance threshold, correlation analysis)
- Train gradient boosting regressor (GBR) or random forest (RF) models
- Screen 10,000+ compositions in coarse-filter approach
Phase 2: Electronic Descriptor Analysis
- Perform DFT optimization on top 5-10% candidates from Phase 1
- Calculate electronic descriptors: d-band center (εd), projected density of states (PDOS)
- Develop customized composite descriptors (e.g., ARSC for dual-atom catalysts)
- Apply recursive feature elimination to identify minimal predictive descriptor set
Phase 3: Microenvironment Refinement
- For most promising candidates, analyze local coordination environments
- Quantify metal-support interactions, strain effects, solvation models
- Validate descriptor transferability across material classes
Customized Composite Descriptors: The ARSC Framework
For complex catalytic systems like dual-atom catalysts (DACs), customized composite descriptors integrate multiple physical effects into compact, interpretable expressions. The ARSC descriptor framework exemplifies this approach, combining:
- Atomic property effects via d-band shape parameters (ϕxx)
- Reactant-based screening for heteronuclear systems (ϕopt)
- Synergistic effects through physics-guided feature engineering (ϕxy)
- Coordination effects with experimental verification (Φ)
This methodology achieved accuracy comparable to ~50,000 DFT calculations while training on fewer than 4,500 data points, dramatically accelerating the exploration of 840 transition metal DACs for ORR, OER, CO2RR, and NRR [8].
Research Reagent Solutions: Essential Computational Tools
Table 3: Key Software and Methodological "Reagents" for ML-DFT Workflows
| Research Reagent | Type | Primary Function | Application Context |
|---|---|---|---|
| Deep Potential (DP) | MLIP Framework | Generates neural network potentials from DFT data | Large-scale MD with quantum accuracy [7] |
| DP-GEN | Automated Workflow | Implements active learning for MLIP development | Adaptive sampling of configuration space [7] |
| EMTO-CPA | DFT Code | Exact muffin-tin orbital method with coherent potential approximation | Alloy formation enthalpy calculations [6] |
| Skala Functional | ML-XC Functional | Deep-learning-powered exchange-correlation functional | High-accuracy DFT without Jacob's ladder trade-offs [9] |
| ARSC Descriptor | Composite Descriptor | Encodes atomic, reactant, synergistic, and coordination effects | Rapid screening of dual-atom catalysts [8] |
| TDDFT-GPU | GPU Implementation | Time-dependent DFT on massively parallel GPUs | Excited-state calculations for large systems [10] |
Integrated Workflow for Materials Discovery
The most impactful applications of ML-DFT integration combine multiple methodologies into cohesive discovery pipelines. The following workflow exemplifies this integrated approach, synthesizing elements from across the protocols detailed in this document.
This integrated workflow enables researchers to efficiently navigate vast chemical spaces, from initial screening of thousands of candidates to detailed investigation of selected leads with DFT-level accuracy at molecular dynamics scale. The continuous feedback loop ensures iterative improvement of both models and descriptors, accelerating the discovery cycle for advanced materials targeting specific application requirements.
In the framework of density functional theory (DFT), the electron density, denoted as ρ(r), is the fundamental variable that uniquely determines all ground-state properties of an interacting electron system, as established by the Hohenberg-Kohn theorems [11] [12]. This foundational principle enables the replacement of the complex many-body wavefunction with the electron density as the central quantity of interest, significantly simplifying computational approaches. The Kohn-Sham equations transform this theoretical framework into a practical tool by mapping the system of interacting electrons onto a fictitious system of non-interacting electrons moving within an effective potential [13]. This effective potential comprises the external potential (from atomic nuclei), the Hartree potential (electron-electron repulsion), and the exchange-correlation potential, which encapsulates all many-body effects not captured by the other terms [11].
The accuracy of DFT calculations critically depends on the approximations used for the exchange-correlation functional, which remains unknown in its exact form [13]. The hierarchy of approximations ranges from the Local Density Approximation (LDA), which depends only on the local electron density, to Generalized Gradient Approximations (GGA) that incorporate density gradients, meta-GGAs that additionally include the kinetic energy density, and hybrid functionals that mix a portion of exact Hartree-Fock exchange with DFT exchange [12] [13]. The pursuit of more accurate functionals represents an active research frontier, directly impacting the reliability of predicted material properties, reaction mechanisms, and electronic behaviors in computational materials science and drug development [14] [15].
Table 1: Hierarchy of Exchange-Correlation Functionals in DFT
| Functional Type | Dependence | Key Characteristics | Example Functionals |
|---|---|---|---|
| Local Density Approximation (LDA) | Local electron density ρ(r) | Simple, efficient, often over-binds | SVWN [13] |
| Generalized Gradient Approximation (GGA) | ρ(r), ∇ρ(r) | Improved molecular geometries & energies | PBE, BLYP [13] |
| meta-GGA | ρ(r), ∇ρ(r), kinetic energy density | Better for properties sensitive orbital shapes | SCAN [12] |
| Hybrid | ρ(r), ∇ρ(r), exact exchange | Mixes DFT & Hartree-Fock exchange | B3LYP, PBE0 [12] [13] |
Machine Learning Charge Density Prediction with Δ-SAED Protocol
Background and Principle
The accurate prediction of electron density is paramount in DFT, as it forms the basis for calculating all other ground-state properties. Traditional machine learning approaches for charge density prediction have targeted the total charge density (TCD) directly [16]. However, the Δ-SAED method introduces a paradigm shift by leveraging physical prior knowledge. Instead of learning the total charge density from scratch, Δ-SAED learns the difference charge density (DCD), defined as the difference between the TCD and the superposition of atomic electron densities (SAED) [16]. This approach effectively incorporates the physical ansatz that the electron density of a molecular or solid-state system can be reasonably initialized as a sum of isolated atomic densities, with the machine learning model capturing the complex redistribution due to chemical bonding.
This Δ-learning strategy has demonstrated robust improvements in prediction accuracy across diverse benchmark datasets, including organic molecules (QM9), battery cathode materials (NMC), and inorganic crystals (Materials Project) [16]. By reducing the complexity of the function that the neural network must approximate, Δ-SAED enhances data efficiency and model transferability, which is particularly valuable when training data is limited or when applying models to unseen chemical spaces in high-throughput screening applications.
Detailed Δ-SAED Protocol
Objective: To accurately predict the ground-state electron density of a molecular or solid-state system using a machine learning model trained on difference charge density.
Workflow Overview: The diagram below illustrates the integrated machine learning and DFT workflow for the Δ-SAED protocol:
Step-by-Step Procedures:
Reference Data Generation via DFT:
- Perform DFT calculations using established codes (Quantum ESPRESSO, VASP, CP2K) for a diverse set of atomic structures relevant to your research domain [17].
- Critical Step: During the calculation setup, ensure the output includes not only the final self-consistent total charge density (ρ(t)) but also the initial superposition of atomic electron densities (ρ(a)) used to start the calculation.
- Compute the difference charge density (DCD, ρ(d)) for each structure according to: ρ(d)(r) = ρ(t)(r) - ρ(a)(r). This ρ(d) becomes the target for machine learning training [16].
Model Training:
- Architecture Selection: Employ an E(3)-equivariant neural network architecture, such as Charge3Net, which incorporates high-order messages to accurately capture complex chemical environments [16]. Grid-based methods are recommended over basis-based methods for their higher accuracy, despite increased computational cost [16].
- Training Target: Optimize model parameters by minimizing the loss function between predicted DCD (ρ̂(d)) and true DCD (ρ(d)) from DFT. The mean absolute error (MAE) metric is commonly used [16]:
ε_mae = [∫_Ω d³r |ρ(r) - ρ̂(r)|] / [∫_Ω d³r ρ_t(r)] × 100%
Prediction and Reconstruction:
- For a new atomic structure, first compute its SAED (ρ(a)) using the same atomic densities as in training.
- Pass the structure through the trained model to obtain the predicted DCD (ρ̂(d)).
- Reconstruct the predicted total charge density by adding the SAED to the predicted difference: ρ̂(t)(r) = ρ(a)(r) + ρ̂(d)(r) [16].
Validation and Quality Control:
- Benchmark the predicted charge densities against held-out DFT calculations.
- For non-self-consistent (single-shot) DFT calculations using the ML-predicted density, verify that derived properties (e.g., formation energies, band structures) achieve chemical accuracy compared to fully self-consistent DFT results [16].
- Monitor the radial distribution of prediction errors to ensure the model correctly captures both short-range (near nuclei) and long-range (bonding regions) electronic interactions.
Table 2: Performance of Δ-SAED vs Traditional TCD Learning on Benchmark Datasets
| Dataset | System Type | MAE Reduction with Δ-SAED | Structures with Improved Accuracy |
|---|---|---|---|
| QM9 | Organic Molecules | Significant reduction [16] | >99% [16] |
| NMC | Battery Cathode Materials | Significant reduction [16] | >99% [16] |
| Materials Project | Inorganic Crystals | Significant reduction [16] | ~90% [16] |
| Si Allotropy | Silicon Polymorphs | Enables transferability to chemical accuracy for derived properties [16] | Nearly 100% for non-self-consistent calculations [16] |
Automated DFT Workflows for High-Throughput Screening
Workflow Architecture and Design Principles
Automated DFT workflows are computational frameworks designed to manage, execute, and document high volumes of DFT calculations with minimal manual intervention [17]. These workflows are essential for robust and reproducible computational research, particularly in machine learning where large, consistent datasets are required. The architecture is typically layered and modular, often implemented in Python and built on workflow engines like AiiDA, JARVIS-Tools, or pyiron [17]. Key design principles include engine-agnostic interfaces (compatible with multiple DFT codes like VASP, Quantum ESPRESSO, CP2K), protocol-driven calculations (using standardized "fast," "moderate," and "precise" settings), and comprehensive provenance tracking to ensure full reproducibility [17].
A representative automated workflow for high-throughput screening might encompass structure generation, parameter convergence, job submission, error handling, and post-processing analysis. The diagram below illustrates such a workflow:
Protocol for High-Throughput Screening
Objective: To systematically screen a large number of materials structures for target properties (e.g., band gaps, adsorption energies, formation energies) using automated DFT workflows.
Step-by-Step Procedures:
Structure Generation and Input Preparation:
- Structure Sources: Import structures from databases (Materials Project, COD) or generate them using algorithms for defect enumeration, surface slab creation, or substitutional doping [17].
- Input Parameter Convergence: Automatically converge critical parameters including k-point mesh density and plane-wave cutoff energy. The workflow should iteratively refine these parameters until threshold changes in total energy (e.g., < 1 meV/atom) are achieved [17].
- Standardization: Utilize standardized input objects (e.g., ASE Atoms class, OPTIMADE format) to ensure consistency across different DFT codes [17].
Job Execution and Error Handling:
- HPC Integration: Interface natively with HPC schedulers (SLURM, PBS) for job submission, monitoring, and resubmission [17].
- Robust Error Handling: Implement automated handlers for common DFT errors:
- SCF non-convergence: Increase electronic step limits, switch mixing algorithms (e.g., from Pulay to RMM-DIIS), or modify mixing parameters [17].
- Geometry optimizer stalls: Change optimization algorithms (e.g., from BFGS to FIRE) or adjust step sizes [17].
- Wall-time failures: Implement automatic checkpointing and restarts [17].
Post-processing and Property Extraction:
- Property Computation: Automatically calculate target properties such as band structures, density of states (DOS), elastic constants, Bader charges, and formation energies [17].
- Thermodynamic Analysis: Perform energy-volume equation of state fitting (e.g., using Birch-Murnaghan formalism) and compute thermodynamic properties under the quasi-harmonic approximation (QHA) if needed [17].
- Validation: Implement automated validation checks comparing calculated properties to experimental data or high-fidelity references where available [17].
Data Management and Provenance:
- Structured Storage: Deposit all inputs, outputs, and calculated properties into structured databases (SQL, NoSQL, HDF5) with comprehensive metadata indexing [17].
- Provenance Tracking: Ensure full reproducibility by recording a complete provenance graph of all calculations, including code versions, parameters, and intermediate results [17].
Table 3: Key Computational Tools and Resources for DFT-ML Research
| Tool Category | Specific Examples | Function and Application |
|---|---|---|
| DFT Codes | VASP, Quantum ESPRESSO, CP2K, CASTEP | Perform the core quantum mechanical calculations to generate reference electronic structure data and properties [17]. |
| Workflow Managers | AiiDA, pyiron, JARVIS-Tools | Automate calculation workflows, ensure reproducibility, and manage computational provenance [17]. |
| ML Charge Density Models | Charge3Net, Δ-SAED method | Predict accurate electron densities for structures, enabling rapid non-self-consistent property calculations [16]. |
| Exchange-Correlation Functionals | PBE (GGA), SCAN (meta-GGA), B3LYP (Hybrid) | Define the approximation for the exchange-correlation energy, critically determining calculation accuracy [12] [13]. |
| Basis Sets | Plane Waves, Gaussian Basis Sets (cc-pVTZ, pc-n) | Represent the Kohn-Sham orbitals; choice affects convergence and accuracy, with large sets needed for accurate densities [12]. |
| Structure Manipulation | Pymatgen, ASE | Create, manipulate, and analyze atomic structures; crucial for input preparation and post-processing [17]. |
Applications in Nanomaterials and Drug Development Research
The integration of DFT with machine learning is particularly impactful in the field of nanomaterials research and drug development. ML-driven charge density models and automated workflows enable the rapid screening and design of novel nanomaterials with tailored electronic, catalytic, and optical properties [15]. Specific applications include predicting band gaps for optoelectronic materials, calculating adsorption energies for catalytic applications, and elucidating reaction mechanisms at nanomaterial surfaces [15].
In drug development contexts, DFT-based molecular dynamics (AIMD) simulations provide insights into drug-receptor interactions, solvation effects, and reaction pathways in complex biomolecular systems [14]. The combination of these accurate but expensive simulations with machine learning potentials has opened new possibilities for simulating larger systems and longer timescales, directly impacting rational drug design [14] [15]. Furthermore, the calculation of NMR and EPR parameters using relativistic DFT provides crucial spectroscopic information that can be directly compared with experimental results, aiding in compound characterization and verification [14].
Why Now? The Convergence of Big Quantum Data and Advanced Algorithms
The integration of quantum computing into machine learning workflows, particularly for domains like Density Functional Theory (DFT), is transitioning from theoretical exploration to practical application. This shift is driven by a critical convergence of three factors: the emergence of large-scale, high-quality quantum data from advanced hardware; significant algorithmic breakthroughs that leverage this data; and the maturation of the software and control systems needed to run these workflows efficiently and reliably. For researchers in drug development and materials science, this creates an unprecedented opportunity to tackle computational problems that have historically been intractable for classical computers alone, such as highly accurate molecular simulations. The following application notes and protocols detail the quantitative evidence, experimental methodologies, and essential tools enabling this transition.
Quantitative Landscape: Market and Performance Data
The following tables summarize key quantitative data that underscores the rapid advancement and commercial potential of quantum technologies in scientific domains.
Table 1: Quantum Technology Market Projections and Investment (2024-2035)
| Metric | 2024 Value | 2035 Projection | Key Context & Sources |
|---|---|---|---|
| Total Quantum Tech (QT) Market | Not Specified | Up to $97B [18] | Encompasses computing, sensing, and communication. |
| Quantum Computing Market | ~$4B [18] | $28B - $72B [18] | Captures the bulk of the QT revenue. |
| Quantum Sensing Market | Not Specified | $7B - $10B [18] | |
| Value in Life Sciences | Not Specified | $200B - $500B [19] | Specific to quantum computing in pharma R&D. |
| Annual QT Start-up Funding | ~$2.0B [18] | N/A | 50% increase from $1.3B in 2023 [18]. |
| Public QT Funding (Gov't) | $1.8B (announced) [18] | N/A | Japan announced a further $7.4B in 2025 [18]. |
Table 2: Documented Quantum Application Performance (2024-2025)
| Application Area | Organization | Quantum System Used | Reported Performance / Milestone |
|---|---|---|---|
| Financial Trading | HSBC | IBM Heron [20] | 34% improvement in bond trading predictions vs. classical alone [20]. |
| Engineering Simulation | Ansys | IonQ [20] | 12% speedup in fluid interaction analysis for medical devices [20]. |
| Production Logistics | Ford Otosan | D-Wave [20] | Reduced scheduling times from 30 minutes to under 5 minutes; deployed in production [20]. |
| Chemical Simulation | IBM & RIKEN | IBM Heron + Fugaku Supercomputer [20] | Simulated molecules "beyond the ability of classical computers alone" at utility scale [20]. |
| Computer Calibration | Quantum Machines | QUAlibrate Framework [21] | Reduced calibration of superconducting qubits from hours to 140 seconds [21]. |
| Algorithm Speed | Google Quantum AI | DQI Algorithm (Theoretical) [22] | Certain optimization problems require ~million quantum ops vs. >10^23 classical ops [22]. |
Experimental Protocols for Quantum-Enhanced DFT and ML Workflows
This section provides detailed methodologies for key experiments and workflows that integrate quantum computing with machine learning for molecular simulation.
Protocol 1: Quantum-Accelerated Computational Chemistry Workflow
This protocol is adapted from industry collaborations, such as that between AstraZeneca, Amazon Web Services, and IonQ, to demonstrate a quantum-accelerated workflow for studying chemical reactions relevant to drug synthesis [19].
- Objective: To model the energy profile of a chemical reaction using a hybrid quantum-classical computational chemistry workflow.
Materials & Prerequisites:
- Classical Computing Resources: High-performance computing (HPC) cluster for pre- and post-processing.
- Quantum Computing Access: Cloud access to a quantum processing unit (QPU) or advanced simulator (e.g., via Amazon Braket, IBM Cloud).
- Software Stack: Python with libraries like Qiskit or Pennylane for quantum circuit definition, and classical computational chemistry software (e.g., PySCF).
- Target Molecule: A small molecule or reaction intermediate (e.g., for a Suzuki–Miyaura cross-coupling reaction).
Procedure:
- System Preparation (Classical):
- Define the molecular geometry of reactants and products.
- Use classical methods (e.g., Hartree-Fock) to generate an initial guess for the molecular Hamiltonian.
- Active Space Selection: Identify a subset of molecular orbitals and electrons most relevant to the chemical process (e.g., using a classical Complete Active Space SCF calculation).
- Hamiltonian Mapping (Classical):
- Transform the electronic Hamiltonian of the selected active space into a qubit representation using a fermion-to-qubit mapping (e.g., Jordan-Wigner or Bravyi-Kitaev).
- Quantum Circuit Execution (Hybrid):
- Algorithm Selection: Employ the Variational Quantum Eigensolver (VQE) algorithm.
- Ansatz Design: Construct a parameterized quantum circuit (ansatz) suitable for chemical systems, such as the Unitary Coupled Cluster (UCC) ansatz.
- Parameter Optimization: On the classical computer, use an optimizer (e.g., COBYLA, SPSA) to variationally minimize the energy expectation value. Each optimization step requires multiple executions of the parameterized quantum circuit on the QPU to estimate the energy.
- Result Analysis (Classical):
- Use the optimized parameters to compute the final, high-accuracy ground state energy.
- Compare the quantum-computed reaction energy profile with results from classical methods like Density Functional Theory (DFT) and experimental data.
- System Preparation (Classical):
Protocol 2: Quantum Machine Learning for Molecular Property Prediction
This protocol outlines a hybrid quantum-classical machine learning approach for predicting molecular properties, leveraging methodologies explored by companies like Merck KGaA and Amgen in collaboration with QuEra [19].
- Objective: To train a Quantum Neural Network (QNN) to predict the biological activity of a drug candidate based on its molecular descriptors.
Materials & Prerequisites:
- Dataset: A curated set of molecules with known biological activity (e.g., IC50 values). Molecular descriptors or fingerprints must be precomputed.
- Quantum Computing Access: Cloud-based QPU or simulator with machine learning libraries (e.g., TensorFlow Quantum, Pennylane).
- Software: Python with scikit-learn for classical ML baseline, and a QML library.
Procedure:
- Data Preprocessing (Classical):
- Standardize the molecular descriptor data (e.g., normalize features, handle missing values).
- Split the dataset into training, validation, and test sets.
- Data Encoding (Quantum):
- Design a feature map to encode classical molecular descriptor data (vector ( x )) into a quantum state ( |\phi(x)\rangle ). This can be achieved using gates like Pauli rotations (RX, RY, RZ).
- Model Construction (Hybrid):
- Construct a Variational Quantum Circuit (VQC), also known as a Quantum Neural Network.
- The circuit typically consists of the fixed feature map followed by a parameterized circuit (e.g., composed of layers of rotational gates and entangling gates).
- Model Training (Hybrid):
- Define a cost function (e.g., mean squared error for regression, cross-entropy for classification).
- Use a classical optimizer to tune the parameters of the VQC. The quantum device evaluates the cost function for given parameters, and the classical optimizer suggests updates.
- Model Evaluation (Classical):
- Use the trained model to make predictions on the held-out test set.
- Benchmark the performance (e.g., R² score, ROC-AUC) against classical machine learning models like Random Forests or Gradient Boosting.
- Data Preprocessing (Classical):
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Key Tools for Quantum-Enhanced DFT and ML Research
| Item / Solution | Category | Function & Application |
|---|---|---|
| QUAlibrate [21] | Control Software | An open-source framework that automates and drastically reduces quantum computer calibration time, essential for maintaining QPU performance for long-running chemistry simulations [21]. |
| Qiskit [23] [24] | Software SDK | An open-source full-stack SDK for creating, simulating, and running quantum circuits on IBM hardware or simulators. Includes Qiskit Machine Learning for building QML models [23]. |
| TensorFlow Quantum [24] | Software Library | A library for prototyping hybrid quantum-classical ML models. Enables the integration of quantum circuits and models with the classical TensorFlow ecosystem [24]. |
| PennyLane [23] [24] | Software Library | A cross-platform Python library for differentiable quantum computing, allowing seamless training of quantum circuits using classical automatic differentiation, ideal for VQE and QNNs [23]. |
| Amazon Braket / IBM Cloud | Cloud Platform | Provide cloud-based access to simulators and various QPU backends (e.g., from Rigetti, OQC, IonQ, IBM), lowering the barrier to entry for experimental workflows [23]. |
| Variational Quantum Eigensolver (VQE) | Algorithm | A leading hybrid quantum-classical algorithm for finding approximate eigenvalues of molecular Hamiltonians, making it a cornerstone for near-term quantum chemistry [25]. |
| Quantum Support Vector Machine (QSVM) | Algorithm | A quantum-enhanced kernel method for classification that can efficiently handle high-dimensional feature spaces, potentially useful for classifying molecular properties [23] [25]. |
| Error Suppression & Mitigation | Software/Technique | Techniques (e.g., those developed by Q-CTRL, or embedded in vendor SDKs) to reduce the impact of noise on current-generation "noisy" quantum processors, improving result fidelity [18]. |
Critical Pathways: Error Correction and Quantum Control
The viability of complex simulations hinges on the stability and accuracy of quantum computations. Recent breakthroughs in quantum error correction (QEC) and control are therefore foundational to "Why Now?"
- The Problem: Physical qubits are sensitive to environmental noise, leading to high error rates that corrupt calculations [20].
- The Solution - QEC: QEC uses multiple physical qubits to form one more-stable logical qubit. By continuously measuring syndromes to detect errors and applying real-time corrections, the integrity of the logical qubit is maintained [18] [20].
- Recent Progress (2024-2025): In 2024, Google's Willow chip demonstrated significant advancements in error correction with a low error rate [18]. Companies including IBM, Quantinuum, and QuEra have all announced new QEC architectures and logical processors, indicating that building a fault-tolerant quantum computer is now primarily an engineering challenge [18] [20]. This progress directly enables the longer, more complex computations required for accurate DFT simulations.
From Theory to Practice: A Guide to ML-DFT Workflows and Their Biomedical Applications
Density Functional Theory (DFT) has established itself as the cornerstone of computational materials science and drug discovery, providing essential insights into electronic structures that govern material and molecular properties. The integration of machine learning (ML) with DFT has emerged as a transformative approach, overcoming DFT's traditional computational bottlenecks and enabling investigations at unprecedented scales. This application note details protocols for constructing end-to-end ML-driven DFT emulation frameworks, validated through case studies in energetic materials and pharmaceutical design. We present quantitative performance benchmarks, standardized workflows for system mapping and property prediction, and a comprehensive toolkit for researchers. The documented methodologies achieve up to three orders of magnitude speedup while maintaining DFT-level accuracy, opening new frontiers in predictive materials modeling and rational drug design.
Density Functional Theory revolutionized computational chemistry and materials science by formulating electronic structure calculations in terms of electron density rather than complex wavefunctions [26]. This fundamental principle enables the prediction of material and molecular properties from first principles, making DFT an indispensable tool across scientific disciplines. In pharmaceutical research, DFT provides quantum mechanical precision for studying drug-receptor interactions, molecular reactivity, and material properties at electronic scales [27] [28]. However, conventional DFT calculations face significant computational constraints due to their cubic scaling with system size, typically limiting routine applications to systems of a few hundred atoms [26].
Machine learning frameworks now circumvent these limitations through local environment mapping and neural network surrogates. By leveraging the "nearsightedness" of electronic interactions—where local electronic structure depends primarily on nearby atomic environments—ML models can predict electronic properties with DFT-level accuracy while achieving linear scaling [26]. This paradigm shift enables electronic structure calculations for systems containing hundreds of thousands of atoms, bridging atomic-scale interactions with macroscopic material behaviors.
Computational Frameworks for DFT Emulation
Foundational ML-DFT Architectures
Local Density of States (LDOS) Learning Framework: The Materials Learning Algorithms (MALA) package implements an end-to-end workflow where bispectrum coefficients encode atomic positions relative to each point in real space, and neural networks map these descriptors to the local density of states [26]. This approach separates the problem into local mappings, enabling parallel processing and system-size independence. The LDOS encodes the local electronic structure and serves as the fundamental quantity from which observables like electronic density, density of states, and total free energy are derived [26].
Neural Network Potentials (NNPs) for Energetic Materials: The EMFF-2025 framework demonstrates a specialized approach for C, H, N, O-based high-energy materials (HEMs), leveraging transfer learning to achieve DFT-level accuracy in predicting structures, mechanical properties, and decomposition characteristics [7]. Built upon the Deep Potential (DP) scheme, this model combines high accuracy with computational efficiency, enabling large-scale molecular dynamics simulations of complex reactive processes [7].
Active Learning Integration: Simple Active Learning (SAL) workflows implement on-the-fly training of ML potentials during molecular dynamics simulations, continuously improving model accuracy through targeted DFT calculations [29]. This approach combines the efficiency of ML potentials with the accuracy of reference DFT calculations, automatically identifying configurations where the model requires refinement and retraining accordingly [29].
Quantitative Performance Benchmarks
Table 1: Accuracy Benchmarks for ML-DFT Frameworks
| Framework | System Type | Energy MAE | Force MAE | Property Predictions | Reference |
|---|---|---|---|---|---|
| EMFF-2025 | C,H,N,O HEMs | < 0.1 eV/atom | < 2.0 eV/Å | Mechanical properties, decomposition pathways | [7] |
| MALA (Beryllium) | Metallic systems | - | - | Electronic density, free energy, forces | [26] |
| B3LYP Functional | Molecular drugs | ~2.2 kcal/mol (atomization) | - | Geometries, transition barriers, ionization | [28] |
Table 2: Computational Efficiency Comparisons
| Method | System Size | Calculation Time | Scaling Behavior | Hardware Requirements |
|---|---|---|---|---|
| Conventional DFT | 256 atoms | Reference | ~N³ | High-performance computing |
| MALA ML-DFT | 131,072 atoms | 48 minutes | ~N | 150 standard CPUs |
| EMFF-2025 | 20 HEMs | Efficient screening | - | - |
Protocol: End-to-End DFT Emulation Workflow
System Preparation and Descriptor Calculation
Step 1: Atomic Configuration Preprocessing
- Generate initial atomic configurations through molecular dynamics or experimental coordinates
- Apply periodic boundary conditions appropriate for the target system
- For crystalline materials, include defect structures and surfaces relevant to properties of interest
Step 2: Descriptor Computation
- Calculate bispectrum coefficients for each point in real space using LAMMPS integration [26]
- Set cutoff radius consistent with the nearsightedness principle (typically 4-6 Å)
- Encode atomic density within local environments using order parameters that respect physical symmetries
Neural Network Training and Validation
Step 3: Initial Model Training
- Implement feed-forward neural networks using PyTorch framework [26]
- Train on diverse reference systems (256 atoms for bulk materials)
- Use energy and force predictions from DFT calculations as training targets
- Employ stratified sampling to ensure adequate representation of different bonding environments
Step 4: Active Learning Implementation
- Integrate Simple Active Learning workflow for molecular dynamics simulations [29]
- Configure triggering criteria for reference DFT calculations (uncertainty thresholds, regular intervals)
- Implement rewind mechanism to continue from last accurate simulation point
- Set convergence criteria for model accuracy and property prediction stability
Property Prediction and Validation
Step 5: Electronic Structure Analysis
- Compute electronic density and density of states from predicted LDOS [26]
- Derive total free energy and atomic forces for molecular dynamics simulations
- Calculate band structures and density of states for crystalline materials
Step 6: Experimental Validation
- Compare predicted crystal structures and mechanical properties with experimental data [7]
- Validate thermal decomposition pathways against experimental observations
- Benchmark electronic properties against spectroscopic measurements
Application in Pharmaceutical Sciences
Drug Design and Optimization
DFT calculations provide critical insights for pharmaceutical development by elucidating electronic interactions between drug molecules and biological targets. The exceptional accuracy of DFT (approximately 0.1 kcal/mol) enables precise reconstruction of molecular orbital interactions, facilitating rational drug design [27].
Table 3: DFT Applications in Drug Development
| Application Area | DFT Methodology | Key Parameters | Impact on Drug Development |
|---|---|---|---|
| API-Excipient Compatibility | Fukui function analysis | Reactive site identification | Guided stability-oriented co-crystal design [27] |
| Nanodrug Delivery Systems | van der Waals & π-π stacking calculations | Interaction energies | Optimized carrier surface distribution [27] |
| Solubility & Release Kinetics | COSMO solvation models | ΔG solvation | Controlled-release formulation design [27] |
| Reaction Mechanism Elucidation | Transition state modeling | Activation energies | Enzyme inhibition optimization [28] |
COVID-19 Research Applications
DFT has played a critical role in pandemic response through rapid screening of therapeutic candidates. Researchers have employed DFT to study amino acids as immunity boosters, identify arginine as particularly effective, and analyze tetrazole derivatives for anti-COVID-19 activity [28]. These applications demonstrate DFT's versatility in addressing emergent health challenges through electronic structure analysis.
The Scientist's Toolkit: Essential Research Reagents
Table 4: Computational Tools for DFT Emulation
| Tool/Category | Specific Implementation | Function | Application Context |
|---|---|---|---|
| DFT Engines | Quantum ESPRESSO, ADF, BAND | Reference calculations | Provide training data for ML potentials [26] [29] |
| ML Potential Frameworks | MALA, EMFF-2025, DP-GEN | Surrogate model training | Large-scale property prediction [7] [26] |
| Descriptor Calculators | LAMMPS | Atomic environment encoding | Convert atomic positions to feature vectors [26] |
| Active Learning Workflows | Simple Active Learning (SAL) | On-the-fly training | Self-improving MD simulations [29] |
| Force Field Integrators | ONIOM, M3GNet | Multiscale simulations | Hybrid QM/MM calculations [27] |
| Analysis Packages | PyTorch, scikit-learn | Model evaluation | Performance validation [26] |
Future Perspectives and Development
The integration of machine learning with DFT continues to evolve, with several emerging trends shaping future developments. Hybrid methodologies that combine ML efficiency with DFT accuracy are expanding to more complex systems, including heterogeneous interfaces and disordered materials. Deep learning approaches are being applied directly to approximate kinetic energy density functionals, potentially overcoming fundamental limitations of traditional exchange-correlation functionals [27]. The development of transferable potential frameworks, demonstrated by EMFF-2025's application across multiple high-energy materials, points toward more generalized models that maintain accuracy across diverse chemical spaces [7].
As these methodologies mature, ML-enhanced DFT emulation will increasingly serve as the foundation for predictive materials science and pharmaceutical development, enabling first-principles accuracy at scales previously inaccessible to computational investigation. This paradigm shift promises to accelerate the design cycle for functional materials and therapeutic compounds, fundamentally transforming computational discovery processes.
Learning the Exchange-Correlation Functional with Neural Networks
Density Functional Theory (DFT) is a cornerstone computational method for solving the many-body Schrödinger equation, with unparalleled impact across quantum chemistry, materials science, and drug discovery [30] [31]. Its practical success hinges entirely on the exchange-correlation (XC) functional, which encapsulates complex electron interactions. The exact form of this functional remains unknown, forcing approximations and limiting accuracy, particularly for systems with strong electron correlations [32].
Traditional approaches to developing XC functionals, like the Local Density Approximation (LDA) and Generalized Gradient Approximation (GGA), rely on heuristic rules and analytic solutions to specific limits. These forms are inherently inflexible, making it difficult to incorporate new numerical data from high-level quantum theories [30]. Machine learning (ML), particularly neural networks (NNs), offers a path beyond these constraints. NN XC functionals provide a universal, highly flexible framework for interpolation and approximation, capable of learning directly from data generated by quantum Monte Carlo or post-Hartree-Fock methods, promising a new frontier of accuracy in DFT [30] [31].
This document details the application notes and protocols for constructing, training, and implementing neural network-based exchange-correlation functionals, contextualized within a broader research thesis on machine learning-enhanced DFT workflows.
Neural Network Architectures for XC Functionals
The design of the neural network architecture is critical for accurately representing the physical relationship between the electron density and the XC functional. The following table compares the primary architectures explored in recent literature.
Table 1: Comparison of Neural Network Architectures for XC Functionals
| Architecture Name | Input Features | Output(s) | Key Features & Advantages | Example System Tested |
|---|---|---|---|---|
| Point-to-Point (LDA-like) [30] | Electron density ((n)) at a single point in space. | XC potential ((v{xc})) or energy density ((\epsilon{xc})) at that point. | Simple, fully-connected network; mirrors locality of LDA. | 3D inhomogeneous electron gas in a harmonic oscillator potential. |
| Region-to-Point (GGA-like) [30] | Electron density in a 5×5×5 cube surrounding a point (enables gradient calculation). | XC potential ((v{xc})) or energy density ((\epsilon{xc})) at the central point. | Captures inhomogeneity; learns gradients without explicit feature engineering. | 3D inhomogeneous electron gas; Crystalline Silicon. |
| Two-Component (NN-E & NN-V) [31] | NN-E: (n), (\sigma) (gradient squared).NN-V: (\epsilon_{xc}), (n), (\sigma), (\gamma), (\nabla^2 n). | NN-E: (\epsilon{xc}).NN-V: (v{xc}). | Separates energy and potential; ensures correct physical link; "economical" for memory. | Crystalline silicon, benzene, ammonia, atoms/molecules from IP13/03 dataset. |
| Differentiable Neural Functional (Grad DFT) [32] | Features for a given approximation (e.g., for GGA: (n), (\sigma)). | Exchange-correlation energy ((E_{xc})). | Fully differentiable library (JAX); enables end-to-end training against energies and properties. | Experimental dissociation energies of dimers, including transition metals. |
The logical flow and data transformation within a Two-Component NN architecture can be visualized as follows:
The workflow for developing and deploying an NN XC functional, from data generation to self-consistent calculation, is outlined below:
Performance and Validation
Trained NN XC functionals must be quantitatively evaluated against traditional functionals and reference data. The following table summarizes key performance metrics from documented experiments.
Table 2: Quantitative Performance of NN XC Functionals on Test Systems
| NN Functional Type | Training System | Test System | Key Metric: MAE (vs. Reference) | Performance Summary |
|---|---|---|---|---|
| NN LDA [30] | 3D electron gas (harmonic potential) | Crystalline Silicon (diamond) | Vxc MAE: 0.6 mHa × Bohr³ | Excellent agreement with reference Octopus data. |
| NN GGA [30] | 3D electron gas (harmonic potential) | Crystalline Silicon (diamond) | Vxc MAE: 18.1 mHa × Bohr³ | Reasonable agreement; errors in high-density-variation regions. |
| Two-Component NN [31] | Crystalline Si, Benzene, NH3 (PBE) | Atoms/Molecules (IP13/03 dataset) | Total Energy Relative Error: ~0.01% | Small error on unseen data; functional works in self-consistent cycle. |
Experimental Protocols
Protocol 1: Data Generation for Training
Objective: To generate a high-quality dataset of electron densities and their corresponding XC potentials/energies for training NN models.
Materials:
- Software: A real-space DFT code like Octopus [30] [31].
- XC Functional: A standard functional (e.g., PBE [31] or LDA [30]) to generate target data.
- System Selection: A diverse set of systems (e.g., 3D electron gases in harmonic potentials [30], simple atoms, molecules like benzene and ammonia, and crystalline solids like silicon [31]).
Procedure:
- Define Calculation Parameters:
- For each system, define the simulation box (e.g., a 40 × 40 × 40 Bohr parallelepiped).
- Set a real-space grid (e.g., 32 × 32 × 32 points, corresponding to a ~1.25 Bohr spacing).
- Run DFT Calculations:
- Perform a self-consistent field (SCF) calculation for each system using the chosen standard XC functional.
- Upon convergence, output the final electron density ((n(\mathbf{r}))) and the corresponding XC potential ((v_{xc}(\mathbf{r}))) for every grid point in the system.
- Calculate Additional Features (for GGA and beyond):
- From the electron density grid, compute its derivatives:
- (\sigma = |\nabla n|^2) (squared gradient)
- (\gamma = \langle \nabla \sigma, \nabla n \rangle)
- (\nabla^2 n) (Laplacian)
- From the electron density grid, compute its derivatives:
- Dataset Assembly:
Protocol 2: Two-Component NN Training
Objective: To train a two-component neural network that separately predicts the XC energy density and the XC potential while preserving their physical relationship [31].
Materials:
- Software: A deep learning framework like TensorFlow/PyTorch.
- Dataset: Generated from Protocol 1.
Procedure: Stage 1: Pre-train the NN-V Component
- Fix NN-E: Do not train NN-E at this stage. Use the reference (\epsilon_{xc}^{PBE}) from your dataset as input to NN-V.
- Configure NN-V Inputs: For each point, feed the following features into NN-V: reference (\epsilon_{xc}), (n), (\sigma), (\gamma), and (\nabla^2 n).
- Train NN-V: Minimize a loss function (e.g., Mean Squared Error) between the NN-V output ((v{xc}^{predicted})) and the reference potential ((v{xc}^{PBE})) from the dataset. The goal is to teach NN-V the precise mapping from energy density to potential.
- Freeze NN-V: Once NN-V is trained, freeze its weights.
Stage 2: Train the NN-E Component
- Connect the Network: The output of NN-E ((\epsilon_{xc}^{predicted})) is now fed as input to the frozen NN-V.
- Define the Composite Loss Function: The total loss for training NN-E is a weighted sum of:
- Potential Loss: ((v{xc}^{PBE} - v{xc}^{predicted})^2). This ensures the final potential is accurate.
- Boundary Condition Losses:
- ((\epsilon_{xc}^{predicted}(n \to 0) - 0)^2): Ensures energy density vanishes as density vanishes.
- ((\epsilon{xc}^{predicted}(\sigma \to 0) - \epsilon{xc}^{LDA}(n))^2): Ensures the functional reduces to LDA where the density is uniform.
- Train NN-E: Minimize the composite loss function by updating only the weights of NN-E. This trains NN-E to produce an energy density that, when passed through the physically-grounded NN-V, yields a correct potential and obeys key physical constraints.
Protocol 3: Self-Consistent Implementation and Testing
Objective: To integrate a trained NN XC functional into a DFT code and run self-consistent calculations to validate its performance and transferability.
Materials:
- DFT Platform: A code that allows for user-defined XC functionals, such as Octopus.
- Trained NN Model: The final model from Protocol 2.
Procedure:
- Integration:
- Implement a wrapper routine in the DFT code that, given an electron density grid (n(\mathbf{r})), calls the trained NN model to compute (v_{xc}(\mathbf{r})) at every point.
- Self-Consistent Cycle:
- Initialize the electron density (e.g., from superposition of atomic densities).
- Begin the Kohn-Sham cycle. Instead of calling a standard LibXC functional, use the NN wrapper to compute the XC potential.
- Construct the Kohn-Sham Hamiltonian, solve for the orbitals, and compute a new electron density.
- Repeat until the density or total energy converges.
- Validation:
- Energy Accuracy: Compare the total energy and XC energy from the NN-driven calculation against results from high-level theories or experimental data (e.g., dissociation energies of dimers [32]).
- Transferability: Test the NN functional on systems not included in the training set (e.g., new molecules or materials) to assess its generalizability [30] [31].
- Property Prediction: Evaluate the model's performance on predicting properties beyond total energy, such as forces, dipole moments, and electronic eigenvalues.
The Scientist's Toolkit
Table 3: Essential Software and Data Resources for NN XC Functional Development
| Resource Name | Type | Primary Function | Relevant Citation |
|---|---|---|---|
| Octopus | Software | Real-space DFT code used for generating training data and implementing NN XC functionals in self-consistent field calculations. | [30] [31] |
| Grad DFT | Software | A fully differentiable, JAX-based library for quick prototyping and training of machine learning-enhanced XC energy functionals. | [32] |
| LibXC | Software | A standard library of exchange-correlation functionals; used to generate target data for pre-training stages and for benchmark comparisons. | [31] |
| TensorFlow / PyTorch | Software | Deep learning frameworks used for constructing, training, and deploying neural network models for the XC functional. | [30] |
| Quantum Monte Carlo / Post-HF Data | Data | High-precision data from advanced electronic structure methods, serving as the ultimate target for training highly accurate NN functionals. | [30] [31] |
| Quantum Chemical Databases (e.g., IP13/03) | Data | Curated datasets of molecular properties (e.g., energies) for validating the transferability and accuracy of developed NN functionals. | [31] |
Atom-Centered Fingerprints and Density Descriptors for Model Input
In machine learning for chemistry and materials science, descriptors transform raw atomic Cartesian coordinates into a numerical representation that encodes essential invariances and physical properties. The accuracy, speed, and reliability of machine learning interatomic potentials (MLIPs) depend strongly on this choice of input representation. Effective descriptors must be invariant to fundamental symmetries: translation and rotation of the entire system, and permutation of like atoms. Atom-centered fingerprints and electronic density descriptors have emerged as powerful classes of representations that fulfill these requirements while capturing the local chemical environment or global electronic structure critical for predicting material properties.
Atom-centered descriptors typically encode the local atomic environment around a central atom using a structural representation, while electronic density descriptors capture features related to the electron density distribution. These representations enable machine learning models to bypass the explicit, computationally expensive solution of the Kohn-Sham equations in Density Functional Theory (DFT), achieving orders of magnitude speedup while maintaining chemical accuracy. This protocol details the application of these descriptors within DFT-machine learning workflows.
Atom-Centered Symmetry Functions and Fingerprints
Definition and Purpose
Atom-centered fingerprints are fixed-length numerical vectors that describe the chemical environment surrounding each atom in a structure. They serve as input for machine learning models that predict atomic-scale properties, effectively replacing the explicit calculation of electronic structure in DFT. Their primary function is to convert the atomic configuration into a machine-readable format that respects physical symmetries.
Key Methodologies and Protocols
Automated Fingerprint Selection Protocol: A critical advancement is the automated selection of optimal fingerprints from a large pool of candidates. The following protocol, adapted from Imbalzano et al., streamlines the construction of neural network potentials [33]:
- Generate a Large Candidate Pool: Create an extensive initial set of symmetry functions or fingerprint candidates. These often include radial functions (describing the density of neighboring atoms at various distances) and angular functions (describing the angular distribution of atom triplets).
- Compute Correlation Matrix: Calculate the intrinsic correlations between all fingerprint candidates across the training dataset of atomic configurations.
- Select Informative Subset: Identify a minimal subset of fingerprints that maintains low mutual correlation while maximally spanning the variability in the training data. This prevents redundancy and overfitting.
- Validate and Balance: Evaluate the selected subset on a validation set to ensure the resulting ML potential strikes the desired balance between computational efficiency and predictive accuracy.
AGNI Fingerprints Workflow: The AGNI (Atom-Centered Neural Network) fingerprints represent a specific implementation widely used for creating ML force fields [2]. The protocol for their application in an ML-DFT framework is as follows:
- Input: Atomic configuration of the system.
- Descriptor Calculation: For each atom
i, the fingerprint is computed by summing Gaussian functions centered on neighboring atomsjwithin a cutoff radius. The functions incorporate interatomic distancesR_ijand can be extended to include angular information via three-body terms involving atomsjandk. - Output: A translation, permutation, and rotation-invariant vector for each atom
i, describing its local chemical environment.
Application Notes
Automated fingerprint selection can greatly simplify the construction of neural network potentials and accelerate the evaluation of Gaussian approximation potentials (GAP) based on the smooth overlap of atomic positions (SOAP) kernel [33]. These fingerprints have been successfully applied to diverse systems, including water, Al-Mg-Si alloys, and small organic molecules [33] [2].
Electronic Density-Based Descriptors
Density of States (DOS) Similarity Descriptor
The DOS provides a comprehensive summary of a material's electronic structure. A tailored fingerprint has been developed to facilitate quantitative comparison of DOS spectra across different materials [34].
Protocol: Constructing a DOS Fingerprint
This protocol transforms a continuous DOS, ρ(ε), into a binary-valued 2D map [34].
- Energy Reference Shift: Shift the energy spectrum so that the reference energy (e.g., the Fermi level) is at
ε_ref = 0. - Non-Uniform Energy Discretization: Integrate the DOS over
N_εintervals of variable width,Δε_i, to create a histogram{ρ_i}. The interval widths are defined to provide finer discretization around the feature region (|ε| < W, whereWis a tunable parameter) and coarser discretization away from it. This focuses the descriptor on physically relevant energy regions.Δε_i = n(ε_i, W, N) * Δε_minwherenis an integer-valued function that increases from 1 to a maximumNas|ε|increases beyondW. - Histogram Intensity Discretization: Discretize each column
iof the histogram into a grid ofN_ρintervals of variable heightΔρ_i. This step uses a similar non-uniform scaling governed by parametersW_HandN_Hto accentuate features in the high-density regions. - Binary Image Generation: For each column
i, the number of "filled" pixels is given bymin(⌊ρ_i / Δρ_i⌋, N_ρ). This generates a 2D raster image withN_ε × N_ρpixels. - Vector Encoding: Flatten the 2D image into a binary-encoded vector
f, where each componentf_αis 1 if the pixel is filled and 0 otherwise.
Similarity Metric: The similarity between two materials i and j with fingerprints f_i and f_j is quantified using the Tanimoto coefficient (Tc) [34]:
S(f_i, f_j) = (f_i · f_j) / (|f_i|^2 + |f_j|^2 - f_i · f_j)
Charge Density Descriptors
In an end-to-end ML framework to emulate DFT, the electron charge density itself is predicted first and then used as a descriptor for other properties [2].
Protocol: Two-Step ML-DFT Emulation
Step 1: Predict Charge Density
- Input: Atomic configuration, represented using atomic fingerprints (e.g., AGNI).
- Model: A deep neural network maps the atomic fingerprints to a set of coefficients and exponents for a decomposition of the atomic charge density in terms of Gaussian-type orbitals (GTOs). The model learns the optimal GTO basis set from the data.
- Output: A predicted electron charge density, obtained by projecting the atomic GTOs onto a real-space grid. This step requires transforming from the internal, rotation-invariant atomic reference frame to the global Cartesian frame.
Step 2: Predict Other Properties
- Input: The original atomic fingerprints and the predicted charge density descriptors.
- Model: A second deep neural network uses this combined input to predict a host of electronic and atomic properties, such as Density of States (DOS), band gap, total potential energy, atomic forces, and stress tensor.
Application Notes
The DOS similarity descriptor enables unsupervised learning and clustering of large materials databases, revealing groups of materials with similar electronic properties that may not be obvious from their composition or crystal structure alone [34]. The ML-DFT approach with charge density descriptors provides a complete DFT emulation, allowing for molecular dynamics simulations with linear scaling with system size and a small prefactor [2].
Workflow Integration and Advanced Applications
Active Learning for Potential Generation
Advanced workflows like AL4GAP integrate active learning for the efficient generation of Gaussian Approximation Potentials (GAP) for complex systems [35].
Diagram 1: Active learning workflow for generating machine learning potentials.
Protocol: The AL4GAP Workflow for Molten Salts [35]
- Combinatorial Space Setup: Define the chemical space of interest, e.g., charge-neutral mixtures of 11 cations and 4 anions.
- Configurational Sampling: Perform molecular dynamics or Monte Carlo sampling using low-cost empirical potentials to explore the configuration space of the mixtures.
- Active Learning: Use an ensemble of MLIPs to down-select the most informative or uncertain configurations from the sampled pool for costly DFT calculations. This step maximizes the information gain per DFT calculation.
- DFT Calculations: Perform single-point DFT calculations (e.g., using the SCAN functional) on the selected configurations to obtain accurate energies and forces.
- Hyperparameter Tuning: Employ Bayesian optimization to tune the parameters of the two-body and many-body GAP models.
- Iteration and Validation: Iterate steps 2-5 until the GAP model achieves the desired accuracy on a hold-out test set.
Application-Specific Functional Descriptors
For specific applications like screening ionic liquids for CO₂ capture, customized functional structure descriptors (FSD) can be constructed. These are based on a group contribution method and can be combined with dimensionless descriptors like "CORE" to build highly accurate quantitative structure-property relationship (QSPR) models using ensemble learning methods (e.g., CatBoost) [36].
The Scientist's Toolkit: Research Reagent Solutions
Table 1: Essential computational tools and descriptors for MLIP development.
| Name/Type | Brief Function/Explanation | Example Application Context |
|---|---|---|
| Atom-Centered Symmetry Functions | Invariant descriptors of the local atomic density and angular distribution within a cutoff radius. | Core input for high-dimensional neural network potentials (HDNNPs) and AGNI force fields [2]. |
| Smooth Overlap of Atomic Positions (SOAP) | A unified descriptor that generalizes atom-centered density correlations, providing a body-ordered representation of the atomic environment. | Basis for Gaussian Approximation Potentials (GAP) [33]. |
| Density-of-States (DOS) Fingerprint | A binary-encoded 2D map that allows for quantitative similarity comparison between materials based on electronic structure [34]. | Unsupervised clustering of materials databases for discovery and analysis [34]. |
| Gaussian-Type Orbitals (GTO) Descriptors | A basis set used to represent the electron charge density around an atom. The coefficients and exponents can be learned by a model. | Predicting the electron charge density from atomic structure in ML-DFT emulation [2]. |
| Active Learning Workflow (AL4GAP) | A software workflow that automates the iterative process of building accurate ML potentials with minimal DFT data [35]. | Generating DFT-accurate potentials for combinatorial molten salt mixtures [35]. |
| Functional Structure Descriptor (FSD) | A descriptor based on group contribution methods, designed for interpretability and application-specific tasks. | Screening and design of ionic liquids for CO₂ capture [36]. |
Quantitative Comparison of Descriptor Performance
Table 2: Performance metrics of selected descriptor-driven models.
| Descriptor / Model | System / Property | Key Performance Metric | Reference |
|---|---|---|---|
| Automated Fingerprint Selection | Neural Network Potentials for Water, Al-Mg-Si Alloy | Greatly simplified model construction; orders of magnitude acceleration for GAP evaluation. | [33] |
| CatBoost with FSD | CO₂ Solubility in Ionic Liquids | R² = 0.9945, MAE = 0.0108 | [36] |
| CatBoost with CORE | CO₂ Solubility in Ionic Liquids | R² = 0.9925, MAE = 0.0120 | [36] |
| Two-Step ML-DFT | Organic Molecules, Polymer Chains/Crystals (C, H, N, O) | Predicts charge density, DOS, energy, forces with chemical accuracy; linear scaling. | [2] |
| AL4GAP GAP Models | Multicomposition Molten Salt Mixtures (e.g., NaCl-CaCl₂, KCl-NdCl₃) | Accurately predicts structure with DFT-SCAN accuracy, captures intermediate range ordering. | [35] |
Generalized Protocol for Implementing Atom-Centered Fingerprints in an ML Potential:
- System Definition: Define the chemical system and the target properties (e.g., energy, forces).
- Reference Data Generation: Use DFT-based molecular dynamics to generate a diverse set of atomic configurations and their corresponding DFT-calculated target properties.
- Fingerprint Calculation & Selection: Compute a large pool of candidate fingerprints for all atoms in all configurations. Apply an automated selection protocol to identify a robust and non-redundant subset.
- Model Training: Train a machine learning model (e.g., neural network, Gaussian process regression) to map the selected fingerprints to the target properties.
- Active Learning (Optional): For complex systems, use an active learning workflow to iteratively improve the model by selectively adding new DFT calculations in underrepresented or uncertain regions of the configuration space.
- Validation: Rigorously test the final ML potential on a held-out test set of configurations not seen during training.
Diagram 2: Generalized protocol for building a machine learning interatomic potential.
The integration of machine learning (ML) with foundational computational chemistry principles like density functional theory (DFT) is revolutionizing the drug discovery pipeline. This paradigm addresses the high costs and long timelines traditionally associated with bringing a new drug to market, which can exceed 10-15 years and $2.5 billion [37]. ML models serve as powerful in-silico surrogates, predicting molecular properties to prioritize the most promising candidates for synthesis and experimental testing [38]. DFT provides the crucial chemical accuracy needed to understand electronic structures and reaction mechanisms at enzyme active sites, a level of detail unattainable with classical molecular mechanics (MM) methods [39] [40]. This case study explores how integrating ML-predicted properties with DFT-based validation creates a powerful, accelerated workflow for modern drug discovery, with a specific application in designing inhibitors for SARS-CoV-2.
Integrated DFT and ML Workflow: Methodology and Protocol
The synergy between DFT and ML can be structured into a cohesive, iterative workflow. The following diagram illustrates this integrated pipeline, from initial molecular screening to validated lead compounds.
Protocol 1: ML-Based Virtual Screening and Property Prediction
The initial phase leverages ML to rapidly evaluate vast chemical libraries for desirable drug-like properties.
- Objective: To screen extensive molecular libraries in silico to identify hits with predicted high binding affinity and optimal ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) properties [41].
- Input: A library of molecular structures, typically represented as SMILES (Simplified Molecular Input Line Entry System) strings or 2D/3D structural files [42].
- Molecular Embedding: Convert molecular structures into machine-readable numerical representations. Common techniques include:
- Model Training and Prediction: Implement supervised ML models, typically tree-based ensembles like XGBoost, CatBoost, or LightGBM, to predict properties of interest [42]. The model is trained on curated datasets, such as those from the CRC Handbook of Chemistry and Physics, containing known molecular properties [42].
- Output: A prioritized list of candidate molecules ranked by their predicted binding affinity and ADMET scores [41].
Table 1: Key Molecular Properties for ML Prediction in Drug Discovery
| Property | Description | Impact on Drug Discovery |
|---|---|---|
| Binding Affinity | Predicted strength of interaction with a target protein. | Primary indicator of compound efficacy [41]. |
| Solubility | Ability of a compound to dissolve in aqueous solution. | Critical for drug absorption and bioavailability [41]. |
| Melting/Boiling Point | Fundamental physicochemical properties. | Informs synthesis and formulation processes [42]. |
| ADMET Profile | Composite of absorption, distribution, metabolism, excretion, and toxicity. | Key determinant of in-vivo safety and pharmacokinetics [41] [37]. |
Protocol 2: DFT Validation of Electronic Properties and Interactions
Top-ranking candidates from ML screening undergo rigorous quantum mechanical analysis using DFT.
- Objective: To provide chemical accuracy for studying drug-target interactions, particularly the electronic properties and reaction mechanisms at enzyme active sites [39] [40].
- System Setup: Select a representative model of the enzyme's active site, including key amino acid residues, and the bound ligand.
- DFT Calculations: Perform electronic structure calculations using a suitable functional (e.g., PBE) and basis set [39]. Key analyses include:
- Electronic Structure Analysis: Calculation of frontier molecular orbitals (HOMO-LUMO) to understand reactivity [40].
- Reaction Mechanism Elucidation: Modeling the formation and breaking of bonds during enzyme inhibition, such as the covalent linkage formation with the Cys-His catalytic dyad in SARS-CoV-2 Mpro [39].
- Binding Energy Decomposition: Detailed analysis of interaction energies between the drug and key residues in the target.
- Output: Atomic-level insight into the mechanism of action and validation of the binding mode predicted by ML.
Protocol 3: Experimental Validation via Label-Free Phenotypic Screening
Computational predictions require experimental confirmation. Label-free high-content screening provides a robust method for this validation.
- Objective: To experimentally detect cellular drug responses without the need for fluorescent labeling, using high-throughput bright-field imaging and ML [43].
- Cell Culture and Treatment: Culture model cell lines (e.g., MCF-7 breast cancer cells). Treat with candidate drugs and appropriate controls (e.g., paclitaxel as a positive control, DMSO as a negative control) [43].
- High-Throughput Imaging: Acquire bright-field images of cells at high speed (up to 10,000 cells/s) using optofluidic time-stretch microscopy or similar platforms [43].
- Image Analysis and Classification: Extract hundreds of morphological features (geometry, granularity, texture) from single-cell images. Train a Support Vector Machine (SVM) classifier to distinguish between drug-treated and untreated cell populations based on these subtle morphological changes [43].
- Output: A classification accuracy score, where a higher score indicates a more pronounced and detectable drug-induced phenotypic change, providing experimental validation of compound efficacy [43].
The following diagram details this experimental protocol.
Application: SARS-CoV-2 Antiviral Development
This integrated workflow has been successfully applied to accelerate the discovery of therapeutics for emerging diseases like COVID-19.
- Target Identification: Two primary SARS-CoV-2 targets are the Main Protease (Mpro/3CLpro), essential for viral replication, and the RNA-dependent RNA Polymerase (RdRp) [39].
- ML and DFT in Action: ML models screened vast compound libraries for potential Mpro and RdRp inhibitors. Promising hits, including natural products and repurposed drugs, were then studied with DFT. DFT calculations provided atomic-level insight into how these inhibitors interact with the catalytic dyad of Mpro or act as nucleotide analogs in RdRp [39].
- Multi-Objective Optimization: During the lead optimization stage, ML models balanced the prediction of multiple properties simultaneously—such as optimizing for high binding affinity while maintaining a favorable ADMET profile—to ensure the development of well-rounded drug candidates [41].
Performance Metrics and Data
The performance of ML models in this pipeline is critical. Rigorous benchmarking using domain-appropriate metrics is essential for reliable adoption in drug discovery [38].
Table 2: Performance Metrics for ML Molecular Property Prediction
| Model / Framework | Property Predicted | Performance Metric & Result | Key Findings |
|---|---|---|---|
| ChemXploreML (with Mol2Vec) | Critical Temperature (CT) | R² = 0.93 [42] | Mol2Vec (300D) offered slightly higher accuracy, while VICGAE (32D) provided greater computational efficiency. |
| SVM Classifier (Image-Based) | Phenotypic Drug Response | 92% Accuracy [43] | Achieved in distinguishing paclitaxel-treated vs. untreated MCF-7 cells using label-free bright-field images. |
| General ML Models | Various ADMET Endpoints | N/A | Neural networks are flexible but do not always outperform simpler models; high-quality training data is paramount [41]. |
Successful implementation of this workflow relies on a suite of software, data, and computational resources.
Table 3: Essential Research Reagents and Computational Tools
| Category | Item / Software | Function / Description |
|---|---|---|
| Computational Chemistry | DFT Software | Performs quantum mechanical calculations to determine electronic structure and reaction mechanisms [39]. |
| RDKit | Open-source cheminformatics toolkit for working with molecular structures and data [42]. | |
| Machine Learning | ChemXploreML | A modular desktop application for building custom ML pipelines for molecular property prediction [42]. |
| Scikit-learn, XGBoost, PyTorch | Open-source libraries for implementing standard and deep learning ML algorithms [44] [42]. | |
| Data & Databases | CRC Handbook Dataset | A reliable source of fundamental molecular properties for training and validating ML models [42]. |
| Broad Bioimage Benchmark Collection (BBBC) | A collection of published image sets for developing and testing image analysis algorithms [45]. | |
| Experimental Screening | CellProfiler / Analyst | Open-source software for automated image analysis of cellular phenotypes [45]. |
| Optofluidic Microscopy | Enables high-throughput, label-free bright-field imaging for phenotypic screening [43]. |
The fusion of DFT's chemical accuracy with ML's predictive speed creates a powerful engine for accelerating drug discovery. This case study demonstrates a robust protocol where ML rapidly identifies promising candidates, DFT provides deep mechanistic validation, and label-free experimental screens offer efficient phenotypic confirmation. This integrated approach, exemplifying the core thesis of hybrid DFT-ML workflows, enhances precision, reduces development costs and timelines, and is poised to tackle complex challenges in pharmaceutical research, from antiviral development to personalized medicine.
The convergence of density functional theory (DFT) and machine learning (ML) is revolutionizing the computational design and analysis of nanomaterials for applications in electronics and medicine. While DFT provides a quantum mechanical framework to model material properties at the atomic scale, its predictive accuracy is often limited by approximations in the exchange-correlation functionals and substantial computational costs, particularly for complex nanomaterial systems [15] [4]. Machine learning workflows address these limitations by creating data-driven surrogate models trained on DFT datasets, enabling high-throughput screening and accurate property prediction at a fraction of the computational expense [15] [6]. This integrated approach is accelerating the development of advanced nanomaterials, from electronic components with tailored band gaps to nanomedicines with optimized biological interactions.
Table 1: ML-Corrected Formation Enthalpy Performance for Ternary Alloys (0 K)
| System | Application Context | DFT Mean Absolute Error (eV/atom) | ML-Corrected Mean Absolute Error (eV/atom) | Key ML Model Parameters |
|---|---|---|---|---|
| Al-Ni-Pd [6] | High-temperature protective coatings (aerospace) | 0.082 | 0.021 | Multi-layer perceptron (MLP) regressor, 3 hidden layers, LOOCV |
| Al-Ni-Ti [6] | High-strength, low-density aerospace alloys | 0.075 | 0.018 | Multi-layer perceptron (MLP) regressor, 3 hidden layers, LOOCV |
Table 2: Optimized Hubbard U Parameters for Metal Oxide Band Gaps
| Material | Application Context | Optimal Ud/f (eV) | Optimal Up (eV) | Resulting Band Gap Accuracy vs. Experiment |
|---|---|---|---|---|
| Rutile TiO₂ [46] | Electronics, Photocatalysis | 8 | 8 | Closely matched |
| Anatase TiO₂ [46] | Electronics, Photocatalysis | 6 | 3 | Closely matched |
| c-CeO₂ [46] | Catalysis, Biomedical | 12 | 7 | Closely matched |
| c-ZnO [46] | Sensors, Electronics | 12 | 6 | Closely matched |
Experimental Protocols
Protocol 1: ML-Augmented DFT for Formation Enthalpy Correction
Application Note: This protocol details a method to correct systematic errors in DFT-calculated formation enthalpies of binary and ternary alloys, which is crucial for accurate prediction of phase stability in high-performance nanomaterials [6].
Methodology:
Reference Data Curation:
- Compile a dataset of experimentally measured formation enthalpies ((H_{f,exp})) for binary and ternary alloys of interest.
- Calculate the corresponding DFT formation enthalpies ((H_{f,DFT})) for all entries in the dataset using a consistent computational setup (e.g., EMTO method with GGA-PBE functional) [6].
- Compute the target value for ML training: (\Delta Hf = H{f,exp} - H_{f,DFT}).
Feature Engineering:
- For each compound, construct an input feature vector containing:
- Elemental concentrations ((xA, xB, x_C)).
- Weighted atomic numbers ((xA ZA, xB ZB, xC ZC)).
- Non-linear interaction terms between elemental features [6].
- Normalize all input features to a common scale.
- For each compound, construct an input feature vector containing:
Model Training and Validation:
- Implement a Multi-Layer Perceptron (MLP) regressor with three hidden layers.
- Employ Leave-One-Out Cross-Validation (LOOCV) and k-fold cross-validation to prevent overfitting and ensure model generalizability [6].
- Train the model to predict the enthalpy discrepancy (\Delta H_f) based on the input features.
Prediction and Correction:
- For a new nanomaterial, calculate (H_{f,DFT}) using standard DFT.
- Use the trained ML model to predict the correction (\Delta H_{f,ML}).
- Obtain the corrected formation enthalpy: (H{f,corrected} = H{f,DFT} + \Delta H_{f,ML}).
Protocol 2: Hybrid DFT+U/ML Workflow for Band Gap Prediction
Application Note: This protocol leverages a hybrid DFT+U and ML approach to accurately predict the band gaps of metal oxides, a critical property for electronic devices and catalytic nanomaterials [46].
Methodology:
DFT+U Parameter Space Exploration:
- Select a metal oxide system (e.g., TiO₂, ZnO, CeO₂).
- Perform a grid of DFT+U calculations, systematically varying the Hubbard U parameters for metal d/f orbitals ((U{d/f})) and oxygen p orbitals ((Up)) [46].
- For each (U({d/f}), U(p)) pair, compute the electronic band gap and lattice parameters.
Benchmarking and Optimal U Selection:
- Identify the (U({d/f}), U(p)) pair that yields band gaps and lattice parameters closest to experimental values for a subset of materials [46].
Machine Learning Model Development:
- Use the dataset of (U({d/f}), U(p)) inputs and resulting band gap outputs from the DFT+U calculations to train a supervised ML regression model.
- Simple regression algorithms (e.g., linear models, decision trees) have proven effective for this task, generalizing well to related polymorphs at a minimal computational cost [46].
Deployment for High-Throughput Screening:
- The trained ML model can rapidly predict the band gap of new, related metal oxide nanomaterials for any given U parameter set, bypassing the need for explicit, computationally expensive DFT+U calculations [46].
Workflow Visualization
The Scientist's Toolkit
Table 3: Essential Research Reagents and Computational Tools
| Item Name | Function/Brief Explanation | Application Context |
|---|---|---|
| VASP (Vienna Ab initio Simulation Package) [46] | Performs DFT and DFT+U calculations to compute total energies, electronic structures, and geometric properties. | General purpose nanomaterial modeling. |
| EMTO-CPA Code [6] | Models disordered alloys within the coherent potential approximation (CPA), essential for realistic alloy nanomaterial simulations. | Alloy formation enthalpy calculations. |
| ACBN0 Pseudo-Hamiltonian [46] | Computes Hubbard U parameters ab initio, reducing empiricism in DFT+U. | Metal oxide band gap correction. |
| MLP Regressor [6] | A class of neural network used as a non-linear regression model to predict continuous properties from material descriptors. | Correcting formation enthalpies and other properties. |
| k-fold Cross-Validation [6] [47] | A resampling procedure used to evaluate ML models on limited data samples, ensuring robustness and mitigating overfitting. | Model validation across all applications. |
Navigating Pitfalls: Strategies for Robust and Transferable ML-DFT Models
In computational chemistry, transferability refers to the ability of a model to make accurate predictions on systems that differ from those it was trained on. This capability is a significant hurdle in integrating machine learning (ML) with electronic structure calculations like Density Functional Theory (DFT). The primary challenge lies in the fact that many ML models experience a substantial drop in performance when applied to larger molecular structures, different basis sets, or alternative exchange-correlation functionals not represented in the training data [48]. Overcoming this challenge is critical for developing robust, general-purpose ML tools that can accelerate quantum chemistry calculations in real-world research and drug discovery applications. This note explores the principles and methodologies for achieving transferable accuracy, with a specific focus on ML-accelerated DFT.
The Core Principles of Transferable ML for DFT
The choice of the target property for the machine learning model is paramount for achieving transferability. Predictions can fail on unseen systems due to numerical instability or the intrinsic non-transferable nature of the target quantity [48].
- Electron Density as a Transferable Target: The electron density, ( \rho(\mathbf{r}) ), is a fundamental physical property. Its local nature makes it highly suitable for ML models. Since the density around an atom is largely influenced by its local chemical environment, a model trained on small molecules can learn local patterns that generalize effectively to larger systems. This approach is more data-efficient and scalable than alternatives [48].
- Challenges with Hamiltonian and Density Matrix Prediction: Many existing methods focus on predicting the Hamiltonian or density matrices. However, the Hamiltonian matrix is "intrinsically non-transferable," and both it and the density matrix can suffer from numerical instability. Small prediction errors in individual matrix elements can be magnified into large, physically nonsensical errors for the system as a whole, hindering application on larger molecules [48].
Table 1: Comparison of ML Prediction Targets for DFT Acceleration
| Prediction Target | Transferability | Scalability | Numerical Stability | Key Advantage |
|---|---|---|---|---|
| Electron Density (in auxiliary basis) | High | Linear with system size | High | Local, fundamental property; compact representation [48] |
| Hamiltonian Matrix | Low | Quadratic with system size | Low (small errors magnified) | Directly used in SCF [48] |
| Density Matrix | Low (basis-set dependent) | Quadratic with system size | Low (especially with diffuse functions) [48] | Directly used in SCF [48] |
Quantitative Performance of Transferable Methods
Recent research demonstrates that an electron-density-centric approach can successfully address the transferability challenge. One study trained an E(3)-equivariant neural network to predict electron density using a compact auxiliary basis representation exclusively on small molecules (up to 20 atoms). When applied to significantly larger systems (up to 60 atoms), the model achieved an average reduction of 33.3% in self-consistent field (SCF) cycles required for convergence. This level of acceleration remained nearly constant with increasing system size, showing remarkable transferability across different orbital basis sets and exchange-correlation functionals [48].
Table 2: Performance of a Transferable Electron Density Model
| Training System Size | Test System Size | Average SCF Reduction | Transferability Across Basis Sets | Transferability Across XC Functionals |
|---|---|---|---|---|
| Up to 20 atoms | Up to 60 atoms | 33.3% | Strong | Strong [48] |
Beyond DFT acceleration, the transferability principle is also being validated in other domains, such as developing Machine Learning Interatomic Potentials (ML-IAPs). The three-step validation approach—assessing basic accuracy/efficiency, benchmarking key properties, and testing on complex defects like dislocations and cracks—highlights that low RMSE on a test set does not automatically guarantee transferability. Model performance must be rigorously validated on the specific, complex systems of interest [49].
Experimental Protocol: A Transferable Workflow for DFT Acceleration
This protocol details the methodology for employing an ML-predicted electron density to generate a high-quality initial guess for SCF calculations, based on the paradigm-shifting work of Liu et al. [48].
The following diagram illustrates the comparative workflow between a traditional SCF procedure and the ML-accelerated approach with a transferable electron density prediction.
Step-by-Step Procedure
Step 1: Data Generation and Model Training
- Dataset Curation: Assemble a dataset of reference molecules and their corresponding electron densities. The
SCFbenchdataset, which includes molecules composed of up to seven different elements, serves as an example [48]. - Target Representation: Instead of the density on a real-space grid, compute the expansion coefficients ( ck ) of the electron density in a compact auxiliary basis ( {\chik(\mathbf{r})} ) (e.g.,
def2-universal-jfitor an even-tempered basis). The density is approximated as ( \rho(\mathbf{r}) \approx \tilde{\rho}(\mathbf{r}) = \sumk ck \chi_k(\mathbf{r}) ) [48]. - Model Selection and Training: Train an E(3)-equivariant neural network to predict the auxiliary basis coefficients ( c_k ) from the molecular structure (atomic numbers and coordinates). The training should be performed exclusively on small molecules (e.g., up to 20 atoms) [48].
Step 2: Initial Guess Generation for a New System
- Input Preparation: For a new, unseen molecular system, provide its atomic numbers, coordinates, and specify the orbital and auxiliary basis sets.
- Density Prediction: Pass the molecular structure through the pre-trained E(3)-equivariant network to obtain the predicted auxiliary coefficients ( {c_k} ) for the electron density.
- Hamiltonian Construction: Use the predicted density coefficients to directly construct the initial Kohn-Sham Hamiltonian matrix, ( \mathbf{H} ).
- The Coulomb matrix (( \mathbf{J} )) is computed efficiently from the coefficients ( {ck} ) using the density fitting approximation.
- The exchange-correlation matrix (( \mathbf{V}{XC} )) is evaluated using the predicted electron density and its gradient, which are readily available from the auxiliary basis expansion. This makes the approach suitable for generalized gradient approximation (GGA) functionals [48].
Step 3: Running the SCF Calculation
- SCF Initialization: Begin the SCF cycle using the ML-predicted Hamiltonian to generate the initial density matrix.
- Convergence: Proceed with the standard SCF iteration (( \mathbf{D} \rightarrow \mathbf{H} \rightarrow \mathbf{C'} \rightarrow \mathbf{D'} )) until convergence is achieved. The high quality of the ML-generated initial guess results in a significantly reduced number of SCF iterations [48].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools for Transferable ML-DFT
| Tool / Reagent | Type | Function in the Workflow |
|---|---|---|
| SCFbench Dataset [48] | Dataset | Provides electron density coefficients for molecules of various sizes and elements, enabling model training and benchmarking. |
| E(3)-Equivariant Neural Network [48] | Software Model | The core architecture that learns to predict electron density in a way that respects physical symmetries (rotations, translations, reflections). |
Auxiliary Basis Set (e.g., def2-universal-jfit) [48] |
Basis Set | Provides a compact, atom-centered representation for expanding the electron density, crucial for efficiency and transferability. |
| Density Fitting Approximation [48] | Mathematical Method | Enables efficient computation of the Coulomb matrix directly from the predicted electron density coefficients. |
| PySCF [48] | Quantum Chemistry Package | A popular Python library used to perform the underlying DFT calculations, including the SCF cycle and integral computation. |
| Model Uncertainty Quantification [49] | Analysis Method | Helps assess the reliability of ML model predictions on new systems, flagging when a prediction might be unreliable. |
The integration of large-scale Density Functional Theory (DFT) datasets with machine learning (ML) workflows is revolutionizing computational materials science and drug discovery. This synergy addresses one of the most significant bottlenecks in traditional DFT calculations: their computational expense, which limits routine calculations to systems of a few hundred atoms [26]. ML models trained on comprehensive DFT datasets can achieve near-DFT accuracy at a fraction of the computational cost, enabling predictions at previously inaccessible scales [15]. The emergence of high-quality, chemically diverse datasets calculated at high levels of DFT theory is foundational to developing robust, generalizable ML interatomic potentials (MLIPs) and electronic structure models. These resources are paving the way for accelerated discovery in fields ranging from catalyst design to battery development and molecular drug discovery [50] [4].
The predictive power of ML models in materials science is fundamentally constrained by the quality and scope of the underlying training data [51]. Several recent datasets exemplify the trend toward larger volumes, improved chemical diversity, and higher levels of DFT theory.
Table 1: Key Characteristics of Prominent DFT Datasets for MLIP Training
| Dataset Name | DFT Level | System Types | Approx. Size | Element Coverage | Key Features |
|---|---|---|---|---|---|
| MP-ALOE [52] | r2SCAN meta-GGA | Primarily off-equilibrium crystals | ~1M calculations from 303k relaxations | 89 elements | Broad pressure/force sampling; Active learning generation |
| OMol25 [50] | ωB97M-V/def2-TZVPD | Molecules, biomolecules, metal complexes | 83M unique molecular systems | 83 elements (H-Bi) | Extensive chemical diversity; Charge/spin states |
| MatPES [52] | r2SCAN meta-GGA | Near-equilibrium solids | Not Specified | Not Specified | Sampled from 300K MD trajectories |
| Compact Datasets [53] | PBE GGA | Various solids | ~4,000 structures avg. | Most of periodic table | Curated for minimalism and high transferability |
These datasets highlight a strategic movement beyond the pervasive Perdew-Burke-Ernzerhof (PBE) generalized gradient approximation (GGA). Meta-GGA functionals like r2SCAN systematically improve over GGAs, reducing mean absolute errors (MAEs) in solid-state formation enthalpies from approximately 150 meV/atom to about 100 meV/atom [52]. For molecular systems, high-level, range-separated hybrid functionals like ωB97M-V provide superior accuracy, particularly for properties involving electronic excitations or non-covalent interactions [50].
Experimental Protocols for Leveraging DFT Datasets
The effective utilization of these datasets requires structured methodologies. The following protocols outline standard workflows for training and benchmarking ML models.
Protocol 1: Training a Universal Machine Learning Interatomic Potential (MLIP)
Application Note: This protocol describes the procedure for training a universal MLIP on a solid-state dataset like MP-ALOE, enabling large-scale molecular dynamics simulations with near-DFT accuracy [52].
Materials:
- Hardware: High-performance computing (HPC) cluster with multiple GPUs.
- Software: MLIP code (e.g., MACE [52]), DFT calculation code (e.g., VASP, Quantum ESPRESSO), data processing scripts.
- Data: A large-scale DFT dataset (e.g., MP-ALOE, MatPES) containing calculated energies, atomic forces, and stresses.
Procedure:
- Data Curation and Combination: If combining multiple datasets (e.g., MP-ALOE and MatPES), ensure compatibility of DFT parameters and functional. Filter out structures with positive cohesive energies if they are not relevant to the target application [52].
- Data Splitting: Partition the data into training, validation, and test sets using a strategy that assesses extrapolation capability, such as splitting by chemical composition or crystal structure family [50].
- Model Selection and Configuration: Choose an MLIP architecture, such as MACE (Message Passing with Equivariant Embeddings) or a Graph Neural Network. Configure model hyperparameters (e.g., interaction blocks, feature dimensions, radial basis functions).
- Model Training: Execute the training job on an HPC cluster. The loss function (L) is typically a weighted sum of errors in energy, forces, and stress:
L = λ_E * (E_pred - E_DFT)² + λ_F * Σ|F_pred - F_DFT|² + λ_S * (S_pred - S_DFT)²Monitor the loss on the validation set to avoid overfitting. - Model Validation: Benchmark the trained potential on a series of tasks:
- Equilibrium Properties: Calculate formation energies of stable crystals and compare to DFT [52].
- Off-Equilibrium Forces: Evaluate force error on far-from-equilibrium structures [52].
- MD Stability: Run molecular dynamics at high temperatures/pressures to test for instability or unphysical behavior [52].
Protocol 2: Correcting DFT Thermodynamics with Machine Learning
Application Note: This protocol uses a smaller, targeted dataset to train an ML model that corrects systematic errors in DFT-calculated formation enthalpies, improving the prediction of phase stability [6].
Materials:
- Software: Python with ML libraries (e.g., Scikit-learn, PyTorch).
- Data: A curated dataset of DFT-calculated and experimentally measured formation enthalpies for a class of materials (e.g., binary and ternary alloys).
Procedure:
- Feature Engineering: For each material, construct a feature vector that encapsulates chemical identity and interactions. This may include:
- Target Definition: The ML model's target is the error between DFT and experiment:
ΔH_f = H_f(DFT) - H_f(Experiment). - Model Training: Train a neural network (e.g., a multi-layer perceptron) to learn the mapping from the feature vector to
ΔH_f. Use cross-validation (e.g., k-fold or leave-one-out) to optimize hyperparameters and prevent overfitting, which is crucial for small datasets [6]. - Prediction and Correction: For a new material, calculate
H_f(DFT)and use the ML model to predict the errorΔH_f_pred. The corrected enthalpy is:H_f(corrected) = H_f(DFT) - ΔH_f_pred.
Protocol 3: Predicting Electronic Structures at Large Scales
Application Note: This protocol employs the Materials Learning Algorithms (MALA) package to predict the local electronic structure, enabling the calculation of observables for systems of >100,000 atoms [26].
Materials:
- Software: MALA software package (interfaces with LAMMPS, PyTorch, and Quantum ESPRESSO) [26].
- Data: DFT-calculated Local Density of States (LDOS) on real-space grids for a training set of small simulation cells (~100-200 atoms).
Procedure:
- Descriptor Calculation: Use LAMMPS to compute bispectrum coefficients for points on the real-space grid. These descriptors encode the atomic environment around each point [26].
- Network Training: Train a feed-forward neural network to map the bispectrum descriptors to the LDOS at a given energy value for each grid point [26].
- Inference on Large Systems: For a large atomic system, calculate bispectrum descriptors for all points in its real-space grid. Pass these through the trained network to predict the full LDOS.
- Post-processing: Use Quantum ESPRESSO utilities within MALA to reconstruct key observables from the predicted LDOS, including the electron density, density of states, and total free energy [26].
Workflow Visualization
The following diagram illustrates the integrated computational workflow for large-scale electronic structure prediction, as implemented in protocols 1 and 3.
ML Workflow for Large-Scale Simulation
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Software and Data Resources for DFT-ML Research
| Tool Name | Type | Primary Function | Relevance to DFT-ML Workflows |
|---|---|---|---|
| MACE [52] | Software / ML Model | A state-of-the-art MLIP architecture. | Used for training universal MLIPs on datasets like MP-ALOE; provides high accuracy and data efficiency [52]. |
| MALA [26] | Software Package | An end-to-end workflow for ML-based electronic structure prediction. | Predicts the local density of states and derived properties for very large systems, circumventing DFT's scaling limit [26]. |
| MP-ALOE [52] | Dataset | A dataset of ~1M r2SCAN calculations. | Provides high-quality, off-equilibrium data for training highly transferable MLIPs across the periodic table [52]. |
| OMol25 [50] | Dataset | A massive molecular dataset at the ωB97M-V level. | Enables training of generalizable ML models for molecular properties, drug discovery, and catalysis [50]. |
| Quantum ESPRESSO [26] | Software Suite | A popular open-source package for DFT calculations. | Often used to generate reference data and for post-processing in ML workflows (e.g., in MALA) [26]. |
| Active Learning [52] | Methodology | A sampling technique to iteratively improve datasets. | Key to building MP-ALOE; used to systematically augment data in uncertain regions of chemical space [52]. |
Mitigating DFT's Inherited Biases for Complex Chemical Systems
Density Functional Theory (DFT) has become an indispensable computational tool for predicting material properties and reaction mechanisms across chemistry, materials science, and drug development. Despite its widespread success, DFT possesses inherent limitations that introduce systematic biases into computational predictions. These biases stem from approximations in the exchange-correlation functionals, which can lead to significant errors in formation enthalpies, reaction barriers, and electronic properties [6] [54]. For complex chemical systems such as ternary alloys, transition metal complexes, and organic reaction pathways, these errors become particularly problematic, limiting DFT's predictive reliability in high-stakes applications like catalyst design and pharmaceutical development.
The integration of machine learning (ML) with DFT has emerged as a transformative approach for mitigating these inherited biases. By leveraging data-driven corrections, researchers can now address systematic errors in DFT calculations while maintaining computational efficiency. This application note outlines structured methodologies and protocols for implementing ML-corrected DFT workflows, providing researchers with practical tools to enhance predictive accuracy across diverse chemical domains. We focus on three principal strategies: error-correcting models, functional-correcting approaches, and consensus frameworks that collectively offer a pathway to more reliable computational predictions [6] [55] [56].
Functional-Driven and Density-Driven Errors
DFT errors can be systematically categorized and quantified to enable targeted corrections. The total error in DFT calculations can be decomposed into two primary components: functional error (ΔEfunc) arising from imperfections in the exchange-correlation functional, and density-driven error (ΔEdens) resulting from inaccuracies in the self-consistent electron density [54]. This decomposition is formally represented as:
ΔE = EDFT[ρDFT] - E[ρ] = ΔEdens + ΔEfunc
where EDFT[ρDFT] is the energy computed with the self-consistent DFT density, and E[ρ] is the exact energy for that density [54]. For challenging chemical systems, both error components can contribute significantly to overall uncertainties. For example, in organic reactions involving bond formation and cleavage, functional errors of 8-13 kcal/mol have been observed even with modern hybrid functionals like ωB97X-D and B3LYP-D3 [54].
System-Specific Error Manifestations
The magnitude and nature of DFT biases vary considerably across chemical systems. In transition metal complexes (TMCs), properties such as spin-state splitting energies (ΔEH-L) show extreme sensitivity to the choice of functional, with variations exceeding 50 kcal/mol across different density functional approximations (DFAs) [56]. For alloy formation enthalpies, intrinsic energy resolution errors particularly affect ternary phase stability calculations, where errors in formation enthalpies can alter predicted stable phases [6]. In molecular datasets used for machine learning interatomic potentials (MLIPs), force component errors averaging 1.7-33.2 meV/Å have been identified, potentially propagating into trained ML models [57].
Table 1: Quantitative Analysis of DFT Errors Across Chemical Systems
| Chemical System | Property | Error Magnitude | Primary Error Source |
|---|---|---|---|
| Organic Reactions [54] | Reaction Energy | 8-13 kcal/mol | Functional approximation |
| Ternary Alloys [6] | Formation Enthalpy | Significant for phase stability | Intrinsic energy resolution |
| Transition Metal Complexes [56] | Spin-State Splitting | >50 kcal/mol variation | HF exchange fraction |
| Molecular Datasets [57] | Force Components | 1.7-33.2 meV/Å | Numerical convergence |
| Main Group Chemistry [54] | Barrier Heights | 2-5 kcal/mol | Density-driven errors |
Machine Learning Correction Strategies
Error-Correcting Models
The error-correcting approach trains ML models to predict the discrepancy between DFT-calculated and reference values (either experimental or high-level computational). This strategy has been successfully applied to improve formation enthalpy predictions for binary and ternary alloys. The implementation typically involves:
A neural network model (e.g., multi-layer perceptron) trained to predict the discrepancy between DFT-calculated and experimentally measured enthalpies for binary and ternary alloys and compounds [6]. The model utilizes a structured feature set comprising elemental concentrations, atomic numbers, and interaction terms to capture key chemical and structural effects. Input features are normalized to prevent variations in scale from affecting model performance [6].
The training process employs rigorous validation techniques including leave-one-out cross-validation (LOOCV) and k-fold cross-validation to prevent overfitting, which is particularly important when working with limited experimental datasets [6]. For the Al-Ni-Pd and Al-Ni-Ti systems relevant to high-temperature applications, this approach has demonstrated significant improvements in predicting phase stability [6].
Functional-Correcting Approaches
Functional-correcting methods employ ML to directly improve the exchange-correlation functional itself, creating ML-corrected density functional approximations. This approach has been demonstrated for popular functionals like B3LYP, where an ML model learns the deviation between the approximate functional and the exact exchange-correlation functional [55].
The key innovation in this approach is the focus on absolute energies rather than energy differences during training, which eliminates reliance on error cancellation between chemical species [55]. The ML model represents a density-dependent correction term that bridges the approximate functional and the exact functional:
EXC^exact[ρ] = EXC^DFA[ρ] + E_ML[ρ]
This strategy involves a double-cycle protocol that incorporates self-consistent-field calculations into the training workflow, ensuring self-consistency between the electron density and the ML correction [55]. Numerical tests demonstrate that ML-corrected functionals trained solely on absolute energies improve accuracy for both thermochemical and kinetic energy calculations, offering a versatile alternative to standard DFAs [55].
Multi-DFA Consensus Methods
For systems where reference data is scarce, consensus approaches across multiple density functional approximations provide an effective strategy for mitigating individual functional biases. This method involves:
Property evaluation across 23+ representative DFAs spanning multiple rungs of Jacob's ladder, from semi-local to double hybrids, to quantify DFA dependence [56]. Although absolute property values differ significantly across functionals, high linear correlations generally persist between DFA pairs, enabling robust comparative analysis.
Artificial neural network (ANN) models are trained independently for each DFA, then used to predict properties for large chemical libraries [56]. By requiring consensus across ANN-predicted DFA properties, researchers can identify compounds with robust property predictions that are invariant to functional choice.
This approach has demonstrated improved correspondence with experimental observations for transition metal complexes, particularly for spin-state splitting energies where DFA dependence is most pronounced [56].
Table 2: Machine Learning Strategies for DFT Bias Mitigation
| Strategy | Mechanism | Best-Suited Applications | Key Advantages |
|---|---|---|---|
| Error-Correcting Models [6] | Predicts DFT-reference discrepancy | Formation enthalpies, phase stability | Direct experimental alignment |
| Functional-Correcting [55] | Learns XC functional deviation | Broad thermochemistry, reaction barriers | Self-consistent, transferable |
| Multi-DFA Consensus [56] | Consensus across functionals | Transition metal complexes, novel materials | No high-level reference needed |
| Δ-Machine Learning [55] | Corrects specific DFA outputs | Targeted property improvement | Preserves physical constraints |
Experimental Protocols
Protocol 1: ML Correction for Formation Enthalpies
This protocol outlines the procedure for implementing ML corrections to DFT-calculated formation enthalpies of alloys and compounds, adapted from established methodologies [6].
Data Curation and Feature Engineering
- Reference Data Collection: Compile a dataset of experimentally measured formation enthalpies for binary and ternary compounds. Filter out unreliable or missing values to ensure data quality.
- DFT Calculations: Perform DFT total energy calculations for all compounds and their constituent elements in ground-state structures using consistent computational parameters (e.g., EMTO-CPA method with GGA-PBE functional [6]).
- Feature Construction: For each compound, compute the following features:
- Elemental concentration vector: x = [xA, xB, x_C, ...]
- Weighted atomic numbers: z = [xA·ZA, xB·ZB, xC·ZC, ...]
- Interaction terms: xi·xj·|Zi - Zj| for element pairs
- Target Variable Calculation: Compute the target correction as ΔHf = Hf^DFT - H_f^experimental
Model Training and Validation
- Data Normalization: Apply standard scaling to all input features to prevent scale-related biases.
- Network Architecture: Implement a multi-layer perceptron (MLP) with three hidden layers using ReLU activation functions.
- Validation Strategy: Employ leave-one-out cross-validation (LOOCV) and k-fold cross-validation to assess model performance and prevent overfitting.
- Application: Apply the trained model to predict corrections for new DFT-calculated formation enthalpies.
Protocol 2: ML-Corrected Functional Implementation
This protocol details the implementation of an ML-corrected density functional approximation, specifically for B3LYP [55].
Reference Data Generation
- Absolute Energy Computation: Generate highly accurate absolute energies for a diverse set of molecular structures using coupled cluster theory [CCSD(T)] or other high-level methods.
- B3LYP Calculations: Perform self-consistent B3LYP calculations for the same structures to obtain baseline energies and electron densities.
- Target Definition: Define the ML learning target as the difference between the reference energy and B3LYP energy: ΔE = Eref - EB3LYP
Model Training with Double-Cycle Protocol
- Feature Representation: Utilize electron density descriptors as input features for the ML model.
- Double-Cycle Training:
- Inner Cycle: Perform self-consistent field calculations with the current ML-corrected functional.
- Outer Cycle: Update the ML model parameters to minimize the difference between corrected energies and reference energies.
- Convergence Testing: Iterate until energy changes between cycles fall below a predetermined threshold (typically 10^-6 Ha).
Protocol 3: Multi-DFA Consensus Workflow
This protocol describes the implementation of a multi-DFA consensus approach for robust property prediction [56].
Multi-DFA Property Evaluation
- Functional Selection: Choose 10-20 representative DFAs spanning multiple rungs of Jacob's ladder, including GGAs, meta-GGAs, hybrids, and double hybrids.
- Consistent Computational Setup: Perform property calculations for all compounds in the dataset using each DFA with consistent geometries and basis sets.
- Correlation Analysis: Compute Pearson correlation coefficients between all DFA pairs to identify consistent trends.
Consensus Model Implementation
- Individual DFA Models: Train separate artificial neural network models for each DFA to predict properties from chemical features.
- Latent Space Alignment: Implement regularization techniques to maintain comparable latent spaces across different DFA-specific models.
- Consensus Prediction: For new compounds, generate predictions from all DFA-specific models and apply consensus criteria (e.g., mean prediction, agreement within uncertainty thresholds).
- Experimental Validation: Prioritize compounds with strong consensus predictions for experimental verification.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Computational Research Reagents for ML-DFT Workflows
| Tool/Resource | Function | Application Context |
|---|---|---|
| EMTO-CPA Code [6] | DFT calculations for alloys | Formation enthalpy correction |
| Local Natural Orbital CCSD(T) [54] | Gold-standard reference energies | Functional correction training |
| Neural Network MLP Regressor [6] | Error prediction model | Formation enthalpy correction |
| Density Error Estimation [54] | Quantifies density-driven errors | Functional diagnostics |
| Artificial Neural Networks (ANNs) [56] | DFA-specific property prediction | Multi-DFA consensus |
| Graph Neural Networks (GNNs) [58] | Molecular structure representation | Bias-corrected property prediction |
Workflow Integration and Decision Pathways
Successful implementation of ML-corrected DFT requires careful selection of appropriate strategies based on the specific research context. The following workflow diagram outlines the decision process for selecting and implementing the most suitable bias mitigation approach:
The integration of machine learning with density functional theory represents a paradigm shift in addressing the long-standing challenge of DFT biases. The methodologies outlined in this application note—error-correcting models, functional-correcting approaches, and multi-DFA consensus strategies—provide researchers with a comprehensive toolkit for enhancing predictive accuracy across diverse chemical systems. As these techniques continue to mature, we anticipate increased focus on model interpretability, uncertainty quantification, and automated workflow integration. The future of computational materials discovery lies in the synergistic combination of physical principles with data-driven insights, enabling more reliable predictions for complex chemical systems relevant to energy applications, catalysis, and pharmaceutical development.
In the realms of computational chemistry and drug development, the principle of Fit-for-Purpose (FfP) modeling advocates for the careful alignment of model complexity with specific application goals and key questions of interest (QOI). This approach is central to the Model-Informed Drug Development (MIDD) framework, which employs modeling and simulation to enhance drug development efficiency and regulatory decision-making [59]. Within the context of density functional theory (DFT) coupled with machine learning (ML) workflows, FfP principles guide the selection of methodologies—from quantitative structure-activity relationship (QSAR) to complex quantitative systems pharmacology (QSP) models—based on the stage of development and the required predictive accuracy [59]. This document outlines detailed application notes and protocols for implementing FfP modeling in research, providing scientists with structured guidelines and practical tools.
Application Notes
The Strategic Imperative of Fit-for-Purpose Modeling
Fit-for-Purpose modeling provides a strategic blueprint for leveraging modeling tools across the drug development lifecycle, from early discovery to post-market management [59]. Its core objective is to ensure that the chosen model's sophistication and resource demands are justified by the specific decision-making needs at each stage. This prevents both the under-utilization of powerful tools and the wasteful application of overly complex models to simple problems.
In the context of DFT/ML workflows, this philosophy translates to selecting the appropriate level of theory, ML algorithm, and dataset based on the specific material property or reaction mechanism being investigated. For instance, a high-level quantitative systems pharmacology (QSP) model is not fit-for-purpose for early-stage lead compound optimization, just as a double-hybrid DFT functional is not practical for the initial screening of millions of candidate molecules [59] [60].
Quantitative Impact of Method Selection
The selection of computational methods directly impacts predictive accuracy and computational cost. The following table summarizes the performance of different ML and DFT approaches for key tasks, demonstrating the trade-offs inherent in FfP decision-making.
Table 1: Performance Comparison of Different ML and DFT Methodologies
| Application Area | Methodology / Descriptor Type | Reported Performance | Key FfP Considerations |
|---|---|---|---|
| Electrocatalysis - Adsorption Energy Prediction | Gradient Boosting Regressor (GBR) with electronic structure descriptors [8] | Test RMSE = 0.094 eV for CO adsorption on Cu SACs | Optimal for medium-to-large datasets (N~2,669); captures non-linear relationships. |
| Electrocatalysis - Overpotential Prediction | Support Vector Regression (SVR) with physics-informed features [8] | Test R² up to 0.98 with ~200 DFT samples | Highly effective in small-data regimes; requires strong feature design. |
| DFT Error Correction for Alloy Enthalpies | Neural Network (MLP) correction of DFT-calculated enthalpies [6] | Significant improvement over uncorrected DFT or linear models | Fit-for-purpose when experimental reference data is available; reduces systematic DFT errors. |
| Universal Interatomic Potentials | Graph-Network Potentials trained on the MAD dataset [61] | Rivals models trained on 100-1000x larger datasets | Designed for robustness across organic/inorganic systems and diverse configurational space. |
| Transition-Metal Complex (TMC) Property Prediction | Artificial Neural Networks (ANNs) informed by 23 different DFAs [60] | Improved consensus predictions for challenging spin-state energies | Mitigates single-DFA bias; fit-for-purpose for systems with strong electronic correlation. |
Essential Research Reagents and Computational Tools
A Fit-for-Purpose toolkit for DFT/ML workflows comprises carefully selected descriptors, datasets, and software. The table below details key "research reagents" essential for conducting experiments in this field.
Table 2: Key Research Reagent Solutions for DFT/ML Workflows
| Reagent / Resource | Type | Primary Function and FfP Rationale |
|---|---|---|
| Intrinsic Statistical Descriptors (e.g., Magpie) [8] | Descriptor | Enable rapid, low-cost coarse screening of vast chemical spaces; require no DFT calculations. |
| Electronic Structure Descriptors (e.g., d-band center, orbital occupancy) [8] | Descriptor | Encode reactivity for accurate predictions in fine screening; require DFT but offer high interpretability. |
| Geometric/Microenvironmental Descriptors (e.g., local strain, coordination number) [8] | Descriptor | Capture structure-activity relationships in complex environments like supports and interfaces. |
| MAD (Massive Atomic Diversity) Dataset [61] | Dataset | Trains robust, universal interatomic potentials; compact size (<100k structures) reduces training cost while ensuring massive configurational diversity. |
| OMol25 Dataset [62] | Dataset | Provides massive scale (83M systems) for training data-intensive models on molecular systems using consistent, high-quality hybrid DFT data. |
| Custom Composite Descriptors (e.g., ARSC, FCSSI) [8] | Descriptor | Combine multiple physical effects into low-dimensional, interpretable features for specific chemistries (e.g., dual-atom catalysts), reducing model complexity and data needs. |
Experimental Protocols
Protocol 1: Correcting DFT Formation Enthalpies using Machine Learning
This protocol details a methodology to improve the predictive accuracy of density functional theory for alloy formation enthalpies using a neural network-based error correction model [6].
1. Objective: To systematically reduce the error between DFT-calculated and experimentally measured formation enthalpies ((H_f)) for binary and ternary alloys.
2. Materials & Software:
- DFT Code: Exact Muffin-Tin Orbital (EMTO) method in combination with the full charge density technique or similar DFT package [6].
- ML Framework: Python with scikit-learn or a similar library for implementing a Multi-Layer Perceptron (MLP) regressor.
- Training Data: A curated set of binary and ternary alloys with reliably known experimental formation enthalpies.
3. Procedure:
- Step 1: Data Curation and Input Feature Engineering
- Compile a dataset of alloys with known experimental (Hf).
- For each material
i, calculate the DFT-derived (Hf^{DFT}(i)) using Equation 1 (see Appendix). - Define the target variable for ML as the error: (\Delta Hf(i) = Hf^{exp}(i) - Hf^{DFT}(i)).
- For each alloy, construct a feature vector that includes:
- Normalize all input features to a common scale.
Step 2: Model Training and Validation
- Implement an MLP regressor with three hidden layers.
- Use Leave-One-Out Cross-Validation (LOOCV) and k-fold cross-validation to optimize hyperparameters and prevent overfitting.
- Train the model to predict (\Delta H_f) using the structured feature set.
Step 3: Prediction and Validation
- For a new alloy, compute (Hf^{DFT}) and the ML-predicted error (\Delta Hf^{ML}).
- The corrected formation enthalpy is: (Hf^{corrected} = Hf^{DFT} + \Delta H_f^{ML}).
- Validate the model's performance on a hold-out test set of alloys not used in training.
4. Diagram: Workflow for ML-Correction of DFT Enthalpies
Protocol 2: Implementing a Fit-for-Purpose ML Screening for Electrocatalysts
This protocol outlines a tiered screening strategy for electrocatalyst discovery, moving from low-cost coarse screening to high-fidelity refinement [8].
1. Objective: To efficiently identify lead electrocatalyst candidates for reactions like ORR, OER, and CO2RR by leveraging a combination of descriptor types and ML models.
2. Materials & Software:
- Descriptor Libraries: Tools to compute intrinsic elemental properties (e.g., Magpie).
- DFT Software: For calculating electronic structure descriptors.
- ML Algorithms: Gradient Boosting Regressors (GBR) for medium-large data; Support Vector Regression (SVR) for small-data regimes.
3. Procedure:
- Step 1: Coarse Screening with Intrinsic Descriptors
- Define a large search space of candidate materials (e.g., thousands to millions).
- For each candidate, compute low-cost intrinsic statistical descriptors (e.g., elemental composition, valence-orbital information) [8].
- Train a model (e.g., GBR) on available data linking these descriptors to the target property (e.g., adsorption energy).
- Screen the vast search space and identify a narrowed-down set of promising candidates (e.g., a few hundred) for further analysis.
Step 2: Refined Screening with Electronic and Geometric Descriptors
- For the shortlisted candidates, perform DFT calculations to obtain electronic structure descriptors (e.g., d-band center, orbital occupancies) [8].
- If applicable, compute geometric/microenvironmental descriptors (e.g., local coordination environments, strain) [8].
- Train a new, more accurate ML model using these advanced descriptors on a smaller, refined dataset.
- Re-rank the candidates based on predictions from this higher-fidelity model.
Step 3: Validation and Lead Identification
- Select the top-ranked candidates from the refined screening.
- Perform direct, full DFT calculations to validate the predicted properties.
- The final validated candidates constitute the lead compounds for experimental synthesis and testing.
4. Diagram: Tiered Screening Workflow for Electrocatalysts
Adhering to Fit-for-Purpose principles ensures that computational resources are deployed efficiently and that models yield actionable, reliable insights for drug development and materials discovery. By strategically selecting from a toolkit of descriptors—ranging from low-cost intrinsic properties to high-fidelity electronic structure features—and aligning ML algorithms with data availability and task complexity, researchers can construct robust predictive workflows. The protocols outlined herein provide a concrete foundation for implementing these strategies, enabling the acceleration of discovery while maintaining scientific rigor.
Density Functional Theory (DFT) serves as the workhorse for quantum mechanical calculations in materials science and drug discovery, with nearly a third of U.S. supercomputer time dedicated to molecular modeling [4]. However, conventional DFT approximations suffer from a fundamental limitation: the unknown universal form of the exchange-correlation (XC) functional, which describes how electrons interact [4]. This limitation becomes particularly problematic in systems with strong electron correlation—such those with strained chemical bonds, open-shell radicals, diradicals, or metal-organic bonds to open-shell transition-metal centers—where standard DFT functionals often yield inaccurate results or fail completely [63].
The emergence of machine learning (ML) has introduced powerful new approaches to these challenges. ML-accelerated discovery workflows, however, inherit the biases of their DFT training data and frequently attempt calculations destined for failure [63]. This combinatorial challenge necessitates a robust filtering mechanism. "Decision engines" represent a sophisticated class of ML models that act as this crucial filter, predicting the likelihood of DFT calculation success and diagnosing the presence of strong correlation before computationally expensive simulations are launched [63]. By enabling rapid diagnoses and adaptation strategies, these systems form the foundation for autonomous workflows that minimize expert intervention and maximize research efficiency in computational chemistry and drug development.
Quantitative Performance of Decision Engine Models
The performance of various ML approaches for building decision engines can be evaluated based on their accuracy, computational cost, and applicability. The table below summarizes key quantitative findings from different methodologies.
Table 1: Performance Metrics of ML-Enhanced DFT and Diagnostic Models
| Model / Approach | Key Performance Metric | Training Data | Computational Efficiency | Limitations / Scope |
|---|---|---|---|---|
| ML-Enhanced XC Functional [4] | Outperformed/matched widely used XC approximations | Exact energies and potentials from QMB calculations for 5 atoms & 2 simple molecules | Kept computational costs in check | Preliminary testing; effective for light atoms, expansion to solids needed |
| Decision Engine Workflow [63] | Enabled quantitative sensitivity analysis; predicted calculation failure | Multiple DFT method results and calculation outcomes | Reduced failed calculations in high-throughput screens | Requires series of tests for trustworthiness; aims for autonomous workflows |
| LSTM Network for Decision Prediction [64] | Predicted target selection decisions preceding conscious human intent; expertise-specific models | Sequences of herder and target state input features | Not specified | Model expertise-specific (expert vs. novice); requires sequential input data |
These models demonstrate a common theme: achieving high accuracy and computational efficiency by being trained on compact, high-quality datasets. The ML-enhanced XC functional, for instance, achieved its striking accuracy using data from only five atoms and two simple molecules [4]. This principle is central to decision engines, which must be lightweight enough to provide rapid diagnostics without becoming a computational bottleneck themselves.
Experimental Protocols for Decision Engine Development and Validation
Protocol: Developing an ML-Enhanced Exchange-Correlation Functional
This protocol is adapted from research that used machine learning to discover more universal XC functionals, bridging the accuracy of quantum many-body (QMB) methods with the simplicity of DFT [4].
Data Acquisition from QMB Calculations:
- Perform exact QMB calculations on a small set of simple, representative systems (e.g., 5 atoms and 2 simple molecules).
- Extract two key types of data for training:
- Interaction energies of electrons.
- Potentials that describe how that energy changes at each point in space. The inclusion of potentials provides a stronger foundation for training by highlighting small differences in systems more clearly than energies alone [4].
Model Training:
- Train a machine learning model (e.g., a neural network) to approximate the XC functional using the compact dataset of exact energies and potentials.
- The objective is for the ML model to learn a mapping from the electron density to a highly accurate XC functional.
Validation and Testing:
- Use the newly developed ML-learned functional in DFT calculations for systems outside the training set.
- Evaluate performance by comparing the results against high-accuracy QMB benchmarks for accuracy and against standard DFT functionals for computational cost.
- Assess the functional's ability to avoid producing unphysical or meaningless results, a common limitation of earlier ML models [4].
Protocol: Building a Decision Engine for Calculation Success and Strong Correlation
This protocol outlines the steps for creating an ML model that predicts the robustness of a DFT calculation for a given material system [63].
Curate a Benchmark Dataset:
- Assemble a diverse dataset of molecular or material systems where the outcomes of DFT calculations are well-characterized.
- Each data point must be labeled with:
- Calculation Success/Failure: Whether a standard DFT calculation converges to a physically meaningful result.
- Correlation Strength: An indicator of the strength of electron correlation (e.g., categorized as "weak," "strong," or "multireference").
- This labeling often requires expert knowledge or higher-level quantum chemistry methods for ground truth.
Feature Engineering:
- Compute a set of input features (descriptors) for each system that the ML model will use to make its prediction. Key features may include:
- Electronic features: Molecular composition, orbital occupations, spin states, and symmetry.
- Geometric features: Bond lengths, angles, and coordination numbers.
- Pre-calculation metrics: Simple, low-cost quantum chemical descriptors that can be computed before a full DFT simulation.
- Compute a set of input features (descriptors) for each system that the ML model will use to make its prediction. Key features may include:
Model Training and Selection:
- Train a suite of supervised ML classifiers (e.g., Decision Trees, Random Forests, or Gradient Boosting machines) on the benchmark dataset.
- The model's task is a two-step classification: first, predicting the probability of calculation success, and second, diagnosing the likelihood of strong correlation.
- Use cross-validation to prevent overfitting and to select the model with the best generalizability.
Workflow Integration and Autonomous Operation:
- Integrate the trained decision engine into a high-throughput computational screening workflow.
- Configure the workflow so that every new material candidate first passes through the decision engine.
- Based on the model's prediction:
- If the candidate has a high probability of success and low correlation, proceed with standard DFT.
- If the candidate is flagged for probable failure or strong correlation, the workflow can automatically trigger an adaptation strategy, such as using a more robust (but expensive) quantum chemistry method or a specialized DFT functional [63].
Workflow Visualization
The following diagram illustrates the integrated autonomous workflow for DFT calculations, incorporating the decision engine as a critical gating mechanism.
Diagram 1: Autonomous DFT workflow with a decision engine. The ML model routes calculations based on predicted success and correlation diagnosis.
The logical structure of the decision engine itself, which powers the workflow above, can be broken down into its core analytical steps as shown in the diagram below.
Diagram 2: Decision engine's internal logic for diagnosing calculation robustness.
The Scientist's Toolkit: Essential Research Reagents
Implementing decision engines and ML-accelerated DFT workflows requires a suite of computational "reagents." The following table details these essential components.
Table 2: Key Research Reagent Solutions for ML-DFT Workflows
| Research Reagent | Function / Explanation | Examples / Notes |
|---|---|---|
| High-Quality Benchmark Datasets | Serves as the ground truth for training and validating ML models. The data quality dictates model performance. | Data from highly accurate QMB methods [4]; datasets covering diverse chemical spaces including transition metals and solids [65]. |
| Feature Descriptor Libraries | Translates chemical structures into numerical inputs that ML models can process. | Electronic structure descriptors (e.g., orbital occupations, electron density moments); composition-based features; geometric descriptors. |
| ML Model Architectures | The core algorithm that learns the complex relationships between material features and DFT outcomes. | Graph Neural Networks for molecular structures [66]; LSTMs for sequential data [64]; ensemble methods like Random Forests for robust classification. |
| Δ-Machine Learning (Δ-ML) | A technique where ML learns the difference (Δ) between a low-level and high-level method, refining DFT results towards wavefunction accuracy at a lower cost [65]. | Used to create correction models that bridge the gap between approximate and accurate quantum methods. |
| Causal AI Techniques | Helps move beyond correlation to identify true cause-and-effect relationships in data, which is crucial for reliable trial design and understanding biological mechanisms [67]. | Emerging as a tool to uncover the true drivers of disease progression and drug response, potentially improving success rates in drug development [67]. |
| Explainable-AI (XAI) Tools | Makes the predictions of "black-box" ML models interpretable to scientists, building trust and providing insights. | SHAP (SHapley Additive exPlanations) [64]; LIME. Critical for understanding which features led to a diagnosis of strong correlation or predicted failure. |
Decision engines represent a transformative advancement in the pursuit of robust and autonomous computational materials and drug discovery. By leveraging machine learning to predict calculation success and diagnose strong correlation, these systems directly address the combinatorial challenge and inherent biases of traditional high-throughput DFT screening. The integration of ML-enhanced XC functionals and diagnostic models into a cohesive, adaptive workflow, as visualized in this document, promises to significantly accelerate research while ensuring greater reliability. As these tools mature, fueled by high-quality data and advanced AI techniques like causal inference and explainable AI, they will move the field closer to fully autonomous discovery cycles, empowering researchers and drug developers to explore complex chemical spaces with unprecedented confidence and efficiency.
Benchmarking Performance: Validating ML-DFT Accuracy and Computational Efficiency
Density functional theory (DFT) is a cornerstone of computational chemistry, materials science, and drug development, enabling the simulation of molecular and material properties at the quantum mechanical level. However, its accuracy is inherently limited by approximations in the exchange-correlation (XC) functional, which describes how electrons interact [4] [68]. Machine learning (ML) is now revolutionizing computational chemistry by integrating with DFT to enhance its predictive power, offering pathways to chemical accuracy while maintaining manageable computational costs [68] [15].
This application note provides a structured benchmark of ML-accelerated DFT (ML-DFT) methodologies against standard quantum chemistry methods. We summarize quantitative performance data, detail experimental protocols for key implementations, and visualize the core benchmarking workflow to equip researchers with the tools needed to adopt these advanced computational strategies.
Performance Benchmarking Tables
The table below summarizes key quantitative benchmarks comparing ML-DFT approaches to traditional methods on various chemical tasks.
Table 1: Performance Benchmarks of ML-DFT Methods vs. Standard Quantum Chemistry Approaches
| Method / Model | Reference Method | System / Property Tested | Reported Accuracy Metric | Key Performance Result |
|---|---|---|---|---|
| ML-XC Functional [4] [69] | QMB Methods | Light atoms & small molecules (Energy/Potential) | Accuracy vs. QMB | Achieved 3rd-rung DFT accuracy at 2nd-rung computational cost [4] [69] |
| OMol25-trained NNPs (eSEN, UMA) [70] | ωB97M-V/def2-TZVPD (DFT) | Diverse molecular energies (GMTKN55) | WTMAD-2 | Matched or exceeded the high-accuracy DFT reference level [70] |
| EMFF-2025 (NNP) [7] | DFT | CHNO-based Energetic Materials (Energy/Forces) | Mean Absolute Error (MAE) | Energy MAE: < 0.1 eV/atom; Force MAE: < 2 eV/Å [7] |
| Δ-ML (PM6-ML) [71] | MP2/def2-TZVP | Proton Transfer Reactions (Relative Energies) | Mean Unsigned Error (MUE) | MUE = 10.8 kJ/mol (vs. 20.3 kJ/mol for base PM6) [71] |
| Traditional DFT (B3LYP) [71] | MP2/def2-TZVP | Proton Transfer Reactions (Relative Energies) | Mean Unsigned Error (MUE) | MUE = 7.44 kJ/mol [71] |
| Multifidelity ΔML [72] | Coupled Cluster | Organic Molecules (Energies) | Data Efficiency & Accuracy | Outperformed standard Δ-ML for a limited number of predictions [72] |
The table below benchmarks the computational efficiency and data requirements of different approaches.
Table 2: Computational Efficiency and Data Requirements of ML-DFT Models
| Model / Approach | Training Data Scale & Source | Computational Cost | Transferability / Generality Claim |
|---|---|---|---|
| ML-XC Functional [4] | Minimal data (5 atoms, 2 molecules) from QMB | Low (2nd-rung cost, 3rd-rung accuracy) | Accurate for systems different from training data [4] |
| OMol25-based NNPs [70] | Massive (100M+ calculations, ωB97M-V) | High training cost, fast inference | Exceptional chemical diversity (biomolecules, electrolytes, metal complexes) [70] |
| EMFF-2025 [7] | Transfer learning from pre-trained model | DFT-level accuracy, higher efficiency than ReaxFF | General purpose for CHNO HEMs (mechanical & chemical properties) [7] |
| Multifidelity Methods [72] | Multi-level datasets (e.g., QeMFi) | Reduced high-fidelity data needs | Effective knowledge transfer across fidelities; good for diverse predictions [72] |
Experimental Protocols
Protocol 1: Developing a Machine-Learned XC Functional
This protocol outlines the methodology for using machine learning to derive a more universal exchange-correlation functional, as demonstrated by Gavini et al. [4] [69].
Key Resources:
- High-Fidelity Reference Data: Generate quantum many-body (QMB) data for a small set of light atoms (e.g., Li, C, N, O, Ne) and simple molecules (e.g., H₂, LiH) using highly accurate but computationally expensive methods.
- Training Targets: Extract not just the total interaction energies from the QMB calculations, but also the quantum potentials, which describe how the energy changes at each point in space. Using potentials in training provides a stronger foundation and helps the model capture subtle changes more effectively [4].
- ML Model Training: Train a machine learning model (e.g., a neural network) to learn the mapping from the electron density to the XC functional. The model is trained to output the functional that, when used in a DFT calculation, reproduces the QMB energies and potentials.
- Validation: Apply the newly learned ML-XC functional in DFT calculations for systems outside its training set to validate its accuracy and transferability. Benchmark its performance against both standard DFT approximations and higher-level QMB methods [4].
Protocol 2: Training a General Neural Network Potential on a Large-Scale Dataset
This protocol describes the workflow for creating state-of-the-art neural network potentials (NNPs), exemplified by the OMol25 and UMA initiatives [70].
Key Resources:
- Dataset Curation: Compile a massive and chemically diverse dataset of molecular structures. The OMol25 dataset, for instance, contains over 100 million calculations and covers biomolecules, electrolytes, metal complexes, and more [70].
- High-Accuracy Reference Calculations: Perform quantum chemical calculations for all structures in the dataset at a consistently high level of theory (e.g., the ωB97M-V/def2-TZVPD level used for OMol25) to serve as the ground truth [70].
- Model Architecture and Training:
- Architecture Selection: Use an equivariant or invariant neural network architecture, such as eSEN (equivariant Spectral Embedding Network) or the Universal Model for Atoms (UMA), which respects physical symmetries like rotation and translation [70].
- Conservative Force Training: For more robust molecular dynamics simulations, employ a two-phase training scheme. First, train a model to predict forces directly. Then, use this model to initialize a second model that is fine-tuned to predict forces as the negative gradient of the energy (conservative forces), which improves the physical correctness of the potential energy surface [70].
- Benchmarking: Rigorously test the final NNP on established benchmarks (e.g., GMTKN55, Wiggle150) to confirm it matches the accuracy of its high-level DFT training data across a broad chemical space [70].
Protocol 3: Applying Δ-Machine Learning for Error Correction
This protocol covers the use of Δ-ML to correct the errors of lower-level methods, bringing their accuracy closer to that of high-level reference calculations [72] [71].
Key Resources:
- Multi-Level Data Collection: For a set of molecular structures, compute the target property (e.g., energy) using both a low-level quantum method (e.g., PM6, a low-fidelity DFT functional) and a high-level reference method (e.g., MP2, CCSD(T), or a high-fidelity DFT functional).
- Delta Label Calculation: For each structure, calculate the difference (Δ) between the high-level and low-level property values:
Δ = E_high - E_low. This Δ becomes the target for the ML model to learn [72]. - Model Training: Train an ML model to predict the Δ value based on a representation of the molecular structure.
- Inference and Prediction: For a new, unknown structure, the property is predicted as
E_predicted = E_low + Δ_ML, whereE_lowis computed by the fast, low-level method andΔ_MLis the correction predicted by the ML model. This approach can be extended to multifidelity learning, which uses data from several levels of theory to improve data efficiency [72].
Workflow Visualization
The following diagram illustrates the logical workflow for developing and benchmarking an ML-DFT model, integrating the protocols above.
Diagram 1: ML-DFT Model Development and Benchmarking Workflow. This chart outlines the key stages for creating and validating machine learning models to enhance DFT, from initial data generation to final benchmarking against established quantum chemical methods.
The Scientist's Toolkit
The table below lists key computational reagents and resources essential for implementing the ML-DFT protocols discussed in this note.
Table 3: Essential Research Reagents and Computational Resources for ML-DFT
| Resource / Tool | Type | Primary Function in ML-DFT | Example(s) |
|---|---|---|---|
| High-Accuracy Reference Datasets | Dataset | Serves as ground-truth data for training and benchmarking ML models. | OMo25 [70], QeMFi [72] |
| Pre-trained ML Potentials | Software/Model | Provides ready-to-use, accurate force fields for molecular simulations without training from scratch. | eSEN models, UMA (Universal Model for Atoms) [70], EMFF-2025 [7] |
| Δ-ML & Multifidelity Frameworks | Methodology/Code | Corrects systematic errors of fast, low-level quantum methods towards high-level accuracy. | Multifidelity ΔML [72], PM6-ML [71] |
| Quantum Chemistry Codes | Software | Performs standard DFT and post-Hartree-Fock calculations for data generation and benchmarking. | Codes used for ωB97M-V [70], MP2 [71] |
| ML Potential Architectures | Software/Model | Neural network frameworks designed to respect physical symmetries in atomistic systems. | eSEN [70], Equiformer [70], Deep Potential (DP) [7] |
Achieving Chemical Accuracy with Orders-of-Magnitude Speedup
Density Functional Theory (DFT) has long served as a cornerstone of computational chemistry, enabling the prediction of molecular structures, reaction energies, and spectroscopic properties. Despite its widespread adoption, DFT has historically faced a fundamental trade-off: achieving chemical accuracy often requires computationally expensive functionals and basis sets that limit practical application to large systems or long timescales. The emergence of machine learning (ML) is now disrupting this paradigm by creating new pathways to accuracy that bypass traditional computational bottlenecks.
Recent advances demonstrate that ML models can emulate key aspects of DFT calculations while achieving orders-of-magnitude speedup. By learning the complex mapping between atomic structures and electronic properties directly from quantum mechanical data, these approaches maintain the accuracy of high-level DFT calculations while dramatically reducing computational cost. This application note examines the protocols and methodologies driving these breakthroughs, providing researchers with practical guidance for implementing ML-accelerated DFT workflows.
Current Breakthroughs in ML-Accelerated DFT
Deep Learning for Exchange-Correlation Functionals
The Microsoft Research team developed Skala, a deep learning-derived exchange-correlation functional that demonstrates significantly improved accuracy for small molecules. Trained on approximately 150,000 reaction energies for molecules with five or fewer non-carbon atoms, Skala utilizes architecture inspired by large language models [73].
- Performance Metrics: Skala achieves a prediction error for small-molecule energies that is half that of ωB97M-V, previously considered one of the most accurate functionals available [73].
- Computational Efficiency: The functional maintains computational efficiency comparable to or better than existing high-accuracy functionals [73].
- Current Limitations: While excelling with small organic molecules, Skala shows intermediate performance for metal-containing systems outside its training domain, highlighting the importance of training data composition [73].
End-to-End DFT Emulation
A comprehensive deep learning framework demonstrates full emulation of the Kohn-Sham DFT workflow, mapping atomic structure directly to electronic charge density and derived properties [2].
- Speed Advantage: This approach provides orders-of-magnitude speedup with linear scaling system size (with a small prefactor) while maintaining chemical accuracy [2].
- Property Portfolio: The model simultaneously predicts multiple electronic and atomic properties, including density of states, band gap, potential energy, atomic forces, and stress tensor [2].
- Architecture: The method employs a two-step learning procedure that first predicts electronic charge density using Gaussian-type orbital descriptors, then uses these descriptors to predict other properties [2].
Potential-Enhanced Training
Researchers at the University of Michigan developed an ML approach that incorporates not just electron interaction energies but also the potentials describing how energy changes at each point in space [4].
- Training Advantage: Potentials highlight subtle system differences more effectively than energies alone, enabling more accurate functional approximation [4].
- Data Efficiency: This method achieved high accuracy using a compact training set of just five atoms and two simple molecules [4].
- Transferability: The resulting model generalized well beyond its training set while avoiding unphysical results that plagued earlier ML approaches [4].
Table 1: Comparison of Recent ML-DFT Approaches
| Approach | Key Innovation | Accuracy Improvement | System Scope | Computational Efficiency |
|---|---|---|---|---|
| Skala Functional [73] | Deep learning-derived XC functional | 50% error reduction vs. ωB97M-V | Small molecules (≤5 non-C atoms) | Comparable to conventional functionals |
| End-to-End Emulation [2] | Direct mapping from structure to charge density | Chemical accuracy maintained | Organic molecules, polymer chains/crystals | Orders-of-magnitude speedup, linear scaling |
| Potential-Enhanced Training [4] | Incorporation of energy potentials in training | High accuracy vs. conventional functionals | Light atoms, transferable to new systems | Low training cost, avoids unphysical results |
Experimental Protocols and Methodologies
Protocol for ML-Derived Functional Development
The development of machine learning-enhanced functionals like Skala follows a structured workflow that integrates quantum mechanics, data science, and computational chemistry [73].
Step 1: Training Database Construction
- Curate a diverse set of molecular structures representing the target chemical space
- For Skala, researchers created approximately 150,000 reaction energies for small molecules [73]
- Ensure representation of various bonding environments and element types
Step 2: Reference Data Generation
- Perform high-level DFT or quantum many-body calculations for all structures
- Calculate target properties including reaction energies, electronic properties
- Use consistent computational parameters across all systems
Step 3: Machine Learning Model Development
- Select appropriate deep learning architecture (Skala borrowed from language models) [73]
- Implement training with careful regularization to prevent overfitting
- Validate model performance on held-out test sets
Step 4: Functional Integration and Testing
- Incorporate the learned functional into existing DFT codes
- Perform comprehensive benchmarking against standard datasets
- Evaluate transferability to systems outside training data
Protocol for End-to-End DFT Emulation
The ML-DFT framework demonstrated for organic materials provides a complete protocol for bypassing the Kohn-Sham equations [2].
Step 1: Database Creation with Configurational Diversity
- Assemble structures including molecules, polymer chains, and crystals
- Incorporate configurational diversity through MD snapshots at various temperatures
- For the referenced study: 67 molecules, 178 polymer chains, 55 polymer crystals (118,000+ structures) [2]
Step 2: Fingerprinting Atomic Structures
- Employ atom-centered symmetry functions (e.g., AGNI fingerprints) [2]
- Ensure descriptors are translation, permutation, and rotation invariant
- Represent local chemical environments for each atom
Step 3: Charge Density Prediction
- Train deep neural networks to map atomic fingerprints to charge density
- Use Gaussian-type orbitals as flexible basis functions learned from data
- Transform from internal atomic reference systems to global Cartesian coordinates
Step 4: Property Prediction
- Use predicted charge density as input for subsequent property prediction
- Train separate networks for different properties (energy, forces, DOS, etc.)
- Employ multi-task learning where beneficial
Step 5: Model Validation
- Split data using 90:10 training:test partition with 80:20 training:validation split [2]
- Evaluate performance on independent test sets representing different structure types
- Assess transferability to larger systems than those in training data
Specialized Protocol for Energetic Materials Stability
For predicting bond dissociation energies (BDEs) of energetic materials, a specialized protocol demonstrates high accuracy even with limited data [74].
Step 1: Curate Domain-Specific Dataset
- Collect 778 synthesized CHON-containing energetic molecules from literature [74]
- Calculate reference BDEs at B3LYP/6-31G level of theory
- Ensure diversity in molecular weight (61-998 g/mol), atom count (7-87), and trigger bond types
Step 2: Implement Hybrid Feature Representation
- Couple local target bond features with global molecular structure characteristics
- Capture both bond-specific and overall molecular environment information
Step 3: Apply Data Augmentation
- Utilize Pairwise Difference Regression (PADRE) to expand effective dataset size
- Generate new samples through feature vector and label differences
- Reduce systematic errors while improving model robustness
Step 4: Train Ensemble Models
- Employ XGBoost algorithm for final model implementation
- Achieve reported accuracy of R² = 0.98 and MAE = 8.8 kJ mol⁻¹ [74]
- Significantly outperform models trained on general organic molecules
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Computational Tools and Resources for ML-DFT Research
| Tool/Resource | Type | Function/Purpose | Example Applications |
|---|---|---|---|
| Skala Functional [73] | ML-derived XC functional | Improves accuracy for small molecule energy calculations | Reaction energy prediction for organic molecules |
| AGNI Fingerprints [2] | Atomic descriptor | Represents chemical environment for machine learning | Structure-property mapping in organic materials |
| MatSci-ML Studio [75] | GUI workflow toolkit | Democratizes ML application for materials science | Automated model training for property prediction |
| BDE Dataset for EMs [74] | Specialized database | Enables stability prediction for energetic materials | Bond dissociation energy prediction for explosives safety |
| PADRE Augmentation [74] | Data enhancement method | Alleviates limitations of small datasets in specialized domains | Improving model robustness with limited energetic molecules |
Workflow Visualization
ML-DFT Workflow: Structure to Properties
ML-Driven Functional Development
The integration of machine learning with density functional theory represents a paradigm shift in computational chemistry and materials science. The methodologies outlined in this application note demonstrate that achieving chemical accuracy with orders-of-magnitude speedup is not merely theoretical but practically attainable across multiple domains. From specialized functionals like Skala to comprehensive DFT emulation frameworks, these approaches maintain the accuracy of quantum mechanical calculations while dramatically reducing computational cost.
As these protocols continue to mature and accessible tools like MatSci-ML Studio democratize their application, researchers across chemistry, materials science, and drug development can leverage these advancements to explore larger systems, longer timescales, and more complex phenomena. The continued development of ML-DFT workflows promises to accelerate the discovery of new materials, catalysts, and pharmaceutical compounds while deepening our fundamental understanding of chemical behavior.
In computational materials science and drug discovery, representing electronic charge density and molecular structure is a foundational step for predicting material properties and enabling virtual screening. Within the context of machine learning (ML) enhanced density functional theory (DFT), two dominant paradigms have emerged: grid-based and atom-based density representations. Grid-based methods sample properties like electron density onto a discrete three-dimensional lattice, providing a direct and detailed representation of the spatial field [76]. In contrast, atom-based methods describe the density as a sum of atom-centered basis functions, such as Gaussian-type orbitals (GTOs), offering a more compact and analytically tractable representation [2]. The choice between these representations presents a significant trade-off, impacting the accuracy, computational efficiency, and transferability of ML-DFT workflows. This application note provides a detailed comparative analysis of these two approaches, offering structured protocols and resources to guide researchers in selecting and implementing the appropriate representation for their specific applications in materials research and drug development.
Comparative Analysis: Core Principles and Trade-offs
Foundational Methodologies
- Grid-Based Representations: This approach maps molecular systems or simulation data onto a high-resolution 3D grid. Each grid cell contains averaged property information—such as electron density or atomic density—sampled from numerous configurational snapshots of the system [76]. The resolution must be fine enough to resolve the smallest features of interest (e.g., 0.5 Å for atomistic systems, 1.5-2.0 Å for coarse-grained models). The resulting data is a 3D scalar field that can be directly visualized and analyzed [76].
- Atom-Based Representations: Instead of a fixed grid, this method decomposes the electron density into atomic contributions, typically using a basis set like Gaussian-type orbitals (GTOs) [2]. The model learns the optimal parameters (exponents and coefficients) for these basis functions from reference data. The total electron density is reconstructed by summing the contributions from all atoms, with each atom's density described within its own internal reference frame before being transformed to the global Cartesian system [2].
Quantitative Comparison of Key Characteristics
Table 1: Direct comparison of grid-based and atom-based density representation methods.
| Feature | Grid-Based Representation | Atom-Based Representation |
|---|---|---|
| Fundamental Description | 3D scalar field on a discrete lattice [76] | Sum of atom-centered basis functions [2] |
| Information Completeness | High; captures delocalized densities and complex features directly [2] | Lower; accuracy depends on the chosen basis set, struggles with delocalization [2] |
| Computational Scaling | Linear with system size, but with a large prefactor due to high grid-cell count [76] [2] | Linear with system size, with a small prefactor; highly efficient [2] |
| Data Efficiency | Low; requires hundreds to thousands of snapshots for smooth density maps [76] | High; reduced parameter set enables learning from fewer examples [2] |
| System Transferability | Limited; model trained on small systems may not generalize to larger ones [2] | High; inherent atomic description improves transferability across system sizes [2] |
| Primary Advantage | High accuracy and direct interpretability as a spatial field [76] [2] | Computational speed and efficient scaling for large systems [2] |
| Primary Limitation | High computational cost and storage requirements [2] | Lower accuracy, especially for systems with delocalized electron densities [2] |
| Ideal Application Domain | Detailed analysis of local electronic phenomena; visualization of density distributions [76] | High-throughput screening; molecular dynamics; large-scale systems [2] |
Application Protocols
Protocol A: Implementing a Grid-Based Density Workflow
This protocol describes the process for generating and analyzing grid-based density representations from a molecular dynamics (MD) trajectory, suitable for visualizing structural features in large biological systems or analyzing electron density.
Materials and Software Requirements
- Input Data: A molecular dynamics trajectory file (e.g., in GROMACS XTC or TRR format) [76].
- Simulation Software: GROMACS or similar MD package [76].
- Sampling & Visualization Tool: Custom scripts (e.g., in Python) and ParaView visualization software [76].
Step-by-Step Procedure
System Preparation and Grid Definition
- Obtain an initial configuration and ensure your trajectory contains a sufficient number of snapshots. A good rule of thumb is that the total number of particles multiplied by the number of configurations should be at least ten times the number of grid cells [76].
- Define a 3D grid encompassing the simulation box. Select a grid spacing that resolves the smallest feature of interest (e.g., 0.5 Å for atomistic detail, 1.5-2.0 Å for coarse-grained beads) [76].
Trajectory Sampling and Grid Population
- Run a simulation to generate hundreds or thousands of configurations if not already available. The sampling time should be short enough to prevent diffusion from smearing structural features but long enough for representative sampling [76].
- For each snapshot in the trajectory, map the property of interest (e.g., atomic positions for density) onto the defined grid. Accumulate and average the values in each grid cell across all sampled configurations [76].
Optional: Local Averaging for Phase Identification
- To enhance contrast between coexisting phases, apply a 3-dimensional moving average to the raw density grid. The size of the averaging domain should be based on the characteristic length scale of the phases [76].
Visualization and Analysis in ParaView
- Export the final density grid in a format compatible with ParaView (e.g., VTK).
- Use ParaView's visualization tools, such as isosurface rendering, to visualize the density field and identify key structural features [76].
Protocol B: Implementing an Atom-Based ML-DFT Workflow
This protocol outlines the steps for training a deep learning model to predict electron density and related properties using an atom-based representation, as demonstrated in state-of-the-art ML-DFT emulation [2].
Materials and Software Requirements
- Reference Database: A set of atomic structures (molecules, polymers, crystals) with corresponding properties calculated using ab initio DFT (e.g., using VASP) [2].
- Fingerprinting Code: Software to compute atomic fingerprints (e.g., AGNI fingerprints) [2].
- ML Framework: A deep learning framework (e.g., TensorFlow, PyTorch) for building and training neural networks.
Step-by-Step Procedure
Database Curation and Fingerprinting
- Assemble a diverse dataset of atomic structures and perform DFT calculations to obtain reference electronic properties (charge density, density of states, forces, energy) [2].
- For each atom in every structure, compute a rotation-invariant descriptor of its local chemical environment. The AGNI fingerprint is a suitable choice, which sums over Gaussian functions to represent the atomic environment [2].
Model Architecture and Charge Density Prediction
- Step 1 - Predict Charge Density Descriptors: Design a deep neural network that takes the AGNI atomic fingerprints as input and outputs the parameters (exponents and coefficients) for a set of Gaussian-type orbitals (GTOs) that describe the atom's electron density. The model learns the optimal basis set from the data [2].
- Coordinate Transformation: For each atom, define a local orthonormal coordinate system using its two nearest neighbors. Use the transformation matrix from this local system to the global Cartesian system to rotate the predicted GTOs to their correct spatial orientation [2].
- Density Reconstruction: Sum the contributions from all atom-centered GTOs to reconstruct the total electron density of the system on any desired grid for analysis [2].
Property Prediction from Atomic Structure and Density
- Step 2 - Predict Other Properties: Build a second neural network that uses both the original atomic fingerprints and the predicted electron density descriptors as input. This network can then predict a wide range of electronic and atomic properties, such as the density of states, total potential energy, atomic forces, and stress tensor [2]. This two-step approach is consistent with the first principles of DFT and has been shown to improve accuracy and transferability.
Model Validation and Deployment
- Test the model on an independent set of structures not seen during training.
- Deploy the trained model for high-throughput screening or molecular dynamics simulations, leveraging its linear scaling with system size and small computational prefactor [2].
Table 2: Key software tools and computational methods for density representation research.
| Item Name | Type | Function in Research |
|---|---|---|
| GROMACS | Software MD Package | Used to run molecular dynamics simulations and generate trajectory files for grid-based density sampling [76]. |
| ParaView | Software Visualization Tool | Open-source platform for visualizing and analyzing the 3D property grids generated from density sampling [76]. |
| VASP | Software DFT Code | Used to compute reference data (charge density, energies, forces) for training machine learning models like ML-DFT [2]. |
| AGNI Fingerprints | Computational Method | Rotation-invariant atomic descriptors that encode the local chemical environment; used as input for atom-based ML models [2]. |
| ROCS | Software | A widely used program for 3D molecular shape comparison that uses Gaussian functions to represent molecular volume and calculate shape similarity [77]. |
| Gaussian-Type Orbitals (GTOs) | Mathematical Basis Set | Functions used to represent the atomic electron density in atom-based representations; their parameters are learned by the ML model [2]. |
Workflow Visualizations
The integration of machine learning (ML) with density functional theory (DFT) is revolutionizing computational materials science and drug discovery. This fusion addresses one of the most significant challenges in the field: balancing quantum-level accuracy with computational tractability for complex, real-world systems. While DFT serves as the workhorse for quantum mechanical calculations, its traditional limitations in accuracy for certain chemical systems and the high computational cost for large-scale screening have persisted. ML-accelerated workflows now present a viable path forward, but their reliability hinges on rigorous validation across diverse chemical domains. This application note examines the performance and validation of these hybrid DFT-ML approaches across molecules, polymers, and crystalline materials, providing structured data, experimental protocols, and key reagent solutions for researchers.
Performance on Molecules
Validation Data and Performance
The application of ML-improved DFT to molecular systems demonstrates significant advancements in accuracy while maintaining manageable computational costs. Vikram Gavini's team at the University of Michigan has pioneered an approach that uses machine learning to discover more universal exchange-correlation (XC) functionals, creating a crucial bridge between the accuracy of quantum many-body (QMB) methods and the simplicity of DFT [4].
Table 1: Performance Metrics of ML-Improved DFT for Molecular Systems
| Validation Metric | Traditional DFT with Approximated XC | ML-Improved DFT (Gavini et al.) | Assessment Method |
|---|---|---|---|
| Generalizability | Works for spotting trends but unreliable for precise predictions | Works beyond training set; accurate for systems different from training data | Testing on systems not included in training [4] |
| Training Data Efficiency | N/A (pre-defined functionals) | High performance with data from only 5 atoms and 2 simple molecules | Comparison of accuracy vs. amount of QMB training data [4] |
| Physical Soundness | Varies by approximation; can produce unphysical results | Avoids unphysical results by adhering to DFT rules | Evaluation of output adherence to physical constraints [4] |
This approach differs fundamentally from earlier attempts by training ML models not only on the interaction energies of electrons but also on the potentials that describe how that energy changes at each point in space [4]. This provides a stronger foundation for training as potentials highlight small system differences more effectively than energies alone.
Experimental Protocol for Molecular Validation
Protocol 1: Validating ML-Improved XC Functionals on Molecular Systems
- Objective: To train and validate a machine-learned XC functional for accurate molecular modeling.
- Materials/Software: Access to high-performance computing (HPC) resources; quantum many-body (QMB) calculation software (e.g., for coupled cluster calculations); DFT code; ML training framework.
- Procedure:
- Reference Data Generation: Perform high-fidelity QMB calculations on a small, curated set of atoms and simple molecules (e.g., 5 atoms, 2 molecules) to obtain exact energies and potentials [4].
- Model Training: Train a machine learning model using this compact dataset. The model should learn to approximate the XC functional, with training incorporating both the QMB energies and the spatially resolved potentials [4].
- Accuracy Benchmarking: Apply the newly trained ML functional to DFT calculations for molecular systems. Compare the results for formation energies, bond lengths, and other properties against both standard DFT approximations (e.g., PBE) and higher-level QMB reference data not used in training [4].
- Generalizability Test: Evaluate the trained model on molecular systems that are chemically distinct from those in the training set to assess its transferability [4].
- Validation Notes: A successful model will match or outperform widely used XC approximations while maintaining low computational costs and producing physically meaningful results across diverse molecules [4].
Performance on Polymers
Validation Data and Performance
Machine learning has demonstrated profound utility in predicting polymer properties, moving beyond traditional trial-and-error approaches. A landmark study used a deep neural network (DNN) model to establish a structure-property correlation for the glass transition temperature (T_g), a critical parameter determining polymer application temperature ranges [78].
Table 2: ML Performance in Predicting Polymer Glass Transition Temperature (T_g)
| Validation Aspect | Data and Methodology | Performance Outcome | Validation Method |
|---|---|---|---|
| Model Training & Prediction | DNN trained on ~6,923 experimental T_g values from PoLyInfo database using Morgan fingerprint representations [78]. | Reasonable prediction of unknown T_g values for polymers with distinct molecular structures [78]. | Comparison of ML predictions with experimental results and molecular dynamics simulations [78]. |
| High-Throughput Screening | Screening of nearly one million hypothetical polymers [78]. | Identification of >65,000 candidates with T_g > 200°C, vastly expanding the known space of high-temperature polymers [78]. | Comparative analysis against existing known high-temperature polymers (~2,000 in PoLyInfo) [78]. |
Beyond predictive screening, validation extends to real-world synthesis. For instance, an organic-inorganic composite scale inhibitor (CT-5) was synthesized based on monomer selection principles, and its high thermal stability (decomposition temperature of 235.24°C) was confirmed through experimental characterization including FTIR, XRD, and TG-DTG [79].
Experimental Protocol for Polymer Discovery
Protocol 2: High-Throughput Screening of Polymers via ML
- Objective: To rapidly discover new high-temperature polymers using a trained ML model.
- Materials/Software: A large and diverse polymer database (e.g., PoLyInfo); ML framework capable of handling deep neural networks; molecular featurization software (e.g., for generating Morgan fingerprints).
- Procedure:
- Data Curation: Collect a diverse set of known homopolymers with experimentally measured Tg values. A foundation of nearly 13,000 real homopolymers is recommended [78].
- Model Formulation and Training: Train a Deep Neural Network (DNN) model using thousands of experimental Tg values. Use a molecular structure representation like Morgan fingerprints as the feature input [78].
- Model Validation: Validate the model's transferability and generalization ability by comparing its predictions against independent experimental data and/or high-fidelity molecular dynamics simulations for polymers not seen during training [78].
- Virtual Screening: Apply the validated ML model to a large library of hypothetical polymers (e.g., nearly one million candidates) to predict their Tg values [78].
- Candidate Selection: Identify and prioritize candidates that exceed a target property threshold (e.g., Tg > 200°C) for further experimental investigation [78].
- Validation Notes: The success of the workflow is confirmed when a large number of promising candidates are identified, and a subset of these is successfully synthesized and characterized, matching the ML-predicted properties [78].
Performance on Crystals
Validation Data and Performance
For crystalline materials, the development of universal machine learning interatomic potentials (MLIPs) is a key focus. Their validation relies on benchmarking across multiple challenging scenarios. The MP-ALOE dataset, containing nearly 1 million DFT calculations using the accurate r2SCAN meta-GGA, provides a robust foundation for training and testing such models [52].
Table 3: Benchmarking Performance of MLIPs Trained on r2SCAN Data for Crystals
| Benchmark Category | Benchmark Description | Key Finding (MP-ALOE Trained Model) | Implication |
|---|---|---|---|
| Equilibrium Properties | Predicting formation energies and structural properties of ~1000 equilibrium structures from the WBM dataset [52]. | Competitive accuracy in predicting equilibrium energies [52]. | Reliable for calculating standard thermochemical properties. |
| Off-Equilibrium Forces | Predicting forces in far-from-equilibrium structures [52]. | Competitive performance in predicting off-equilibrium forces [52]. | Accurate for modeling defects, reactions, and other non-equilibrium processes. |
| Static Extreme Conditions | Behavior under extreme hydrostatic pressure [52]. | Improved stability and physical soundness of the potential energy surface [52]. | More robust for simulating high-pressure phases. |
| Dynamic Stability | Molecular dynamics (MD) stability under extreme temperatures and pressures [52]. | Improved stability in MD runs under extreme ensemble conditions [52]. | Enables reliable and longer-time MD simulations in harsh conditions. |
The MP-ALOE dataset itself was validated by its content, showing a wider distribution of cohesive energies, forces, and pressures compared to earlier datasets like MatPES, ensuring the trained MLIPs encounter a broader range of physical scenarios [52].
Experimental Protocol for Crystalline Materials
Protocol 3: Benchmarking a Universal ML Interatomic Potential
- Objective: To evaluate the performance and reliability of a trained MLIP across a suite of standardized tests.
- Materials/Software: A trained MLIP (e.g., a MACE model); DFT code; molecular dynamics simulation package; benchmark datasets (e.g., WBM for equilibrium structures).
- Procedure:
- Equilibrium Properties Test:
- Off-Equilibrium Forces Test:
- Generate or select structures that are far from equilibrium (e.g., with strained bonds or defects).
- Calculate forces on atoms in these structures using both the MLIP and direct DFT.
- Quantify the error of the MLIP-predicted forces versus the DFT reference [52].
- Static Extreme Deformations Test:
- Apply extreme hydrostatic pressures to crystal structures in a static calculation.
- Evaluate whether the MLIP produces physically sound potential energy surfaces or exhibits unphysical behavior [52].
- Molecular Dynamic Stability Test:
- Run MD simulations at extreme temperatures and pressures using the MLIP.
- Monitor for simulation crashes or unphysical structural disintegration to assess stability and robustness [52].
- Validation Notes: A high-quality UMLIP should perform competitively on all benchmarks, demonstrating accuracy, robustness, and transferability across different states of matter and conditions [52].
Table 4: Key Resources for DFT-ML Workflows
| Resource Name/Type | Function/Purpose | Example(s) |
|---|---|---|
| High-Performance Computing (HPC) | Provides the computational power required for DFT calculations and training large ML models. | Local clusters, national supercomputing centers, cloud computing platforms. |
| DFT Codes | Software to perform the foundational quantum mechanical calculations that generate training data and reference values. | VASP, Quantum ESPRESSO, CASTEP [52]. |
| Machine Learning Frameworks | Libraries and tools for building, training, and deploying machine learning models. | PyTorch, TensorFlow, JAX [52]. |
| Materials Databases | Curated repositories of computed and/or experimental material properties used for training and validation. | Materials Project (MP) [52], PoLyInfo (for polymers) [78], Alexandria [52]. |
| Accurate DFT Datasets | Large-scale datasets calculated at high levels of theory, used for training more reliable MLIPs. | MP-ALOE (r2SCAN) [52], MatPES (r2SCAN) [52]. |
| MLIP Architectures | Graph-based neural network models designed to represent atomic systems and learn potential energy surfaces. | MACE [52], M3GNet [52]. |
| Benchmarking Suites | Standardized sets of tests and structures to consistently evaluate the performance of different computational methods. | WBM dataset for equilibrium properties [52]. |
Workflow Visualization
The following diagram illustrates a robust, validated DFT-ML workflow for materials discovery, integrating the key stages of data generation, model training, and multi-faceted validation discussed in this note.
The integration of machine learning (ML) with density functional theory (DFT) is fundamentally reshaping the landscape of computational materials science and drug development. This paradigm shift moves research beyond traditional, manually intensive simulation methods toward intelligent, self-correcting systems. Autonomous workflows, capable of managing complex calculation sequences with minimal human intervention, are now being enhanced by ML-driven sensitivity analysis. This powerful combination allows researchers to not only automate tasks but also to understand and optimize the critical parameters governing their simulations [17] [15]. This document outlines application notes and detailed protocols for implementing these advanced techniques, providing a framework for robust, reproducible, and accelerated discovery.
Application Notes
The Architecture of Autonomous DFT Workflows
Autonomous DFT workflows are sophisticated computational frameworks designed to execute multi-step simulation and analysis processes with high efficiency and minimal manual input. Their development is driven by the need for high-throughput screening, reliable defect characterization, and the generation of large datasets for machine learning interatomic potentials (MLIPs) [17].
Core Components and Design Principles: These workflows are built on a foundation of several key principles:
- Modularity: Workflows are composed of reusable, interchangeable components (e.g., for structure relaxation, property calculation, or convergence testing) [17].
- Engine Agnosticism: Protocols are designed to be independent of the specific DFT code used. Frameworks like AiiDA implement engine-agnostic interfaces, allowing the same workflow to run seamlessly across different quantum engines such as Quantum ESPRESSO, VASP, and CASTEP [17] [80].
- Provenance Tracking: Every piece of data, along with all inputs, parameters, and computational steps that produced it, is meticulously recorded. This ensures full reproducibility and auditability of the entire research process [17] [80].
- Robust Error Handling: Workflows incorporate intelligent error handlers (e.g., via tools like Custodian) to manage common calculation failures, such as electronic self-consistency field (SCF) non-convergence, by automatically adjusting parameters and restarting jobs [17].
Interoperability via Universal Standards: A significant challenge in automated workflows is the inconsistency between different software packages. This is being addressed by the development of universal input/output schemas and APIs, such as the OPTIMADE API. These standards allow workflow managers to translate data into code-specific formats internally, enabling true cross-code interoperability and validation [80].
The Role of Sensitivity Analysis in Workflow Optimization
Sensitivity analysis has emerged as a critical tool for enhancing the efficiency and reliability of autonomous workflows. It quantitatively identifies which input parameters most significantly impact the output of a DFT simulation, guiding resource allocation and preventing overfitting in ML-integrated workflows [81].
HSIC: A Kernel-Based Sensitivity Metric: Modern implementations, such as those in the ParAMS software, utilize the Hilbert-Schmidt Independence Criterion (HSIC). The HSIC is a robust, kernel-based statistical measure used to quantify the dependence between input parameters and the resulting loss function or target property [81].
- Calculation Method: The sensitivity for a parameter is calculated by drawing numerous uniform random samples of parameter values, computing the loss function for each set, and then applying the HSIC metric to these pairs of values. This process reveals the strength of the dependency [81].
- Interpretation: The resulting sensitivity values are normalized to sum to one, ranging from 0 (completely insensitive) to 1 (highly sensitive). This intuitive scale allows researchers to immediately identify the parameters that warrant closer scrutiny or inclusion in an optimization loop [81].
Application in Parameter Selection: In complex force field reparameterization or ML model training, it is difficult to know a priori which parameters to optimize. Including too many parameters slows down convergence and increases the risk of overfitting, while too few may limit model accuracy. Sensitivity analysis directly addresses this by pinpointing the most influential parameters, enabling leaner, more effective optimizations [81].
Synergy with Machine Learning Interatomic Potentials
The creation of accurate MLIPs is a primary application driving the adoption of autonomous workflows. MLIPs aim to achieve the accuracy of ab initio methods at a fraction of the computational cost, enabling large-scale and long-time-scale molecular dynamics simulations [8] [82].
Autonomous workflows manage the intricate process of generating training data through high-throughput DFT calculations. They automate structure selection, execute the necessary simulations, and handle error correction. The resulting datasets are then used to train MLIPs, which can be categorized into families such as "general graph-network," "symmetry-equivariant," and "extreme-efficiency" models, each with different trade-offs in accuracy, cost, and scope [8].
Sensitivity analysis contributes to this pipeline by helping to refine the feature space, or descriptors, used by the ML models. By identifying which descriptors (e.g., geometric, electronic structure, or intrinsic elemental properties) have the strongest influence on predicting a target property, researchers can build more efficient and accurate models [8].
Table: Categories of Descriptors for ML in Electrocatalysis
| Descriptor Category | Description | Examples | Computational Cost | Primary Use |
|---|---|---|---|---|
| Intrinsic Statistical | Fundamental, readily available elemental properties. | Magpie attributes, valence-electron count, ionization energy. | Very Low | Rapid, wide-angle coarse screening of chemical space. |
| Electronic Structure | Quantum mechanical quantities from DFT. | d-band center, orbital occupation, magnetic moments, Bader charges. | High (requires DFT) | Fine screening and mechanistic analysis. |
| Geometric/Microenvironmental | Local structural and chemical environment. | Coordination number, interatomic distances, local strain, site indices. | Low to Moderate | Capturing structure-activity trends in complex supports. |
Experimental Protocols
Protocol: Implementing an Engine-Agnostic Relaxation Workflow
This protocol describes how to set up a CommonRelaxWorkChain, a type of autonomous workflow that performs structure relaxation (geometry optimization) using a standardized input schema that can be executed across multiple DFT engines [17].
1. Workflow Configuration and Input Preparation
- Select a Workflow Manager: Choose a framework that supports engine-agnostic protocols, such as AiiDA.
- Define the Calculation Protocol: Specify the desired level of precision (e.g., "fast," "moderate," or "precise"). This will automatically set convergence thresholds for forces, stresses, and energies.
- Prepare the Input Structure: Provide the initial atomic structure in a standard format (e.g., CIF, POSCAR). The workflow will handle conversion to the target engine's format.
- Select Quantum Engines: Specify which DFT codes to use (e.g., CP2K, CASTEP, VASP). The workflow's universal schema ensures consistent settings across codes.
2. Job Submission and Execution Control
- Configure HPC Resources: Define computational resources (e.g., number of CPUs, memory, wall time) and the scheduler (SLURM, PBS).
- Launch the Workflow: Submit the job. The workflow manager will generate all necessary input files, submit the calculation to the queue, and begin monitoring.
- Enable Error Handling: The workflow will automatically detect and attempt to recover from common failures (e.g., SCF non-convergence, wall-time expiration) by adjusting mixer settings or restarting from the last checkpoint.
3. Post-Processing and Validation
- Extract Properties: Upon successful completion, the workflow will parse outputs to extract key properties, including the relaxed atomic structure, total energy, forces, and stress tensor.
- Cross-Code Validation (Optional): To validate results, run the same relaxation protocol using two different DFT engines. Compare the final energies and structures to assess consistency and identify any code-specific idiosyncrasies [80].
- Provenance Recording: All inputs, outputs, and computational steps are automatically stored in a queryable database with full provenance.
Engine-Agnostic DFT Relaxation Workflow
Protocol: Conducting Parameter Sensitivity Analysis for a ReaxFF Training Set
This protocol uses the ParAMS software package to perform a sensitivity analysis, identifying the most sensitive parameters in a ReaxFF force field for a given training set [81].
1. Initial Setup and Parameter Selection
- Load the Training Set: Begin with a set of reference data (e.g., bond lengths, angles, reaction energies, etc.) that defines the loss function.
- Activate Parameters: Select all parameters you wish to investigate for sensitivity. It is preferable to start with a broad set, including potentially irrelevant parameters, as the analysis will naturally filter them out.
2. Generating and Running the Sensitivity Calculation
- Configure Sampling: In the ParAMS input panel, navigate to the sensitivity settings. Set the number of samples (e.g., 2000) to generate. A larger number of samples improves statistical reliability.
- Load or Generate Samples:
- To save time, you can load a pre-computed set of parameter samples and their corresponding loss values from a directory.
- Alternatively, set
RunSampling Yesto generate new samples by drawing uniform random values for the active parameters and computing the loss for each set.
- Set Analysis Parameters:
- Set the number of calculation repeats (e.g., 5) and the number of samples per repeat (e.g., 500). This uses subsets of the total samples to assess the robustness of the sensitivity rankings.
- Keep the default kernel settings (Gaussian for parameters, conjunctive-Gaussian for loss) for most scenarios.
3. Interpreting Results and Refining the Model
- Review Sensitivity Plots: Examine the generated scatter plot, which shows the sensitivity value for each parameter across all repeats. Parameters with consistently high average sensitivity (red line) are the most influential.
- Sort Parameters by Sensitivity: In the Parameters panel, sort the list by sensitivity in decreasing order. The values, summing to 1, indicate each parameter's relative importance.
- Downselect for Optimization: Use the results to deactivate insensitive parameters (sensitivity ~0), creating a smaller, more effective set for subsequent force field optimization, which reduces overfitting risk and improves convergence speed.
Table: Key Inputs for a ReaxFF Sensitivity Analysis in ParAMS
| Input Option | Setting | Purpose & Rationale |
|---|---|---|
| RunSampling | Yes / No |
Generate new parameter samples or load existing ones. |
| NumberSamples | e.g., 2000 |
Size of the initial parameter space sample pool. |
| SaveResiduals | No |
Saves disk space unless detailed per-calculation error data is needed. |
| Repeat calculation | e.g., 5 |
Number of subset analyses to run; checks result stability. |
| Samples per repeat | e.g., 500 |
Size of each subset drawn from the full sample pool. |
| Filter infinite values | Yes |
Removes non-converged parameter sets from the analysis. |
Parameter Sensitivity Analysis Workflow
The Scientist's Toolkit
Table: Essential Research Reagents and Software Solutions
| Tool Name / Category | Function / Description | Application in Autonomous Workflows |
|---|---|---|
| AiiDA | A robust workflow manager and provenance tracking platform. | Orchestrates complex, multi-step computational workflows, ensuring reproducibility and handling job submission across HPC schedulers [17]. |
| JARVIS-Tools | A integrated framework for high-throughput DFT and ML. | Provides automated job management, robust error handling, and a extensive database of computed materials properties [17]. |
| ParAMS | A software package for parameter optimization and sensitivity analysis. | Used for identifying the most sensitive parameters in force fields or ML models using the HSIC metric, guiding efficient optimization [81]. |
| OPTIMADE API | A universal API for exchanging materials data. | Enables interoperability between different databases and workflow managers, facilitating engine-agnostic calculations [80]. |
| Pymatgen & ASE | Python libraries for materials analysis and atomistic simulations. | Core utilities for structure manipulation, file format conversion, and analysis within automated workflows [17]. |
| MLIP Families | Machine learning interatomic potentials (e.g., MACE, NequIP). | Provide quantum-accurate forces and energies for large-scale molecular dynamics, trained on data from autonomous DFT workflows [8] [82]. |
Conclusion
The synergy between Machine Learning and Density Functional Theory marks a paradigm shift in computational science, moving beyond mere acceleration to a more profound and complete emulation of quantum mechanics. By learning the fundamental mappings from atomic structure to electronic density and properties, ML-DFT workflows achieve unprecedented efficiency while maintaining chemical accuracy, as demonstrated across extensive molecular and material databases. For biomedical researchers and drug development professionals, this integration promises to drastically shorten development timelines, reduce costs, and unlock the exploration of complex biological systems previously beyond computational reach. Future progress hinges on enhancing model interpretability, expanding to heavier elements and solid-state systems, and seamlessly integrating these powerful in silico tools into the Model-Informed Drug Development (MIDD) pipeline. As data quality and algorithms continue to advance, ML-DFT is poised to become an indispensable, predictive engine for the design of next-generation therapeutics and biomaterials.
References
- 1. Unsupervised representation learning of Kohn–Sham ... [https://www.nature.com/articles/s41467-024-53748-7]
- 2. A deep learning framework to emulate density functional ... [https://www.nature.com/articles/s41524-023-01115-3]
- 3. Density Functional Theory [https://www.microsoft.com/en-us/research/project/dft/]
- 4. Machine learning helps density functional theory model ... [https://cen.acs.org/physical-chemistry/modeling/Machine-learning-helps-density-functional/103/web/2025/10]
- 5. Overcoming the Chemical Complexity Bottleneck in on-the- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11270744/]
- 6. Machine learning for improved density functional theory ... [https://www.nature.com/articles/s41598-025-02088-7]
- 7. EMFF-2025: a general neural network potential for ... [https://www.nature.com/articles/s41524-025-01809-w]
- 8. From descriptors to machine learning interatomic potentials [https://www.oaepublish.com/articles/aiagent.2025.05]
- 9. Timeline: The continuing evolution of density functional ... [https://www.microsoft.com/en-us/research/articles/timeline-the-continuing-evolution-of-density-functional-theory/]
- 10. Kohn–Sham time-dependent density functional theory with ... [https://www.nature.com/articles/s41524-023-01041-4]
- 11. Density functional theory - Wikipedia [https://en.wikipedia.org/wiki/Density_functional_theory]
- 12. Accuracy of Charge Densities in Electronic Structure ... [https://arxiv.org/html/2410.17866v4]
- 13. Fundamentals of Density Functional Theory: Recent Developments, Challenges and Future Horizons | IntechOpen [https://www.intechopen.com/chapters/78124]
- 14. Density functional theory across chemistry, physics and biology - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3928866/]
- 15. Applications of density functional theory and machine ... [https://www.sciencedirect.com/science/article/pii/S2949822825002011]
- 16. Machine Learning Difference Charge Density [https://arxiv.org/html/2510.00788v1]
- 17. Automated DFT Workflows [https://www.emergentmind.com/topics/automated-dft-workflows]
- 18. The Year of Quantum: From concept to reality in 2025 [https://www.mckinsey.com/capabilities/tech-and-ai/our-insights/the-year-of-quantum-from-concept-to-reality-in-2025]
- 19. Quantum computing in life sciences and drug discovery [https://www.mckinsey.com/industries/life-sciences/our-insights/the-quantum-revolution-in-pharma-faster-smarter-and-more-precise]
- 20. Top quantum breakthroughs of 2025 [https://www.networkworld.com/article/4088709/top-quantum-breakthroughs-of-2025.html]
- 21. Quantum Machines Launches QUAlibrate [https://www.quantum-machines.co/press-release/quantum-machines-launches-qualibrate-an-open-source-framework-that-cuts-quantum-computer-calibration-from-hours-to-minutes/]
- 22. A new quantum toolkit for optimization [https://research.google/blog/a-new-quantum-toolkit-for-optimization/]
- 23. Quantum Machine Learning: The Complete Guide for 2025 [https://morsoftware.com/blog/quantum-machine-learning]
- 24. Quantum Machine Learning Explained: The Ultimate Guide [https://www.guvi.in/blog/quantum-machine-learning/]
- 25. Quantum Machine Learning Algorithms [https://meritshot.com/blogs/quantum-machine-learning-algorithms]
- 26. Predicting electronic structures at any length scale with ... [https://www.nature.com/articles/s41524-023-01070-z]
- 27. Application of Density Functional Theory to Molecular ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11989887/]
- 28. Role of DFT in Drug Design: A Mini Review [https://www.longdom.org/open-access/role-of-dft-in-drug-design-a-mini-review-93540.html]
- 29. General — Workflows 2025.1 documentation - SCM [https://www.scm.com/doc/Workflows/SimpleActiveLearning/General.html]
- 30. Neural network interpolation of exchange-correlation ... [https://www.nature.com/articles/s41598-020-64619-8]
- 31. Application of two-component neural network for exchange ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9391383/]
- 32. Grad DFT: a software library for machine learning ... [https://arxiv.org/html/2309.15127v2]
- 33. Automatic selection of atomic fingerprints and reference configurations for machine-learning potentials - PubMed [https://pubmed.ncbi.nlm.nih.gov/29960368/]
- 34. Density-of-states similarity descriptor for unsupervised ... [https://www.nature.com/articles/s41597-022-01754-z]
- 35. AL4GAP: Active learning workflow for generating DFT ... [https://pubmed.ncbi.nlm.nih.gov/37428051/]
- 36. Machine learning with new functional structure descriptors ... [https://pubs.rsc.org/en/content/articlelanding/2025/cp/d5cp01972a]
- 37. Machine Learning in Drug Discovery: A Review - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8356896/]
- 38. Practically significant method comparison protocols for ... [https://chemrxiv.org/engage/chemrxiv/article-details/672a91bd7be152b1d01a926b]
- 39. Applications of density functional theory in COVID-19 drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8695517/]
- 40. Application of density functional theory (DFT) and response ... [https://www.sciencedirect.com/science/article/pii/B9780323905930000046]
- 41. Machine learning small molecule properties in drug ... [https://www.sciencedirect.com/science/article/pii/S2949747723000209]
- 42. A Machine Learning Pipeline for Molecular Property ... [https://arxiv.org/html/2505.08688v1]
- 43. Label-free detection of cellular drug responses by high- ... [https://www.nature.com/articles/s41598-017-12378-4]
- 44. Applications of machine learning in drug discovery and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6552674/]
- 45. Machine Learning and Assay Development for Image-based ... [https://www.ncbi.nlm.nih.gov/books/NBK589577/]
- 46. Integrating density functional theory with machine learning ... [https://pubs.rsc.org/en/content/articlehtml/2025/cp/d4cp03397c]
- 47. A Comprehensive Review of Machine-Learning ... [https://www.mdpi.com/2073-4352/15/11/925]
- 48. Towards A Universally Transferable Acceleration Method ... [https://arxiv.org/html/2509.25724v1]
- 49. Efficiency, accuracy, and transferability of machine learning ... [https://www.sciencedirect.com/science/article/pii/S135964542400140X]
- 50. OMol25: High-Precision Molecular DFT Dataset [https://www.emergentmind.com/topics/omol25-dataset]
- 51. Data Readiness for Scientific AI at Scale [https://arxiv.org/html/2507.23018v1]
- 52. MP-ALOE: an r2SCAN dataset for universal machine ... [https://www.nature.com/articles/s41524-025-01834-9]
- 53. Highly transferable atomistic machine-learning potentials ... [https://pubs.rsc.org/en/content/articlelanding/2023/dd/d3dd00046j]
- 54. Understanding DFT Uncertainties for More Reliable Reactivity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12529897/]
- 55. Mitigating error cancellation in density functional ... [https://arxiv.org/html/2504.14961v1]
- 56. Machine learning to tame divergent density functional ... [https://pubs.rsc.org/en/content/articlehtml/2021/sc/d1sc03701c]
- 57. How Accurate Are DFT Forces? Unexpectedly Large ... [https://arxiv.org/html/2510.19774v1]
- 58. Chemical property prediction under experimental biases [https://www.nature.com/articles/s41598-022-12116-5]
- 59. Advancing drug development with "Fit-for-Purpose" ... [https://pubmed.ncbi.nlm.nih.gov/40952534/]
- 60. Machine learning to tame divergent density functional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8513898/]
- 61. Massive Atomic Diversity: a compact universal dataset for ... [https://www.nature.com/articles/s41597-025-06109-y]
- 62. The OMol25 Dataset [https://www.faccts.de/omol25-dataset/]
- 63. Putting Density Functional Theory to the Test in Machine- ... [http://hjkgrp.mit.edu/publication/duan-putting-2021/]
- 64. Predicting and understanding human action decisions ... [https://www.nature.com/articles/s41598-023-31807-1]
- 65. Density Functional Theory and Artificial Intelligence ... [https://www.cecam.org/workshop-details/density-functional-theory-and-artificial-intelligence-learning-from-each-other-1281]
- 66. Machine Learning Approaches for Diagnostics and ... [https://arxiv.org/html/2312.16810v3]
- 67. Correlation vs. Causation: How Causal AI is Helping ... [https://globalforum.diaglobal.org/issue/october-2024/correlation-vs-causation-how-causal-ai-is-helping-determine-key-connections-in-healthcare-and-clinical-trials/]
- 68. Machine learning for accuracy in density functional ... [https://arxiv.org/html/2311.00196v2]
- 69. Quantum Chemistry: Making Key Simulation Approach ... [https://thequantuminsider.com/2025/09/22/quantum-chemistry-making-key-simulation-approach-more-accurate/]
- 70. Exploring Meta's Open Molecules 2025 (OMol25) & Universal ... [https://www.rowansci.com/blog/exploring-open-molecules-2025]
- 71. Benchmark of Approximate Quantum Chemical and Machine ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12288007/]
- 72. Benchmarking Data Efficiency in Advanced Machine ... [https://www.peter-zaspel.de/?p=1539]
- 73. Machine learning helps improve accuracy and efficiency of small-molecule calculations [https://cen.acs.org/physical-chemistry/computational-chemistry/Machine-learning-helps-improve-accuracy/103/web/2025/06]
- 74. Exploring an accurate machine learning model to quickly ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10960145/]
- 75. MatSci-ML Studio: an interactive workflow toolkit for ... [https://www.oaepublish.com/articles/jmi.2025.45]
- 76. Density based visualization for molecular simulation [https://pubs.rsc.org/en/content/articlehtml/2014/fd/c3fd00124e]
- 77. Three-Dimensional Compound Comparison Methods and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5005041/]
- 78. Machine learning discovery of high-temperature polymers [https://www.sciencedirect.com/science/article/pii/S2666389921000398]
- 79. Preparation and performance evaluation of polymer ... [https://www.nature.com/articles/s41598-025-99968-9]
- 80. Dft Workflow Interoperability Enables Engine-Agnostic ... [https://quantumzeitgeist.com/codes-dft-workflow-interoperability-enables-engine-agnostic-calculations-across/]
- 81. 5.5. Parameter sensitivity analysis - SCM [https://www.scm.com/doc/params/examples/sensitivity.html]
- 82. Advancing materials discovery through artificial intelligence [https://www.sciencedirect.com/science/article/pii/S2352940725003981]